-Search & filter documents
 FAQ,
FAQ,  News,
News,  Information,
Information,  EM Navigator,
EM Navigator,  Yorodumi,
Yorodumi,  Omokage search,
Omokage search,  Services,
Services,  All docs
All docs
-News (20)
+Feb 9, 2022. New format data for meta-information of EMDB entries
New format data for meta-information of EMDB entries
- Version 3 of the EMDB header file is now the official format.
- The previous official version 1.9 will be removed from the archive.
Related info.: EMDB header
EMDB header
External links: wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
+Oct 5, 2021. Nobel Prize for mechanically activated and temperature-gated ion channels
Nobel Prize for mechanically activated and temperature-gated ion channels
- The Nobel Prize in Physiology or Medicine 2021 was awarded jointly to David Julius and Ardem Patapoutian "for their discoveries of receptors for temperature and touch."
- EM Navigator can help to find cryo-EM structure data by both pioneers.
External links: The Nobel Prize in Physiology or Medicine 2021 - NobelPrize.org /
The Nobel Prize in Physiology or Medicine 2021 - NobelPrize.org /  Structure data by Ardem Patapoutian /
Structure data by Ardem Patapoutian /  Structure data by David Julius
Structure data by David Julius
+Aug 12, 2020. Covid-19 info
Covid-19 info

URL: https://pdbj.org/emnavi/covid19.php
New page: Covid-19 featured information page in EM Navigator.
Related info.: Covid-19 info /
Covid-19 info /  Mar 5, 2020. Novel coronavirus structure data
Mar 5, 2020. Novel coronavirus structure data
+Mar 5, 2020. Novel coronavirus structure data
Novel coronavirus structure data
- International Committee on Taxonomy of Viruses (ICTV) defined the short name of the 2019 coronavirus as "SARS-CoV-2".
- In the structure databanks used in Yorodumi, some data are registered as the other names, "COVID-19 virus" and "2019-nCoV". Here are the details of the virus and the list of structure data.
Related info.: Yorodumi Speices /
Yorodumi Speices /  Aug 12, 2020. Covid-19 info
Aug 12, 2020. Covid-19 info
External links: COVID-19 featured content - PDBj /
COVID-19 featured content - PDBj /  Molecule of the Month (242):Coronavirus Proteases
Molecule of the Month (242):Coronavirus Proteases
+Jul 5, 2019. Downlodablable text data
Downlodablable text data

