6GEK
 
 | |
6GE3
 
 | |
6GGN
 
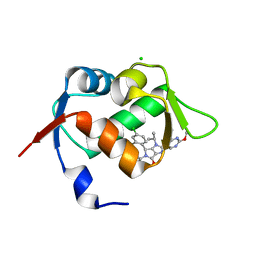 | | In vitro and in vivo characterization of a novel, highly potent p53-MDM2 inhibitor | | Descriptor: | (4~{S})-4-(4-chloranyl-2-methyl-phenyl)-5-(5-chloranyl-2-methyl-phenyl)-2-(2,4-dimethoxypyrimidin-5-yl)-3-propan-2-yl-4~{H}-pyrrolo[3,4-d]imidazol-6-one, CHLORIDE ION, E3 ubiquitin-protein ligase Mdm2 | | Authors: | Kallen, J. | | Deposit date: | 2018-05-03 | | Release date: | 2018-09-26 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | In vitro and in vivo characterization of a novel, highly potent p53-MDM2 inhibitor.
Bioorg. Med. Chem. Lett., 28, 2018
|
|
6GE6
 
 | |
6GEG
 
 | |
6GEE
 
 | |
3HI9
 
 | |
8C17
 
 | | Crystal structure of TEAD4 in complex with peptide 1 | | Descriptor: | GLYCEROL, MYRISTIC ACID, Stapled peptide, ... | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2022-12-20 | | Release date: | 2023-03-15 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Biochemical and Structural Characterization of a Peptidic Inhibitor of the YAP:TEAD Interaction That Binds to the alpha-Helix Pocket on TEAD.
Acs Chem.Biol., 18, 2023
|
|
8CAA
 
 | | Crystal structure of TEAD4 in complex with YTP-13 | | Descriptor: | (2~{R})-2-[2-chloranyl-5-[2-chloranyl-4-(trifluoromethyl)phenoxy]phenyl]sulfanylpropanoic acid, 4-[bis(fluoranyl)methoxy]-2-[(2~{S})-5-chloranyl-6-fluoranyl-2-[[(4-oxidanylcyclohexyl)amino]methyl]-2-phenyl-3~{H}-1-benzofuran-4-yl]-3-fluoranyl-benzamide, PHOSPHATE ION, ... | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2023-01-24 | | Release date: | 2023-04-12 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.999 Å) | | Cite: | Optimization of a Class of Dihydrobenzofurane Analogs toward Orally Efficacious YAP-TEAD Protein-Protein Interaction Inhibitors.
Chemmedchem, 18, 2023
|
|
1CWB
 
 | |
1CWA
 
 | |
8A0V
 
 | | Crystal structure of TEAD3 in complex with CPD2 | | Descriptor: | DIMETHYL SULFOXIDE, MYRISTIC ACID, PHOSPHATE ION, ... | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2022-05-30 | | Release date: | 2022-08-17 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.699 Å) | | Cite: | The First Class of Small Molecules Potently Disrupting the YAP-TEAD Interaction by Direct Competition.
Chemmedchem, 17, 2022
|
|
8A0U
 
 | | Crystal structure of TEAD3 in complex with CPD4 | | Descriptor: | 2-[(2~{S})-2-(aminomethyl)-5-chloranyl-2-phenyl-3~{H}-1-benzofuran-4-yl]benzamide, MYRISTIC ACID, PHOSPHATE ION, ... | | Authors: | Scheufler, C, Kallen, J. | | Deposit date: | 2022-05-30 | | Release date: | 2022-08-17 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.895 Å) | | Cite: | The First Class of Small Molecules Potently Disrupting the YAP-TEAD Interaction by Direct Competition.
Chemmedchem, 17, 2022
|
|
1UOM
 
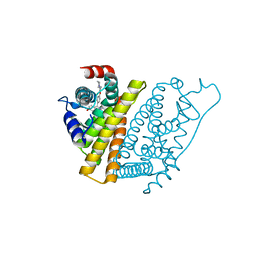 | | The Structure of Estrogen Receptor in Complex with a Selective and Potent Tetrahydroisochiolin Ligand. | | Descriptor: | 2-PHENYL-1-[4-(2-PIPERIDIN-1-YL-ETHOXY)-PHENYL]-1,2,3,4-TETRAHYDRO-ISOQUINOLIN-6-OL, ESTROGEN RECEPTOR | | Authors: | Stark, W, Bischoff, S.F, Buhl, T, Fournier, B, Halleux, C, Kallen, J, Keller, H, Renaud, J. | | Deposit date: | 2003-04-11 | | Release date: | 2003-07-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Estrogen Receptor Modulators: Identification and Structure-Activity Relationships of Potent Eralpha-Selective Tetrahydroisoquinoline Ligands
J.Med.Chem., 46, 2003
|
|
1XDG
 
