8RP5
 
 | | Alpha-Methylacyl-CoA racemase from Mycobacterium tuberculosis (E241A mutant) | | Descriptor: | 1,2-ETHANEDIOL, Alpha-methylacyl-CoA racemase, TRIETHYLENE GLYCOL | | Authors: | Mojanaga, O.O, Acharya, K.R, Lloyd, M.D. | | Deposit date: | 2024-01-12 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | alpha-Methylacyl-CoA Racemase from Mycobacterium tuberculosis -Detailed Kinetic and Structural Characterization of the Active Site.
Biomolecules, 14, 2024
|
|
7T72
 
 | | Epitope-based selection of SARS-CoV-2 neutralizing antibodies from convalescent patients | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Antibody heavy chain, Antibody light chain, ... | | Authors: | Langley, D.B, Christ, D, Rouet, R. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.177 Å) | | Cite: | Broadly neutralizing SARS-CoV-2 antibodies through epitope-based selection from convalescent patients.
Nat Commun, 14, 2023
|
|
2NZ4
 
 | |
7T90
 
 | | Cryo-EM structure of ACh-bound M2R-Go signaling complex in S2 state | | Descriptor: | ACETYLCHOLINE, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Xu, J, Wang, Q, Du, Y, Kobilka, B.K. | | Deposit date: | 2021-12-17 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (3.32 Å) | | Cite: | Structural and dynamic insights into supra-physiological activation and allosteric modulation of a muscarinic acetylcholine receptor.
Nat Commun, 14, 2023
|
|
7T8X
 
 | | Cryo-EM structure of ACh-bound M2R-Go signaling complex in S1 state | | Descriptor: | ACETYLCHOLINE, Antibody fragment, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Xu, J, Wang, Q, Du, Y, Kobilka, B.K. | | Deposit date: | 2021-12-17 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (3.21 Å) | | Cite: | Structural and dynamic insights into supra-physiological activation and allosteric modulation of a muscarinic acetylcholine receptor.
Nat Commun, 14, 2023
|
|
7T9R
 
 | | Crystal structure of Crocodile defensin CpoBD13:phosphatidic acid complex | | Descriptor: | (R)-2-(FORMYLOXY)-3-(PHOSPHONOOXY)PROPYL PENTANOATE, 1,2-ETHANEDIOL, CpoBD13, ... | | Authors: | Kvansakul, M, Williams, S, Hulett, M.D. | | Deposit date: | 2021-12-19 | | Release date: | 2023-01-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crocodile defensin (CpoBD13) antifungal activity via pH-dependent phospholipid targeting and membrane disruption.
Nat Commun, 14, 2023
|
|
7T5O
 
 | | VFLIP Spike Trimer with GAR03 | | Descriptor: | GAR03 Fab heavy chain, GAR03 Fab light chain, Spike glycoprotein | | Authors: | Sobti, M, Stewart, A.G, Rouet, R, Langley, D.B. | | Deposit date: | 2021-12-12 | | Release date: | 2023-01-25 | | Last modified: | 2023-08-09 | | Method: | ELECTRON MICROSCOPY (3.39 Å) | | Cite: | Broadly neutralizing SARS-CoV-2 antibodies through epitope-based selection from convalescent patients.
Nat Commun, 14, 2023
|
|
7T9Q
 
 | | Crystal structure of Crocodile defensin CpoBD13 | | Descriptor: | 1,2-ETHANEDIOL, CITRATE ANION, CpoBD13 | | Authors: | Kvansakul, M, Williams, S, Hulett, M.D. | | Deposit date: | 2021-12-19 | | Release date: | 2023-01-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crocodile defensin (CpoBD13) antifungal activity via pH-dependent phospholipid targeting and membrane disruption.
Nat Commun, 14, 2023
|
|
7T8F
 
 | |
7T96
 
 | | Cryo-EM structure of S2 state ACh-bound M2R-Go signaling complex with a PAM | | Descriptor: | 3-amino-5-chloro-N-cyclopropyl-4-methyl-6-[2-(4-methylpiperazin-1-yl)-2-oxoethoxy]thieno[2,3-b]pyridine-2-carboxamide, ACETYLCHOLINE, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Xu, J, Wang, Q, Du, Y, Kobilka, B.K. | | Deposit date: | 2021-12-18 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (3.22 Å) | | Cite: | Structural and dynamic insights into supra-physiological activation and allosteric modulation of a muscarinic acetylcholine receptor.
Nat Commun, 14, 2023
|
|
7T8E
 
 | | G93A mutant of human SOD1 in P21 space group | | Descriptor: | ACETATE ION, SULFATE ION, Superoxide dismutase [Cu-Zn], ... | | Authors: | Amporndanai, K, Hasnain, S.S. | | Deposit date: | 2021-12-16 | | Release date: | 2023-01-25 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Ebselen analogues delay disease onset and its course in fALS by on-target SOD-1 engagement.
Sci Rep, 14, 2024
|
|
7T94
 
 | | Cryo-EM structure of S1 state ACh-bound M2R-Go signaling complex with a PAM | | Descriptor: | 3-amino-5-chloro-N-cyclopropyl-4-methyl-6-[2-(4-methylpiperazin-1-yl)-2-oxoethoxy]thieno[2,3-b]pyridine-2-carboxamide, ACETYLCHOLINE, Antibody fragment, ... | | Authors: | Xu, J, Wang, Q, Du, Y, Kobilka, B.K. | | Deposit date: | 2021-12-17 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | Structural and dynamic insights into supra-physiological activation and allosteric modulation of a muscarinic acetylcholine receptor.
Nat Commun, 14, 2023
|
|
7T8G
 