Some data of EM Navigator services can be downloaded as text file. Software such as Excel can load the data files.
| Page | Data | Format |
|---|---|---|
| EMN Search | search result | CSV, TSV, or JSON |
| EMN statistics | data table | CSV or TSV |
Related info.: EMN Search /
EMN Search /  EMN Statistics
EMN Statistics
+Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
EMDB accession codes are about to change! (news from PDBe EMDB page)
- The allocation of 4 digits for EMDB accession codes will soon come to an end. Whilst these codes will remain in use, new EMDB accession codes will include an additional digit and will expand incrementally as the available range of codes is exhausted. The current 4-digit format prefixed with “EMD-” (i.e. EMD-XXXX) will advance to a 5-digit format (i.e. EMD-XXXXX), and so on. It is currently estimated that the 4-digit codes will be depleted around Spring 2019, at which point the 5-digit format will come into force.
- The EM Navigator/Yorodumi systems omit the EMD- prefix.
Related info.: Q: What is EMD? /
Q: What is EMD? /  ID/Accession-code notation in Yorodumi/EM Navigator
ID/Accession-code notation in Yorodumi/EM Navigator
External links: EMDB Accession Codes are Changing Soon! /
EMDB Accession Codes are Changing Soon! /  Contact to PDBj
Contact to PDBj
+Feb 20, 2018. PDBj/BINDS workshop in Osaka University
PDBj/BINDS workshop in Osaka University
+Oct 4, 2017. Three pioneers of this field were awarded Nobel Prize in Chemistry 2017
Three pioneers of this field were awarded Nobel Prize in Chemistry 2017
- Jacques Dubochet (University of Lausanne, Switzerland) is a pioneer of ice-embedding method of EM specimen (as known as cryo-EM), Most of 3DEM structures in EMDB and PDB are obtained using his method.
- Joachim Frank (Columbia University, New York, USA) is a pioneer of single particle reconstruction, which is the most used reconstruction method for 3DEM structures in EMDB and EM entries in PDB. And also, he is a develper of Spider, which is one of the most famous software in this field, and is used for some EM Navigor data (e.g. map projection/slice images).
- Richard Henderson (MRC Laboratory of Molecular Biology, Cambridge, UK) was determined the first biomolecule structure by EM. The first EM entry in PDB, PDB-1brd is determinedby him.
External links: The 2017 Nobel Prize in Chemistry - Press Release
The 2017 Nobel Prize in Chemistry - Press Release
+Jul 12, 2017. Major update of PDB
Major update of PDB
- wwPDB released updated PDB data conforming to the new PDBx/mmCIF dictionary.
- This is a major update changing the version number from 4 to 5, and with Remediation, in which all the entries are updated.
- In this update, many items about electron microscopy experimental information are reorganized (e.g. em_software).
- Now, EM Navigator and Yorodumi are based on the updated data.
External links: wwPDB Remediation /
wwPDB Remediation /  Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
+Jun 16, 2017. Omokage search with filter
Omokage search with filter
Result of Omokage search can be filtered by keywords and the database types
Related info.: Omokage search
Omokage search
+Sep 15, 2016. EM Navigator & Yorodumi renewed
EM Navigator & Yorodumi renewed
New versions of EM Navigator and Yorodumi started
Related info.: Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+Aug 31, 2016. New EM Navigator & Yorodumi
New EM Navigator & Yorodumi
- In 15th Sep 2016, the development versions of EM Navigator and Yorodumi will replace the official versions.
- Current version will continue as 'legacy version' for some time.
Related info.: Changes in new EM Navigator and Yorodumi /
Changes in new EM Navigator and Yorodumi /  EM Navigator /
EM Navigator /  Yorodumi
Yorodumi
+Apr 13, 2016. Omokage search got faster
Omokage search got faster
- The computation time became ~1/2 compared to the previous version by re-optimization of data accession.
- Enjoy "shape similarity" of biomolecules, more!
Related info.: Omokage search
Omokage search
+Mar 3, 2016. Presentation PDF file for IPR seminar on Feb 19.
Presentation PDF file for IPR seminar on Feb 19.
External links: IPR seminar Feb 19th, 2016 /
IPR seminar Feb 19th, 2016 /  Presentation PDF
Presentation PDF
+Dec 4, 2015. The article about Omokage search is published online
The article about Omokage search is published online
Omokage search: shape similarity search service for biomolecular structures in both the PDB and EMDB. Suzuki Hirofumi, Kawabata Takeshi, and Nakamura Haruki, Bioinformatics. (2015) btv614
Related info.: Omokage search /
Omokage search /  gmfit
gmfit
External links: Main text (HTML, Open Access) /
Main text (HTML, Open Access) /  Supplementray data (PDF, Open Access)
Supplementray data (PDF, Open Access)
+Nov 28, 2015. Omokage search starts to support SASBDB models
Omokage search starts to support SASBDB models
- Models data in SASBDB are included in the Omokage search database.
- SASBDB is a databank for small angle scattering data.
- A SASBDB model can be used as a search query.
- The search result may include SASBDB models.
Related info.: Omokage search /
Omokage search /  SASBDB /
SASBDB /  Comparison of 3 databanks
Comparison of 3 databanks
+Dec 10, 2014. Multiple "PDB "SPLIT" entries are replaced with single "LARGE" entries
Multiple "PDB "SPLIT" entries are replaced with single "LARGE" entries
- Structure data deposited as multiple PDB entries are replace with single combined entries, which were previously stored as "large structure".
- In EM Navigator, many ribosome and several virus entries are replaced.
External links: Integration of Large Structures with the Main PDB Archive
Integration of Large Structures with the Main PDB Archive
+Sep 22, 2014. New service: Omokage search
New service: Omokage search
- Shape similarity search service, Omokage search has started.
- 形状類似検索「Omokage検索」を開始しました。
Related info.: Omokage search
Omokage search
+Mar 30, 2013. New page of EM Navigator, EMN Statistics
+Jan 16, 2013. New EMDB header format
New EMDB header format
- Format of the EMDB header file (XML based meta data file, not the map data themselves) are updated to version 1.9.
- Now, the contents of EM Navigator are based on the new data. Colors of some parts (e.g. bars in top area of the Detail pages) indicate 3D-reconstruction method, instead of "aggregation states".
External links: Announcement: EMDB header format update
Announcement: EMDB header format update
-FAQ (15)
+Q: How do you make the images for the structure data? What do their colors mean?
