-Search query
-Search result
Showing all 50 items for (author: kjaer & s)

EMDB-29911: 
The Type 9 Secretion System Extended Translocon - delta GldL, Peak I, SprA-PorV-RemZ-PPI focused volume

EMDB-40085: 
The Type 9 Secretion System Extended Translocon - delta GldL, Peak I, SprE-SkpA-Nterm SprA focused volume

EMDB-40086: 
The Type 9 Secretion System Extended Translocon - delta GldL, Peak I, consensus volume

EMDB-40191: 
The Type 9 Secretion System in vitro assembled, RemA-CTD substrate bound complex

EMDB-40194: 
The Type 9 Secretion System Extended Translocon - SprA-PorV-PPI-RemZ-SkpA-SprE complex

EMDB-40195: 
The Type 9 Secretion System in vitro assembled, FspA-CTD substrate bound complex

EMDB-40196: 
The Type 9 Secretion System dGldL peak II, NucA substrate bound complex

EMDB-40199: 
The Type 9 Secretion System in vivo assembled, RemZ substrate bound complex - conformation 1

EMDB-40201: 
The Type 9 Secretion System in vivo assembled, RemZ substrate bound complex - conformation 2

EMDB-15217: 
PAPP-A dimer in complex with a dimer of the inhibitor STC2

EMDB-15219: 
PAPP-A dimer in complex with endogenous STC2 inhibitor.

EMDB-15220: 
Partial dimer complex of PAPP-A and its inhibitor STC2

EMDB-15221: 
PAPP-A dimer in complex with its inhibitor STC2

EMDB-13355: 
Structure of the Caulobacter crescentus S-layer protein RsaA N-terminal domain bound to LPS and soaked with Holmium

PDB-7peo: 
Structure of the Caulobacter crescentus S-layer protein RsaA N-terminal domain bound to LPS and soaked with Holmium

EMDB-12585: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (closed conformation)

EMDB-12586: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (one RBD erect)

EMDB-12587: 
Trimeric SARS-CoV-2 spike ectodomain bound to P008_056 Fab

EMDB-11777: 
RET/GDF15/GFRAL extracellular complex negative stain envelope

EMDB-11822: 
RET/GDNF/GFRa1 extracellular complex Cryo-EM structure

EMDB-10893: 
Structure of GldLM, the proton-powered motor that drives protein transport and gliding motility

EMDB-10959: 
Cryo-EM structure of the ARP2/3 1B5CL isoform complex.

EMDB-10960: 
Cryo-EM structure of the ARP2/3 1A5C isoform complex.

PDB-6yw6: 
Cryo-EM structure of the ARP2/3 1B5CL isoform complex.

PDB-6yw7: 
Cryo-EM structure of the ARP2/3 1A5C isoform complex.

EMDB-4605: 
Human Phenylalanine Hydroxylase (hPAH) apo structure

EMDB-0464: 
Legionella pneumophila bacterial flagellar motor

EMDB-0465: 
Pseudomonas aeruginosa bacterial flagellar motor
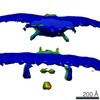
EMDB-0467: 
Shewanella oneidensis MR-1 bacterial flagellar motor

EMDB-3792: 
Electron cryotomogram of Vibrio cholerae O395 N1

EMDB-8603: 
Sub-tomogram average of the archaellar motor in Thermococcus kodakaraensis

EMDB-8492: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in wild-type Vibrio cholerae cells (aligning OM-parts)

EMDB-8493: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in wild-type Vibrio cholerae cells (aligning IM-parts)

EMDB-8494: 
Subtomogram average of the piliated toxin-coregulated pilus machine in wild-type Vibrio cholerae cells (aligning OM-parts)

EMDB-8495: 
Subtomogram average of the piliated toxin-coregulated pilus machine in wild-type Vibrio cholerae cells (aligning IM-parts)

EMDB-8496: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpS knockout (aligning OM-parts)

EMDB-8497: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpS knockout (aligning IM-parts)

EMDB-8498: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpB knockout (aligning OM-parts)

EMDB-8499: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpB knockout (aligning IM-parts)

EMDB-8500: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpD knockout (aligning OM-parts)

EMDB-8501: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpD knockout (aligning IM-parts)

EMDB-8502: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpR knockout (aligning OM-parts)

EMDB-8503: 
Subtomogram average of the non-piliated toxin-coregulated pilus machine in Vibrio cholerae cells with tcpR knockout (aligning IM-parts)

EMDB-3398: 
Electron Cryotomography and subvolume averaging of the cytoplasmic chemoreceptor array in Vibrio cholerae

EMDB-6102: 
Electron cryo-microscopy of DNGR-1 in complex with F-actin

PDB-3j82: 
Electron cryo-microscopy of DNGR-1 in complex with F-actin

EMDB-2712: 
Structure of the RET receptor tyrosine kinase extracellular domain

EMDB-2713: 
Structure of the zebrafish RET tyrosine kinase extracellular domain

PDB-4ux8: 
RET recognition of GDNF-GFRalpha1 ligand by a composite binding site promotes membrane-proximal self-association

EMDB-1099: 
Architecture of CRM1/Exportin1 suggests how cooperativity is achieved during formation of a nuclear export complex.
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