 | |
7T8H
 
 | |
2OO4
 
 | | Structure of LNR-HD (Negative Regulatory Region) from human Notch 2 | | Descriptor: | CALCIUM ION, GLYCEROL, Neurogenic locus notch homolog protein 2, ... | | Authors: | Gordon, W.R, Vardar-Ulu, D, Histen, G, Sanchez-Irizarry, C, Aster, J.C, Blacklow, S.C. | | Deposit date: | 2007-01-25 | | Release date: | 2007-04-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.003 Å) | | Cite: | Structural basis for autoinhibition of Notch
Nat.Struct.Mol.Biol., 14, 2007
|
|
7STF
 
 | | Structure of KRAS G12V/HLA-A*03:01 in complex with antibody fragment V2 | | Descriptor: | Beta-2-microglobulin, HLA class I histocompatibility antigen, A alpha chain, ... | | Authors: | Wright, K.M, Gabelli, S.B, Miller, M. | | Deposit date: | 2021-11-12 | | Release date: | 2023-05-31 | | Last modified: | 2023-10-25 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | Hydrophobic interactions dominate the recognition of a KRAS G12V neoantigen.
Nat Commun, 14, 2023
|
|
7SXJ
 
 | | BIO-2895 (BRD0705) bound GSK3beta-axin complex | | Descriptor: | (4~{S})-4-ethyl-7,7-dimethyl-4-phenyl-2,6,8,9-tetrahydropyrazolo[3,4-b]quinolin-5-one, Glycogen synthase kinase-3 beta, axin peptide | | Authors: | Chodaparambil, J.V. | | Deposit date: | 2021-11-23 | | Release date: | 2023-06-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Elucidation of the GSK3 alpha Structure Informs the Design of Novel, Paralog-Selective Inhibitors.
Acs Chem Neurosci, 14, 2023
|
|
7SXG
 
 | | BIO-8546 bound GSK3alpha-axin complex | | Descriptor: | (4S,5R,8R)-4-ethyl-8-fluoro-4-[3-(3-fluoro-5-methoxypyridin-4-yl)phenyl]-7,7-dimethyl-4,5,6,7,8,9-hexahydro-2H-pyrazolo[3,4-b]quinolin-5-ol, Axin peptide, Glycogen synthase kinase-3 alpha | | Authors: | Chodaparambil, J.V. | | Deposit date: | 2021-11-23 | | Release date: | 2023-06-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Elucidation of the GSK3 alpha Structure Informs the Design of Novel, Paralog-Selective Inhibitors.
Acs Chem Neurosci, 14, 2023
|
|
7SXF
 
 | | BIO-2895 (BRD0705) bound GSK3alpha-axin complex | | Descriptor: | (4~{S})-4-ethyl-7,7-dimethyl-4-phenyl-2,6,8,9-tetrahydropyrazolo[3,4-b]quinolin-5-one, Axin peptide, CALCIUM ION, ... | | Authors: | Chodaparambil, J.V. | | Deposit date: | 2021-11-23 | | Release date: | 2023-06-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Elucidation of the GSK3 alpha Structure Informs the Design of Novel, Paralog-Selective Inhibitors.
Acs Chem Neurosci, 14, 2023
|
|
7SXH
 
 | | BIO-8546 bound GSK3beta-axin complex | | Descriptor: | (4S,5R,8R)-4-ethyl-8-fluoro-4-[3-(3-fluoro-5-methoxypyridin-4-yl)phenyl]-7,7-dimethyl-4,5,6,7,8,9-hexahydro-2H-pyrazolo[3,4-b]quinolin-5-ol, Glycogen synthase kinase-3 beta, axin peptide | | Authors: | Chodaparambil, J.V. | | Deposit date: | 2021-11-23 | | Release date: | 2023-06-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Elucidation of the GSK3 alpha Structure Informs the Design of Novel, Paralog-Selective Inhibitors.
Acs Chem Neurosci, 14, 2023
|
|
7SG4
 
 | |
2OT8
 
 | |
8Q5B
 
 | |
7R7P
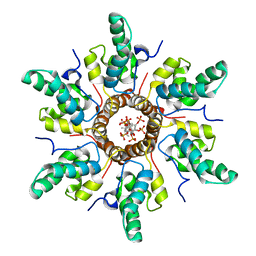 
 | | Immature HIV-1 CACTD-SP1 lattice with Bevirimat (BVM) and Inositol hexakisphosphate (IP6) | | Descriptor: | 3alpha-[(3-carboxy-3-methylbutanoyl)oxy]-8alpha,9beta,10alpha,13alpha,17alpha,19beta-lup-20(29)-en-28-oic acid, Gag polyprotein, INOSITOL HEXAKISPHOSPHATE | | Authors: | Sarkar, S, Zadrozny, K.K, Zadorozhnyi, R, Russell, R.W, Quinn, C.M, Kleinpeter, A, Ablan, S, Meshkin, H, Perilla, J.R, Ganser-Pornillos, B.K, Pornillos, O, Freed, E.O, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2021-06-25 | | Release date: | 2023-02-15 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | Structural basis of HIV-1 maturation inhibitor binding and activity.
Nat Commun, 14, 2023
|
|
7R7Q
 
 | | Immature HIV-1 CACTD-SP1 lattice with Inositol hexakisphosphate (IP6) | | Descriptor: | Gag polyprotein, INOSITOL HEXAKISPHOSPHATE | | Authors: | Sarkar, S, Zadrozny, K.K, Zadorozhnyi, R, Russell, R.W, Quinn, C.M, Kleinpeter, A, Ablan, S, Meshkin, H, Perilla, J.R, Ganser-Pornillos, B.K, Pornillos, O, Freed, E.O, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2021-06-25 | | Release date: | 2023-02-15 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | Structural basis of HIV-1 maturation inhibitor binding and activity.
Nat Commun, 14, 2023
|
|