A: Images of EMDB entries are made for EM Navigator, and ones of PDB entries are for Yorodumi. There are several patterns of coloring.
See following items for the details.
Related info.: Q: How do you make the movies in EM Navigator? /
Q: How do you make the movies in EM Navigator? /  Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.? /
Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.? /  Q: How do you make the images of PDB entries in Yorodumi, Omokage, etc.?
Q: How do you make the images of PDB entries in Yorodumi, Omokage, etc.?
+Q: When the data are updated?
A: EMDB and PDB entries are released/updated every Wednesday at 0:00GMT/9:00JST
- Data in EM Navigator, Yorodumi, and Omokage are updated at the same time.
- SASBDB seems to update irregularly.
Related info.: Q: Where is the official data of EMDB?
Q: Where is the official data of EMDB?
External links: The release schedule of PDB data, the available data at each wwPDB site
The release schedule of PDB data, the available data at each wwPDB site
+Q: Is 3DEM same as electron cryo microscopy (cryoEM)?
A: No, "3DEM" and "cryoEM" are distinct to be exact.
- However, they are closely related. Some people call 3DEM "cryoEM".
- Many but not all the 3DEM analyses are perfomed with cryoEM. In EMDB and PDB, there are some entries, whose structure data were obtained in non-cryo contition.
Related info.: 3D electron microscopy (3DEM) /
3D electron microscopy (3DEM) /  electron cryo microscopy (cryoEM)
electron cryo microscopy (cryoEM)
+Q: Where is the official data of EMDB?
A: Following three are the official, have the same contents, and update at same time
| Country | Organization | HTTP URL | FTP URL |
|---|---|---|---|
| Japan | PDBj | https://ftp.pdbj.org/pub/emdb | ftp://ftp.pdbj.org/pub/emdb |
| UK | EBI PDBe | https://ftp.ebi.ac.uk/pub/databases/emdb | ftp://ftp.ebi.ac.uk/pub/databases/emdb |
| USA | wwPDB & RCSB PDB | https://ftp.wwpdb.org/pub/emdb | ftp://ftp.wwpdb.org/pub/emdb |
Related info.: Q: When the data are updated?
Q: When the data are updated?
External links: About the contents of the PDBj data site - PDBj
About the contents of the PDBj data site - PDBj
+Q: Is an EM map electron-density map?
A: No. But, they are very similar.
- To be exact, 3D map derived by 3D-EM is related to Coulomb potential (or electron potential), rather than electron density.
- The difference seems to be ignored in the most studies of atomic model building and fitting.
- Thers are some reports that protonation state affects the density. (see the ext. link)
Related info.: Q: What is the format for the maps? /
Q: What is the format for the maps? /  EMDB map data format
EMDB map data format
+Q: What is the format for the maps?
A: It is CCP4 map format
It's binary data with header for map geometry and body for density values.
Related info.: Q: Is an EM map electron-density map? /
Q: Is an EM map electron-density map? /  Q: Which software is suitable to view EMDB maps? /
Q: Which software is suitable to view EMDB maps? /  EMDB map data format
EMDB map data format
External links: EMDB Map Distribution Format Description (PDF document) /
EMDB Map Distribution Format Description (PDF document) /  CCP4 map format
CCP4 map format
+Q: Which software is suitable to view EMDB maps?
A: Many software packages can display CCP4 map data
- Some softwares introduced in this page shoud be suitable.
- Movies for EMDB map data in the EM Navigator are made by UCSF-Chimera.
- UCSF-Chimera seems to be the major software in the community.
Related info.: Q: What is the format for the maps? /
Q: What is the format for the maps? /  Q: How do you make the movies in EM Navigator? /
Q: How do you make the movies in EM Navigator? /  EMDB map data format
EMDB map data format
External links: UCSF Chimera Home Page
UCSF Chimera Home Page
+Q: What is EMD?
A: EMD (or emd) is "prefix" for ID number of EMDB
- At the beginning, the EMDB was also called EMD.
- Since the EM Navigator uses PDB and EMDB data, ID codes are indicated as [database name]-[ID], without the prefix. e.g. EMDB-1001, PDB-1brd.
Related info.: EMDB /
EMDB /  EM Navigator /
EM Navigator /  Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page) /
Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page) /  ID/Accession-code notation in Yorodumi/EM Navigator
ID/Accession-code notation in Yorodumi/EM Navigator
+Q: What are the data sources of EM Navigator?
A: EMDB and PDB electron microscopy dat
- Main source:
- all EMDB entries
- PDB entries with method information (exptl.details) including "electron microscopy" or "electron diffraction"
- Some contents and infromation are original of EM Navigator:
- Movies and their snapshot images
- Projection and slice images
- Some information about related data and similar structures
Related info.: EM Navigator /
EM Navigator /  EMDB /
EMDB /  PDB /
PDB /  EMN Search /
EMN Search /  EMN Papers /
EMN Papers /  EMN Statistics
EMN Statistics
+Q: Can I use the movies and their snapshots in the EM Navigator for papers or presentations?
A: Yes, plese
The EM Navigator movies and their snapshots are open to public. It would be appreciated if you would cite it as "EM Navigator, PDBj"
Related info.: Movie viewer
Movie viewer
External links: Terms and conditions - PDBj
Terms and conditions - PDBj
+Q: How do you make the movies in EM Navigator?
A: Series of images are recorded by UCSF-Chimera or Jmol. Then, they are encoded into movie files by FFmpeg
- This is an example of Chimera script.
system mkdir img1 movie reset movie record fformat png directory ./img1/ pattern img* roll y 2 180; wait wait 15 roll x 2 180; wait reset pos1; wait reset pos2 180; wait reset pos1 180; wait reset movie stop
"pos1" and "pos2" are saved position, which are the start and end of sectioning motion. - Chimera session files distributed with the movie files may be helpfull
Related info.: Q: Which software is suitable to view EMDB maps? /
Q: Which software is suitable to view EMDB maps? /  Q: How do you make the images for the structure data? What do their colors mean? /
Q: How do you make the images for the structure data? What do their colors mean? /  Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.? /
Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.? /  Movie viewer /
Movie viewer /  "Movies out of date"
"Movies out of date"
External links: UCSF Chimera Home Page /
UCSF Chimera Home Page /  Jmol: an open-source Java viewer for chemical structures in 3D /
Jmol: an open-source Java viewer for chemical structures in 3D /  FFmpeg
FFmpeg
+Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.?
A: in semi-automatic process with UCSF Chimera
- Method:
