| Entry | Database: PDB / ID: 2cgi
|
|---|
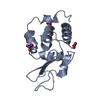
| Title | Siras structure of tetragonal lysozyme using derivative data collected at the high energy remote Holmium Kedge |
|---|
 Components Components | LYSOZYME C |
|---|
 Keywords Keywords | HYDROLASE / HIGH ENERGY / PHASING / MAD / SAD / SIRAS / RADIATION DAMAGE / ABSORPTION / HOLMIUM / GLYCOSIDASE / ALLERGEN / ANTIMICROBIAL / BACTERIOLYTIC ENZYME |
|---|
| Function / homology |  Function and homology information Function and homology information
Lactose synthesis / Antimicrobial peptides / Neutrophil degranulation / beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process / lysozyme / lysozyme activity / killing of cells of another organism / defense response to Gram-negative bacterium / defense response to bacterium ...Lactose synthesis / Antimicrobial peptides / Neutrophil degranulation / beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process / lysozyme / lysozyme activity / killing of cells of another organism / defense response to Gram-negative bacterium / defense response to bacterium / defense response to Gram-positive bacterium / endoplasmic reticulum / : / identical protein binding / cytoplasmSimilarity search - Function Lysozyme - #10 / Glycoside hydrolase, family 22, lysozyme / Glycoside hydrolase family 22 domain / Glycosyl hydrolases family 22 (GH22) domain signature. / Glycoside hydrolase, family 22 / C-type lysozyme/alpha-lactalbumin family / Glycosyl hydrolases family 22 (GH22) domain profile. / Alpha-lactalbumin / lysozyme C / Lysozyme / Lysozyme-like domain superfamily ...Lysozyme - #10 / Glycoside hydrolase, family 22, lysozyme / Glycoside hydrolase family 22 domain / Glycosyl hydrolases family 22 (GH22) domain signature. / Glycoside hydrolase, family 22 / C-type lysozyme/alpha-lactalbumin family / Glycosyl hydrolases family 22 (GH22) domain profile. / Alpha-lactalbumin / lysozyme C / Lysozyme / Lysozyme-like domain superfamily / Orthogonal Bundle / Mainly AlphaSimilarity search - Domain/homology |
|---|
| Biological species |   GALLUS GALLUS (chicken) GALLUS GALLUS (chicken) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SIRAS / Resolution: 1.35 Å SIRAS / Resolution: 1.35 Å |
|---|
 Authors Authors | Jakoncic, J. / Di Michiel, M. / Zhong, Z. / Honkimaki, V. / Jouanneau, Y. / Stojanoff, V. |
|---|
 Citation Citation |  Journal: J.Appl.Crystallogr. / Year: 2006 Journal: J.Appl.Crystallogr. / Year: 2006
Title: Anomalous Diffraction at Ultra-High Energy for Protein Crystallography.
Authors: Jakoncic, J. / Di Michiel, M. / Zhong, Z. / Honkimaki, V. / Jouanneau, Y. / Stojanoff, V. |
|---|
| History | | Deposition | Mar 7, 2006 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | Nov 13, 2006 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Nov 12, 2014 | Group: Database references / Derived calculations ...Database references / Derived calculations / Non-polymer description / Other / Source and taxonomy / Structure summary / Version format compliance |
|---|
| Revision 1.2 | Dec 28, 2016 | Group: Structure summary |
|---|
| Revision 1.3 | Jan 23, 2019 | Group: Data collection / Structure summary / Category: struct / Item: _struct.title |
|---|
| Revision 1.4 | Nov 20, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other / Structure summary
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_entry_details / pdbx_modification_feature / struct_site
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  SIRAS / Resolution: 1.35 Å
SIRAS / Resolution: 1.35 Å  Authors
Authors Citation
Citation Journal: J.Appl.Crystallogr. / Year: 2006
Journal: J.Appl.Crystallogr. / Year: 2006 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2cgi.cif.gz
2cgi.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2cgi.ent.gz
pdb2cgi.ent.gz PDB format
PDB format 2cgi.json.gz
2cgi.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/cg/2cgi
https://data.pdbj.org/pub/pdb/validation_reports/cg/2cgi ftp://data.pdbj.org/pub/pdb/validation_reports/cg/2cgi
ftp://data.pdbj.org/pub/pdb/validation_reports/cg/2cgi Links
Links Assembly
Assembly
 Components
Components
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  ESRF
ESRF  / Beamline: ID15A / Wavelength: 0.219
/ Beamline: ID15A / Wavelength: 0.219  Processing
Processing SIRAS / Resolution: 1.35→19.07 Å / Cor.coef. Fo:Fc: 0.965 / Cor.coef. Fo:Fc free: 0.955 / SU B: 2.053 / SU ML: 0.038 / Cross valid method: THROUGHOUT / ESU R: 0.073 / ESU R Free: 0.06 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
SIRAS / Resolution: 1.35→19.07 Å / Cor.coef. Fo:Fc: 0.965 / Cor.coef. Fo:Fc free: 0.955 / SU B: 2.053 / SU ML: 0.038 / Cross valid method: THROUGHOUT / ESU R: 0.073 / ESU R Free: 0.06 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS. Movie
Movie Controller
Controller

















































































































































































































































 PDBj
PDBj






