+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1gpq | ||||||
|---|---|---|---|---|---|---|---|
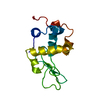
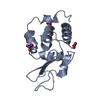
| Title | Structure of ivy complexed with its target, HEWL | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HYDROLASE/INHIBITOR / HYDROLASE-INHIBITOR COMPLEX / LYSOZYME-INHIBITOR COMPLEX / TYPE-C LYSOZYME INHIBITOR / HYDROLASE / GLYCOSIDASE / BACTERIAL TARGETS AT IGS-CNRS / FRANCE / BIGS / STRUCTURAL GENOMICS | ||||||
| Function / homology |  Function and homology information Function and homology informationlysozyme inhibitor activity / negative regulation of catalytic activity / chaperone-mediated protein folding / Lactose synthesis / Antimicrobial peptides / Neutrophil degranulation / beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process / lysozyme / lysozyme activity ...lysozyme inhibitor activity / negative regulation of catalytic activity / chaperone-mediated protein folding / Lactose synthesis / Antimicrobial peptides / Neutrophil degranulation / beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process / lysozyme / lysozyme activity / outer membrane-bounded periplasmic space / defense response to Gram-negative bacterium / killing of cells of another organism / periplasmic space / defense response to Gram-positive bacterium / defense response to bacterium / endoplasmic reticulum / protein homodimerization activity / extracellular space / identical protein binding / cytoplasm Similarity search - Function | ||||||
| Biological species |   | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å MOLECULAR REPLACEMENT / Resolution: 1.6 Å | ||||||
 Authors Authors | Abergel, C. / Monchois, V. / Claverie, J.-M. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2007 Journal: Proc.Natl.Acad.Sci.USA / Year: 2007Title: Structure and Evolution of the Ivy Protein Family, Unexpected Lysozyme Inhibitors in Gram-Negative Bacteria. Authors: Abergel, C. / Monchois, V. / Byrne, D. / Chenivesse, S. / Lembo, F. / Lazzaroni, J. / Claverie, J. #1: Journal: J.Biol.Chem. / Year: 2001 Title: Escherichia Coli Ykfe "Orfan" Gene Encodes a Potent Inhibitor of C-Type Lysozyme Authors: Monchois, V. / Abergel, C. / Sturgis, J. / Jeudy, S. / Claverie, J.-M. #2: Journal: Acta Crystallogr.,Sect.D / Year: 2000 Title: Crystallization and Preliminary Crystallographic Study of B0220, an "Orfan" Protein of Unknown Function from Escherichia Coli Authors: Abergel, C. / Monchois, V. / Chenivesse, S. / Jeudy, S. / Claverie, J.-M. | ||||||
| History |
| ||||||
| Remark 700 | SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN ... SHEET THE SHEET STRUCTURE OF THIS MOLECULE IS BIFURCATED. IN ORDER TO REPRESENT THIS FEATURE IN THE SHEET RECORDS BELOW, TWO SHEETS ARE DEFINED. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1gpq.cif.gz 1gpq.cif.gz | 124.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1gpq.ent.gz pdb1gpq.ent.gz | 98 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1gpq.json.gz 1gpq.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1gpq_validation.pdf.gz 1gpq_validation.pdf.gz | 440.9 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1gpq_full_validation.pdf.gz 1gpq_full_validation.pdf.gz | 449.2 KB | Display | |
| Data in XML |  1gpq_validation.xml.gz 1gpq_validation.xml.gz | 27.9 KB | Display | |
| Data in CIF |  1gpq_validation.cif.gz 1gpq_validation.cif.gz | 41.6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gp/1gpq https://data.pdbj.org/pub/pdb/validation_reports/gp/1gpq ftp://data.pdbj.org/pub/pdb/validation_reports/gp/1gpq ftp://data.pdbj.org/pub/pdb/validation_reports/gp/1gpq | HTTPS FTP |
-Related structure data
| Related structure data |  1uuzC  1xs0C  1helS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14973.754 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Details: COMPLEXED WITH HEWL IN CRYSTAL / Source: (gene. exp.)   #2: Protein | Mass: 14331.160 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: COMPLEXED WITH IVY IN CRYSTAL / Source: (natural)  #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.98 Å3/Da / Density % sol: 38 % Description: UNCOMPLEXED, AND UNREFINED MAD IVY STRUCTURE WAS ALSO USED FOR MOLECULAR REPLACEMENT |
|---|---|
| Crystal grow | pH: 9 Details: 15% PEG 4000, 5% GLYCEROL, CHES BUFFER 0.1M PH 9.0, PROTEIN CONCENTRATION: 5MG.ML-1 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: BM30A / Wavelength: 0.9798 / Beamline: BM30A / Wavelength: 0.9798 |
| Detector | Date: Aug 15, 2000 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9798 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→25 Å / Num. obs: 58982 / % possible obs: 99 % / Redundancy: 3.4 % / Biso Wilson estimate: 18.1 Å2 / Rmerge(I) obs: 0.044 / Net I/σ(I): 6.7 |
| Reflection shell | Resolution: 1.6→1.65 Å / Redundancy: 3.3 % / Rmerge(I) obs: 0.131 / Mean I/σ(I) obs: 4.1 / % possible all: 99 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1HEL Resolution: 1.6→20 Å / Rfactor Rfree error: 0.003 / Data cutoff high absF: 688920.93 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 48.18 Å2 / ksol: 0.38 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 19.4 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.6→1.7 Å / Rfactor Rfree error: 0.007 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller
























































































































































 PDBj
PDBj