- We make been creating the images and movies for structure entries in EM Navigator using UCSF Chimera in semi-automatic process.
- Orientation:
- The structures are manually rotated to be seen in "good" orientation (usually, figures of the paper is refererd), or in similar orientation with similar structure data.
- For the case of icosahedral symmetry structure data, they are viewed normal to their five-fold axis, while typical view of such the structure data are along the two-fold or three-fold axes. This is to unify the orientation with structure data with pseudo-icosahedral symmetry, such as bacteriophage with tail structures.
- Color:
- In EM Navigator, multiple images are made for a single entry. In Omokage search, image of "colored surface view" are shown.
- The surface are colored by height, distance, or cylindrical radius. See the detail page for the coloring of particular images.
Related info.: Q: How do you make the images of PDB entries in Yorodumi, Omokage, etc.? /
Q: How do you make the images of PDB entries in Yorodumi, Omokage, etc.? /  Q: How do you make the movies in EM Navigator? /
Q: How do you make the movies in EM Navigator? /  Q: How do you make the images for the structure data? What do their colors mean?
Q: How do you make the images for the structure data? What do their colors mean?
External links: UCSF Chimera Home Page
UCSF Chimera Home Page
+Q: How do you make the images of PDB entries in Yorodumi, Omokage, etc.?
A: We are making them by full-automatic process using Jmol. Their styles are depend on the data type
- Orientation:
- Icosahedral assembly: Original orientation.
- Helical assembly: 6 orientation (+/- of X, Y, Z direction) images are generated in jpeg format. Then, the largest file is chosen. (large JPEG => many dots/colors)
- Others: by "rotate best" command of Jmol.
- Ribosomes: by "rotate best" command of Jmol. Only RNA coordinates are used for the orientation.
- Color:
- monomer AUs: by N->C (blue->red) rainbow
- BUs of monomer AUs: by "color molecule" (Jmol's coloring)
- multimer AUs and thier BUs (including monomer BUs): by "color chain". For this coloring, Jmol uses not a rainbow color but the colors determined by the chain-ID.
- Problems:
- BU models are generated by Jmol's system. It works well for the most data, but there are some exception.
- Checking and fixing processes of the many (>200,000) images are in progress. There are still some wrong images.
Related info.: Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.?
Q: How do you make the images of EMDB entries in Yorodumi, Omokage, etc.?
External links: Jmol: an open-source Java viewer for chemical structures in 3D
Jmol: an open-source Java viewer for chemical structures in 3D
+Q: Who make these documents? Who develop EM Navigator, Yorodumi, Omokage search, etc?
A: Hirofumi Suzuki (PDBj / IPR, Osaka University)
Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita, Osaka, 565-0871, Japan.
Related info.: PDBj /
PDBj /  Yorodumi annotation
Yorodumi annotation
External links: Contact to PDBj /
Contact to PDBj /  Protein Data Bank Japan at IPR, Osaka-univ.
Protein Data Bank Japan at IPR, Osaka-univ.
+Q: What is difference in PDB coordinate formats?
A: As follows
| Format | mmCIF compatible | Format based on | Provider | Comment |
|---|---|---|---|---|
| PDBx/mmCIF | - | CIF > STAR > plain text | wwPDB | Officail data format |
| PDB | no | plain text | wwPDB | Legacy officail |
| PDBML | yes | XML > plain text | wwPDB | For programers & develpers |
| PDBx/mmJSON | yes | JSON > plain text | PDBj | For programers & develpers |
Related info.: PDBx/mmCIF format /
PDBx/mmCIF format /  PDB format /
PDB format /  PDBx/mmJSON format
PDBx/mmJSON format
-Information (43)
+EMDB map data format
3D density map data in CCP4 format
Related info.: EMDB /
EMDB /  Q: What is the format for the maps? /
Q: What is the format for the maps? /  Q: Which software is suitable to view EMDB maps? /
Q: Which software is suitable to view EMDB maps? /  Q: Is an EM map electron-density map?
Q: Is an EM map electron-density map?
External links: EMDB Map Distribution Format Description /
EMDB Map Distribution Format Description /  Format technical details /
Format technical details /  CCP4 - Wikipedia
CCP4 - Wikipedia
+EMDB header
Meta information in XML format
Data file in XML format containing information associated with the map of EMDB entries, experimental information, detailed sample information, etc.
Related info.: EMDB /
EMDB /  Feb 9, 2022. New format data for meta-information of EMDB entries
Feb 9, 2022. New format data for meta-information of EMDB entries
External links: EMDB Schema v3 /
EMDB Schema v3 /  EMDB Schema v1.9 /
EMDB Schema v1.9 /  Documentation directory
Documentation directory
+Mask map
Map data of mask pattern, sub-region, segmentation, etc
Related info.: EMDB
EMDB
+FSC data file
Fourier shell correlation data in XML format for resolution estimation
Related info.: EMDB
EMDB
External links: Fourier shell correlation - Wikipedia /
Fourier shell correlation - Wikipedia /  FSC-schema
FSC-schema
+EMDB validaton report
Validaton report of EMDB entry by wwPDB and EMDB
Related info.: EMDB /
EMDB /  wwPDB validaton report
wwPDB validaton report
External links: User guide to the EmDataBank map validation reports
User guide to the EmDataBank map validation reports
+PDBx/mmCIF format
Officail data format for PDB entries, in CIF text format, having atomic coordinates and meta information
Related info.: PDB /
PDB /  Q: What is difference in PDB coordinate formats?
Q: What is difference in PDB coordinate formats?
External links: PDBx/mmCIF Dictionary Resources /
PDBx/mmCIF Dictionary Resources /  Crystallographic Information File
Crystallographic Information File
+PDB format
Legacy officail data format for PDB entries, contains atomic coordinates
Related info.: PDB /
PDB /  Q: What is difference in PDB coordinate formats?
Q: What is difference in PDB coordinate formats?
External links: Atomic Coordinate Entry Format Version 3.3
Atomic Coordinate Entry Format Version 3.3
+PDBx/mmJSON format
JSON representation of the PDBx/mmCIF data developed by PDBj
Related info.: PDB /
PDB /  Q: What is difference in PDB coordinate formats?
Q: What is difference in PDB coordinate formats?
External links: mmJSON - PDBj help
mmJSON - PDBj help
+wwPDB validaton report
Validaton report of PDB entry by wwPDB
Related info.: PDB /
PDB /  EMDB validaton report
EMDB validaton report
External links: wwPDB: Validation Reports /
wwPDB: Validation Reports /  wwPDB: validation report FAQs
wwPDB: validation report FAQs
+EM Navigator
3D electron microscopy data browser

