-Search query
-Search result
Showing all 50 items for (author: jobe & a)

EMDB-50849: 
Cryo-EM structure of E. coli transcription factor NrdR in the ATP-bound, filamentous form
Method: single particle / : Martinez-Carranza M, Rozman Grinberg I, Sjoberg BM, Logan DT, Stenmark P

EMDB-50819: 
Cryo-EM structure of E. coli transcription factor NrdR in complex with DNA
Method: single particle / : Banerjee I, Bimai O, Martinez-Carranza M, Stenmark P, Sjoberg BM, Rozman Grinberg I, Logan DT

EMDB-50111: 
Cryo-EM structure of the I923V MDA5-dsRNA filament without nucleotide
Method: helical / : Singh R, Herrero del Valle A, Modis Y

EMDB-50136: 
Cryo-EM structure of the I923V MDA5-dsRNA filament with ADP-AlF4 bound and 81-degree helical twist
Method: helical / : Singh R, Herrero del Valle A, Modis Y

EMDB-50137: 
Cryo-EM structure of the I923V MDA5-dsRNA filament with ADP-AlF4 bound and 88-degree helical twist
Method: helical / : Singh R, Herrero del Valle A, Modis Y

EMDB-50150: 
Cryo-EM structure of the I923V MDA5-dsRNA filament with ADP-AlF4 bound and 73-degree helical twist
Method: helical / : Singh R, Herrero del Valle A, Modis Y

EMDB-50165: 
Cryo-EM structure of the I923V MDA5-dsRNA filament in complex with ATP
Method: helical / : Singh R, Herrero del Valle A, Modis Y

EMDB-50175: 
Cryo-EM structure of the A946T MDA5-dsRNA filament
Method: helical / : Singh R, Herrero del Valle A, Modis Y

PDB-9f0j: 
Cryo-EM structure of the I923V MDA5-dsRNA filament without nucleotide
Method: helical / : Singh R, Herrero del Valle A, Modis Y

PDB-9f1u: 
Cryo-EM structure of the I923V MDA5-dsRNA filament with ADP-AlF4 bound and 81-degree helical twist
Method: helical / : Singh R, Herrero del Valle A, Modis Y

PDB-9f20: 
Cryo-EM structure of the I923V MDA5-dsRNA filament with ADP-AlF4 bound and 88-degree helical twist
Method: helical / : Singh R, Herrero del Valle A, Modis Y

PDB-9f2l: 
Cryo-EM structure of the I923V MDA5-dsRNA filament with ADP-AlF4 bound and 73-degree helical twist
Method: helical / : Singh R, Herrero del Valle A, Modis Y

PDB-9f2w: 
Cryo-EM structure of the I923V MDA5-dsRNA filament in complex with ATP
Method: helical / : Singh R, Herrero del Valle A, Modis Y

PDB-9f3p: 
Cryo-EM structure of the A946T MDA5-dsRNA filament
Method: helical / : Singh R, Herrero del Valle A, Modis Y

EMDB-17360: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its tetrameric state produced in the presence of dATP and CTP
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

EMDB-17361: 
Cryo-EM structure of the dimeric form of the anaerobic ribonucleotide reductase from Prevotella copri produced in the presence of dATP and CTP
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

EMDB-17373: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, ATP/dTTP/GTP-bound state
Method: single particle / : Bimai O, Banerjee I, Sjoberg BM, Logan DT

EMDB-17385: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, dGTP/ATP-bound state
Method: single particle / : Bimai O, Banerjee I, Sjoberg BM, Logan DT

PDB-8p2c: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its tetrameric state produced in the presence of dATP and CTP
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

PDB-8p2d: 
Cryo-EM structure of the dimeric form of the anaerobic ribonucleotide reductase from Prevotella copri produced in the presence of dATP and CTP
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

PDB-8p2s: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, ATP/dTTP/GTP-bound state
Method: single particle / : Bimai O, Banerjee I, Sjoberg BM, Logan DT

