+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0465 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
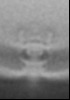
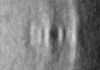
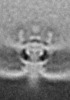
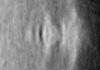
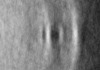
| Title | Pseudomonas aeruginosa bacterial flagellar motor | |||||||||
 Map data Map data | em-volume_P1 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 59.0 Å | |||||||||
 Authors Authors | Kaplan M / Ghosal D / Subramanian P / Oikonomou CO / Kjaer A / Pirbadian S / Ortega DR / Briegel A / El-Naggar MY / Jensen GJ | |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: The presence and absence of periplasmic rings in bacterial flagellar motors correlates with stator type. Authors: Mohammed Kaplan / Debnath Ghosal / Poorna Subramanian / Catherine M Oikonomou / Andreas Kjaer / Sahand Pirbadian / Davi R Ortega / Ariane Briegel / Mohamed Y El-Naggar / Grant J Jensen /  Abstract: The bacterial flagellar motor, a cell-envelope-embedded macromolecular machine that functions as a cellular propeller, exhibits significant structural variability between species. Different torque- ...The bacterial flagellar motor, a cell-envelope-embedded macromolecular machine that functions as a cellular propeller, exhibits significant structural variability between species. Different torque-generating stator modules allow motors to operate in different pH, salt or viscosity levels. How such diversity evolved is unknown. Here, we use electron cryo-tomography to determine the in situ macromolecular structures of three Gammaproteobacteria motors: , , and , providing the first views of intact motors with dual stator systems. Complementing our imaging with bioinformatics analysis, we find a correlation between the motor's stator system and its structural elaboration. Motors with a single H-driven stator have only the core periplasmic P- and L-rings; those with dual H-driven stators have an elaborated P-ring; and motors with Na or Na/H-driven stators have both their P- and L-rings embellished. Our results suggest an evolution of structural elaboration that may have enabled pathogenic bacteria to colonize higher-viscosity environments in animal hosts. #1:  Journal: bioRxiv Journal: bioRxivTitle: The structural complexity of the Gammaproteobacteria flagellar motor is related to the type of its torque-generating stators Authors: kaplan M / Ghosal D / Subramanian P / Oikonomou CO / Kjaer A / Pirbadian S / Ortega DR / El-Naggar MY / Jensen GJ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0465.map.gz emd_0465.map.gz | 1.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0465-v30.xml emd-0465-v30.xml emd-0465.xml emd-0465.xml | 8.6 KB 8.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0465.png emd_0465.png | 65.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0465 http://ftp.pdbj.org/pub/emdb/structures/EMD-0465 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0465 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0465 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0465.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0465.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | em-volume_P1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 13.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
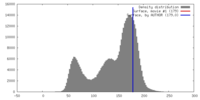
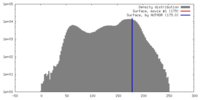
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Pseudomonas aeruginosa
| Entire | Name:  |
|---|---|
| Components |
|
-Supramolecule #1: Pseudomonas aeruginosa
| Supramolecule | Name: Pseudomonas aeruginosa / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 59.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number subtomograms used: 144 |
|---|---|
| Extraction | Number tomograms: 156 / Number images used: 144 |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller





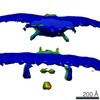



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)