URL: https://pdbj.org/emnavi/.
- EM Navigator is a browser for 3D electron microscopy (3D-EM) data of biological molecules and assemblies.
- It provides EMDB-PDB cross-search, statistical information, and links to similar structures.
- You can easily check the latest EM structural data and structural papers.
Related info.: EMDB /
EMDB /  PDB /
PDB /  PDBj /
PDBj /  Q: What are the data sources of EM Navigator? /
Q: What are the data sources of EM Navigator? /  EMN Search /
EMN Search /  EMN Papers /
EMN Papers /  EMN Gallery /
EMN Gallery /  EMN Statistics /
EMN Statistics /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+EMN Search
3DEM data search

URL: https://pdbj.org/emnavi/esearch.php
Advanced data search for EMDB and EM data in PDB widh various search and display options
Related info.: EMDB /
EMDB /  PDB /
PDB /  EM Navigator /
EM Navigator /  Q: What are the data sources of EM Navigator? /
Q: What are the data sources of EM Navigator? /  Yorodumi Search /
Yorodumi Search /  Jul 5, 2019. Downlodablable text data
Jul 5, 2019. Downlodablable text data
+EMN Papers
Database of articles cited by 3DEM data entries

URL: https://pdbj.org/emnavi/pap.php?em=1
- Database of articles cited by 3DEM data entries in EMDB and PDB
- Using PubMed data
Related info.: EMDB /
EMDB /  PDB /
PDB /  Q: What are the data sources of EM Navigator? /
Q: What are the data sources of EM Navigator? /  EM Navigator /
EM Navigator /  Yorodumi Papers /
Yorodumi Papers /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+EMN Gallery
Image gallery of 3DEM data

URL: https://pdbj.org/emnavi/gallery.php
Categorization is done by EM Navigator manager manually. It is not strict.
Related info.: EM Navigator
EM Navigator
+EMN Statistics
Statistics of 3DEM data in table and graph styles