PDB-8p39: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, dGTP/ATP-bound state
Method: single particle / : Bimai O, Banerjee I, Sjoberg BM, Logan DT

EMDB-17357: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, ATP/CTP-bound state
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

EMDB-17358: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, dATP-bound state
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

EMDB-17359: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its tetrameric, dATP-bound state
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

PDB-8p23: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, ATP/CTP-bound state
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

PDB-8p27: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, dATP-bound state
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

PDB-8p28: 
Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its tetrameric, dATP-bound state
Method: single particle / : Banerjee I, Bimai O, Sjoberg BM, Logan DT

EMDB-13178: 
Streptomyces coelicolor ATP-loaded NrdR
Method: single particle / : Martinez-Carranza M, Stenmark P

EMDB-13179: 
Streptomyces coelicolor dATP/ATP-loaded NrdR in complex with its cognate DNA
Method: single particle / : Martinez-Carranza M, Stenmark P

EMDB-13182: 
Streptomyces coelicolor dATP/ATP-loaded NrdR octamer
Method: single particle / : Martinez-Carranza M, Stenmark P

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
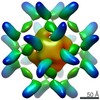
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-12985: 
Low resolution reconstruction of the dATP-inhibited complex of the ribonucleotide reductase NrdA and NrdB proteins from Leeuwenhoekiella blandensis
Method: single particle / : Banerjee I, Rozman Grinberg I, Sjoberg BM, Logan DT

PDB-4v8m: 
High-resolution cryo-electron microscopy structure of the Trypanosoma brucei ribosome
Method: single particle / : Hashem Y, des Georges A, Fu J, Buss SN, Jossinet F, Jobe A, Zhang Q, Liao HY, Grassucci RA, Bajaj C, Westhof E, Madison-Antenucci S, Frank J

EMDB-5936: 
Electron cryo-microscopy of the Moloney murine leukemia virus furin precursor Env in its native form
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-5937: 
Electron cryo-microscopy of the Moloney murine leukemia virus furin precursor Env in its isomerization arrested state (IAS) form
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-5938: 
Electron cryo-microscopy of the Moloney murine leukemia virus furin precursor Env in its native form in complex with 83A25 Fab
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-5939: 
Electron cryo-microscopy of the partially glycosylated gp80 form of the wild type furin precursor Env of Moloney murine leukemia virus
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-2239: 
Cryo-electron microscopy structure of the Trypanosoma brucei 80S ribosome
Method: single particle / : Hashem Y, des Georges A, Fu J, Buss SN, Jossinet F, Jobe A, Zhang Q, Liao HY, Grassucci B, Bajaj C, Westhof E, Madison-Antenucci S, Frank J

EMDB-2065: 
Electron cryo-microscopy of R-peptide precursor of Moloney murine leukemia virus Env in its isomerization arrested intermediate state
Method: single particle / : Loving R, Wu SR, Sjoberg M, Lindqvist B, Garoff H

EMDB-2066: 
Electron cryo-microscopy of the L651A mutant R-peptide precursor Env of Moloney murine leukemia virus in its native state
Method: single particle / : Loving R, Wu SR, Sjoberg M, Lindqvist B, Garoff H

EMDB-2067: 
Electron cryo-microscopy of R-peptide precursor of Moloney murine leukemia virus Env in its native form
Method: single particle / : Loving R, Wu SR, Sjoberg M, Lindqvist B, Garoff H

EMDB-1800: 
Single particle cryo-electron microscopy analysis of CD4 bound HIV-1 Env
Method: single particle / : Wu SR, Loving R, Lindqvist B, Hebert H, Koeck P, Sjoberg M, Garoff H

EMDB-1801: 
Single particle cryo-electron microscopy analysis of unliganded HIV-1 Env
Method: single particle / : Wu SR, Loving R, Lindqvist B, Hebert H, Koeck P, Sjoberg M, Garoff H

EMDB-2863: 
Surface rendered vitrified native Env
Method: single particle / : Wu SR, Sjoberg M, Wallin M, Lindqvist B, Ekstrom M, Hebert H, Koeck P, Garoff H
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