URL: https://pdbj.org/emnavi/stat.php
- The table can be sorted. Click the column header to be sorted. (second click to reverse, [Shift]+click for multi-column sort)
- To show the bar graph in table mode, point the column/row header by mouse courser.
- To search the correspoinding data, click the cell of value.
- Examples:
Related info.: Q: What are the data sources of EM Navigator? /
Q: What are the data sources of EM Navigator? /  EM Navigator /
EM Navigator /  Mar 30, 2013. New page of EM Navigator, EMN Statistics
Mar 30, 2013. New page of EM Navigator, EMN Statistics
+Yorodumi
Thousand views of thousand structures
URL: https://pdbj.org/emnavi/quick.php
- Yorodumi is a browser for structure data from EMDB, PDB, SASBDB, etc.
- This page is also the successor to EM Navigator detail page, and also detail information page/front-end page for Omokage search.
- The word "yorodu" (or yorozu) is an old Japanese word meaning "ten thousand". "mi" (miru) is to see.
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  Yorodumi Search /
Yorodumi Search /  Aug 31, 2016. New EM Navigator & Yorodumi /
Aug 31, 2016. New EM Navigator & Yorodumi /  Yorodumi Papers /
Yorodumi Papers /  Jmol/JSmol /
Jmol/JSmol /  Function and homology information /
Function and homology information /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+Yorodumi Search
Cross-search of EMDB, PDB, SASBDB, etc.
URL: https://pdbj.org/emnavi/ysearch.php
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  EMN Search /
EMN Search /  Yorodumi /
Yorodumi /  Experimental methods, equipments, and software data /
Experimental methods, equipments, and software data /  Function and homology information /
Function and homology information /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+Yorodumi Papers
Database of articles cited by EMDB/PDB/SASBDB data
URL: https://pdbj.org/emnavi/pap.php
- Database of articles cited by EMDB, PDB, and SASBDB entries
- Using PubMed data
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Yorodumi /
Yorodumi /  EMN Papers /
EMN Papers /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+Yorodumi Speices
Taxonomy data in EMDB/PDB/SASBDB
URL: https://pdbj.org/emnavi/taxo.php
Taxonomy database of sample sources of data in EMDB/PDB/SASBDB
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  Mar 5, 2020. Novel coronavirus structure data
Mar 5, 2020. Novel coronavirus structure data
+Omokage search
Search structure by SHAPE

URL: https://pdbj.org/omokage/
- "Omokage search" is a shape similarity search service for 3D structures of macromolecules. By comparing global shapes, and ignoring details, similar-shaped structures are searched.
- The search is performed ageinst >200,000 structure data, which consists of EMDB map data, PDB coordinates (deposited units (asymmetric units, usually), PDB biological units, and SASBDB mdoels).
- For the search query, you can use either a data in the PDB/EMDB/SASBDB or your original model.
- Supported formats are PDB (atomic model, SAXS bead model, etc.) and CCP4/MRC map (3D density map).
- Shape comparison is performed by iDR profile method, which uses 1D-distance profiles of super-simplified models generated by vector quantizaion in Situs package.
Related info.: Comparison of 3 databanks /
Comparison of 3 databanks /  Sep 22, 2014. New service: Omokage search /
Sep 22, 2014. New service: Omokage search /  gmfit /
gmfit /  Re-ranking by gmfit
Re-ranking by gmfit
+Yorodumi Docs
+Covid-19 info
Page for Covid-19 featured contents

URL: https://pdbj.org/emnavi/covid19.php
Related info.: Comparison of 3 databanks /
Comparison of 3 databanks /  Aug 12, 2020. Covid-19 info
Aug 12, 2020. Covid-19 info
+F&H Search
Function & Homology similarity search
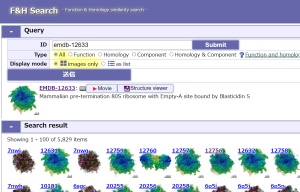
URL: https://pdbj.org/emnavi/fh-search.php
Based on the similarity of Function & Homolog items related to the structure data, similar data are searched from structure databases.
Related info.: Function and homology information
Function and homology information
+Movie viewer
Visualization of 3DEM structure data by movies
- Horizontal- and vertical- rotation motions and "slicing" motion are recorded in the movies.
- As the movie frame can be controlled by mouse/touch position, it can be operated interactively as a molecular viewer.
- According to the mouse/finger position and orientation of the motion on the movie panel, the model rotates on horizontal or vertical direction.
- By horizontal mouse/finger movement on the top part of the movie, the slicing motion is shown.
Related info.: Q: How do you make the movies in EM Navigator? /
Q: How do you make the movies in EM Navigator? /  Q: Can I use the movies and their snapshots in the EM Navigator for papers or presentations? /
Q: Can I use the movies and their snapshots in the EM Navigator for papers or presentations? /  SurfView /
SurfView /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+Jmol/JSmol
An open-source viewer for chemical structures in 3D
- Mouse/touch operation
X-Y rotation X-Y move Zoom in/out Mouse left drag [Ctrl] + right drag center drag (vertical) / wheel Touch single finger double finger pinch - Jmol menu: [Ctrl] + click / right click
Related info.: Molmil /
Molmil /  Yorodumi
Yorodumi
External links: Jmol: an open-source Java viewer for chemical structures in 3D
Jmol: an open-source Java viewer for chemical structures in 3D
+Molmil
WebGL based molecular viewer
- Molmil runs on Web browser on a smartphone or tablet device as well as PC. (Molmil is based on WebGL and Javascript).
- Mouse/touch operation
X-Y rotation X-Y move Zoom in/out Mouse left drag center drag / shift + left drag right drag / wheel Touch single finger double finger pinch - Function menu: top left buttons on viewer screen
Related info.: Jmol/JSmol /
Jmol/JSmol /  SurfView /
SurfView /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
External links: Documentation of Molmil in PDBj site /
Documentation of Molmil in PDBj site /  Molmil page in GitHub
Molmil page in GitHub
+SurfView
EMDB map surface models viewer on Web browsers

- SurfView run on Web browser on a smartphone or tablet device as well as PC.
- SurfView is based on a WebGL and three.js, Javascript 3D library.
- The surface models shwon in this viewer are modified for simplification and data size reduction. To get views of full-resolution data, see movies instead.
- Mouse/touch operation%table-1%
Related info.: Movie viewer /
Movie viewer /  Molmil /
Molmil /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
+Comparison of 3 databanks
Yorodumi and Omokage can cross-search these DBs
| Method | Main data | Str. data format | meta data format | |
|---|---|---|---|---|
| PDB | various | atomic model | PDBx/mmCIF, etc. | PDBx/mmCIF, etc. |
| EMDB | 3DEM | 3D map | CCP4 map | EMDB XML |
| SASBDB | Small Angle Scattering (SAS) | SAS profile (+/- 3D models) | PDB + sasCIF | ASCII + sasCIF |
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Covid-19 info /
Covid-19 info /  ID/Accession-code notation in Yorodumi/EM Navigator /
ID/Accession-code notation in Yorodumi/EM Navigator /  Experimental methods, equipments, and software data
Experimental methods, equipments, and software data
+ID/Accession-code notation in Yorodumi/EM Navigator
| Database | pattern | e.g. | note |
|---|---|---|---|
| EMDB | EMDB-**** | EMDB-1001 | prefix EMD- omitted |
| PDB | PDB-**** | PDB-1a00 | |
| SASBDB | _ | SASDA24 | as it is |
Related info.: Comparison of 3 databanks /
Comparison of 3 databanks /  Q: What is EMD? /
Q: What is EMD? /  Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
+EMDB
Electron Microscopy Data Bank, Databank for 3DEM

URL: https://www.ebi.ac.uk/emdb/
Related info.: 3D electron microscopy (3DEM) /
3D electron microscopy (3DEM) /  PDB /
PDB /  Comparison of 3 databanks
Comparison of 3 databanks
External links: The Electron Microscopy Data Bank - EBI /
The Electron Microscopy Data Bank - EBI /  EMDataResource
EMDataResource
+PDB
Protein Data Bank - The single repository of information about the 3D structures of proteins, nucleic acids, and complex assemblies

URL: https://wwPDB.org/
Related info.: PDBj /
PDBj /  EMDB
EMDB
+PDBj
Protein Data Bank Japan

URL: http://pdbj.org/
A member of wwPDB
Related info.: PDB /
PDB /  Information from PDBMLplus
Information from PDBMLplus
External links: Protein Data Bank Japan at IPR, Osaka-univ. /
Protein Data Bank Japan at IPR, Osaka-univ. /  PDBj@Facebook /
PDBj@Facebook /  PDBj@Twitter
PDBj@Twitter
+SASBDB
Small Angle Scattering Biological Data Bank - Curated repository for small angle scattering data and models

- Develop & manage: BioSAXS group in EMBL
- Since 2014
Related info.: Comparison of 3 databanks
Comparison of 3 databanks
+gmfit
A program for fitting subunits into density map of complex using GMM (Gaussian Mixture Model)

- Developed by Takeshi Kawabata in IPR, Osaka-univ.
- Used from Omokage search.
Related info.: Omokage search /
Omokage search /  Dec 4, 2015. The article about Omokage search is published online /
Dec 4, 2015. The article about Omokage search is published online /  Re-ranking by gmfit
Re-ranking by gmfit
External links: Pairwise gmfit
Pairwise gmfit
+3D electron microscopy (3DEM)
A generic term of electron microscopic analyses to obtain 3D structures
- Analyses such as electron tomography, single particle analysis, and electron diffraction are included.
Aggregation states Methods generally used "individual structure" electron tomography "single particle" single particle analysis "icosahedral" single particle analysis "helical" helical reconstruction / single particle analysis "2D/3D-crystal" electron diffraction / Fourier filtering - Characteristics compared to X-ray crystallography and NMR:
Advantage Wider applicability of sample (not require high-purity or high-concentration smaple, and useful for more huge, complex, flexible, and less uniform sample) Disadvantage (previous) Lower resolution (hard to get atomic-level resolution data) Disadvantage (new) High-performance electron microscopes and their maintenance costs are quite expensive
Related info.: EMDB /
EMDB /  electron cryo microscopy (cryoEM)
electron cryo microscopy (cryoEM)
+electron cryo microscopy (cryoEM)
Electron microscopy where the sample is studied at cryogenic temperatures
The specimen is cooled at 4~100K (-269~-170℃) to keep it in hydrated state in the highly vaqumed environment, and to reduce radiation dammage by electron beam.
Related info.: 3D electron microscopy (3DEM) /
3D electron microscopy (3DEM) /  Q: Is 3DEM same as electron cryo microscopy (cryoEM)?
Q: Is 3DEM same as electron cryo microscopy (cryoEM)?
+"Movies out of date"
Movies with this annotation may not be up-to-date.
EMDB map data are sometimes remediated. The process of making movies in the EM Navigator are not fully automated, and it is very hard to remake the all to catch up. So, some movies and movie parameters are based on the older data, and may be improper for new map data.
Related info.: Q: How do you make the movies in EM Navigator?
Q: How do you make the movies in EM Navigator?
+Carbohydrate representation
Polysaccaride/carbohydrate data representation in PDB
- In July 2020, representation for polysaccaride/carbohydrate information in PDB is improved.
- The polysaccaride figures are generated by GlycanBuilder2.
- See following external links for details.
External links: Implementation of GlycanBuilder to draw a wide variety of ambiguous glycans - ScienceDirect /
Implementation of GlycanBuilder to draw a wide variety of ambiguous glycans - ScienceDirect /  GlycanBuilder2 - RINGS (Resource For INformatics Of Glycomes at Soka) /
GlycanBuilder2 - RINGS (Resource For INformatics Of Glycomes at Soka) /  Carbohydrate Remediation - wwPDB documentation /
Carbohydrate Remediation - wwPDB documentation /  Coming July 29: Improved Carbohydrate Data at the PDB - wwPDB news
Coming July 29: Improved Carbohydrate Data at the PDB - wwPDB news
+Experimental methods, equipments, and software data
Database of experimental methods of EMDB, PDB and SABDB.
URL: https://pdbj.org/emnavi/ysearch.php?&act_tab=met
Related info.: Comparison of 3 databanks /
Comparison of 3 databanks /  Yorodumi Search
Yorodumi Search
+Function and homology information
Molecular function, domain, and homology information from related databases.
- To help to understand the molecular function and to find the related structure data, Yorodumi and Yorodumi Search display and utilize the related database information about function and homology.
- In addition to the information of EC, EMBL, GeneBank, GO, InterPro, UniProt, etc. stored in the EMDB header XML, PDBx/mmCIF and sasCIF original data, information of Pfam, PROSITE, Reactome, UniProkKB, etc. are collected via PDBMLplus, EMDB-PDB fitting data, and/or UniProt.
Category Name of database or definition Function% Gene ontology, Enzyme Commission number, Reactome, etc. Domain/homology CATH, InterPro, SMART, etc. Component UniProt, GenBank, PDB chemical component, etc.
Related info.: F&H Search /
F&H Search /  Information from PDBMLplus /
Information from PDBMLplus /  Yorodumi Search /
Yorodumi Search /  Yorodumi /
Yorodumi /  F&H Search
F&H Search
+Re-ranking by gmfit
The similarity ranking by Omokage search can be re-ordered according to correlation coefficient by gmfit, expecting incorporation of the (potential) benefits of two methods with following properties.
| Name | Method | Speed | Potential accuracy |
|---|---|---|---|
| Omokage search | iDR profile comparison | ++ sub-msec / 1 comparison | - 1D profile comparison |
| gmfit | Gaussian mixture model fitting | + sub-sec / 1 comparison | + comparison in 3D space |
Related info.: gmfit /
gmfit /  Omokage search
Omokage search
+Changes in new EM Navigator and Yorodumi
Changes in new versions of EM Navigator and Yorodumi. (Sep. 2016)
- Changes:
- New user interface unified in EM Navigator, Yorodumi & Omokage search, supporting mobile devices as well as PCs.
- New Yorodumi replace the individual data page ("Detail page" of EM Navigator in the legacy system). It is unified browser for EMDB, PDB, and SASBDB entries integrated with EM Navigator and Omokage search.
- The viewers (structure/movie viewers) appear in pop-up windows. On a PC, multiple viewer windows can be opened. On mobile devices, the viewers support touch operation.
- New features:
- Molmil, molecular structure viewer, and SurfView, surface model viewer for EMDB map data, are available to view the 3D structures. Both viewers support mobile device use.
- Yorodumi support SASBDB entries.
- Some new pages such as Yorodumi Papers, citation database of structure data entries and Yorodumi Search, cross search by keywords
Related info.: EM Navigator /
EM Navigator /  Yorodumi /
Yorodumi /  SurfView /
SurfView /  Molmil /
Molmil /  Movie viewer /
Movie viewer /  EMN Papers /
EMN Papers /  Yorodumi Papers /
Yorodumi Papers /  Yorodumi Search /
Yorodumi Search /  Aug 31, 2016. New EM Navigator & Yorodumi
Aug 31, 2016. New EM Navigator & Yorodumi
+Information from PDBMLplus
PDBMLplus is the XML format file including additional information relating to individual proteins. Currently, PDB files are lacking a detailed description of function, experimental conditions, and the like. And then such information was included to extended XML database, named PDBMLplus.
Related info.: PDBj /
PDBj /  Function and homology information
Function and homology information
External links: PDBMLplus - PDBj helip /
PDBMLplus - PDBj helip /  About Functional Details page - PDBJ help
About Functional Details page - PDBJ help
+Yorodumi annotation
Annotation by Yorodumi/EM Navigator manager
Related info.: Q: Who make these documents? Who develop EM Navigator, Yorodumi, Omokage search, etc?
Q: Who make these documents? Who develop EM Navigator, Yorodumi, Omokage search, etc?
 Movie
Movie Controller
Controller Structure viewers
Structure viewers