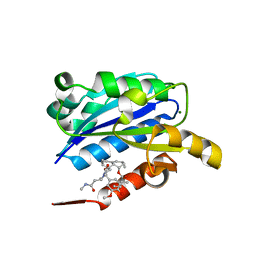 | | X-ray structure of LFA-1 I-domain in complex with LFA878 at 2.1A resolution | | Descriptor: | (1S,3R,8AS)-8-(2-{(4S,6S)-3-(4-HYDROXY-3-METHOXYBENZYL)-4-[2-(METHYLAMINO)-2-OXOETHYL]-2-OXO-1,3-OXAZINAN-6-YL}ETHYL)-3 ,7-DIMETHYL-1,2,3,7,8,8A-HEXAHYDRONAPHTHALEN-1-YL (2R)-2-METHYLBUTANOATE, Integrin alpha-L, MAGNESIUM ION | | Authors: | Weitz-Schmidt, G, Welzenbach, K, Dawson, J, Kallen, J. | | Deposit date: | 2004-09-06 | | Release date: | 2004-09-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Improved lymphocyte function-associated antigen-1 (LFA-1) inhibition by statin derivatives: molecular basis determined by x-ray analysis and monitoring of LFA-1 conformational changes in vitro and ex vivo
J.Biol.Chem., 279, 2004
|
|
1XDD
 
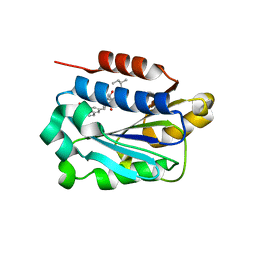 | | X-ray structure of LFA-1 I-domain in complex with LFA703 at 2.2A resolution | | Descriptor: | 8-[2-((2S)-4-HYDROXY-1-{[5-(HYDROXYMETHYL)-6-METHOXY-2-NAPHTHYL]METHYL}-6-OXOPIPERIDIN-2-YL)ETHYL]-3,7-DIMETHYL-1,2,3,7,8,8A-HEXAHYDRONAPHTHALEN-1-YL 2-METHYLBUTANOATE, Integrin alpha-L, MAGNESIUM ION | | Authors: | Weitz-Schmidt, G, Welzenbach, K, Dawson, J, Kallen, J. | | Deposit date: | 2004-09-06 | | Release date: | 2004-09-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Improved lymphocyte function-associated antigen-1 (LFA-1) inhibition by statin derivatives: molecular basis determined by x-ray analysis and monitoring of LFA-1 conformational changes in vitro and ex vivo
J.Biol.Chem., 279, 2004
|
|
1XQC
 
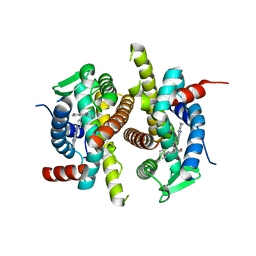 | | X-ray structure of ERalpha LBD bound to a tetrahydroisoquinoline SERM ligand at 2.05A resolution | | Descriptor: | (1S)-1-{4-[(9AR)-OCTAHYDRO-2H-PYRIDO[1,2-A]PYRAZIN-2-YL]PHENYL}-2-PHENYL-1,2,3,4-TETRAHYDROISOQUINOLIN-6-OL, Estrogen receptor | | Authors: | Renaud, J, Bischoff, S.F, Buhl, T, Floersheim, P, Fournier, B, Geiser, M, Halleux, C, Kallen, J, Keller, H.J, Ramage, P. | | Deposit date: | 2004-10-12 | | Release date: | 2005-02-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Selective Estrogen Receptor Modulators with Conformationally Restricted Side Chains. Synthesis and Structure-Activity Relationship of ERalpha-Selective Tetrahydroisoquinoline Ligands
J.Med.Chem., 48, 2005
|
|
1CYN
 
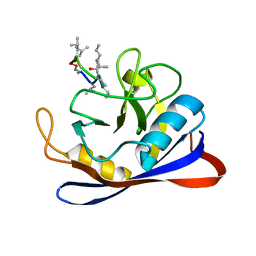 | |
1XUO
 
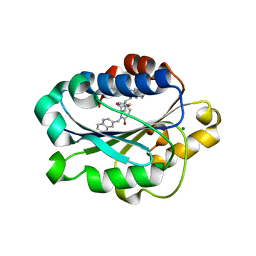 | | X-ray structure of LFA-1 I-domain bound to a 1,4-diazepane-2,5-dione inhibitor at 1.8A resolution | | Descriptor: | (2R)-2-[3-ISOBUTYL-2,5-DIOXO-4-(QUINOLIN-3-YLMETHYL)-1,4-DIAZEPAN-1-YL]-N-METHYL-3-(2-NAPHTHYL)PROPANAMIDE, Integrin alpha-L, MAGNESIUM ION | | Authors: | Wattanasin, S, Kallen, J, Myers, S, Guo, Q, Sabio, M, Ehrhardt, C, Albert, R, Hommel, U, Weckbecker, G, Welzenbach, K. | | Deposit date: | 2004-10-26 | | Release date: | 2005-10-26 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | 1,4-Diazepane-2,5-diones as novel inhibitors of LFA-1
Bioorg.Med.Chem.Lett., 15, 2005
|
|
1CWO
 
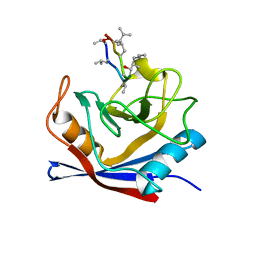 | | HUMAN CYCLOPHILIN A COMPLEXED WITH THR2, LEU5, D-HIV8, LEU10 CYCLOSPORIN | | Descriptor: | CYCLOSPORIN C, PEPTIDYL-PROLYL CIS-TRANS ISOMERASE A | | Authors: | Mikol, V, Kallen, J, Taylor, P, Walkinshaw, M.D. | | Deposit date: | 1998-06-05 | | Release date: | 1998-08-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Conformational Differences of an Immunosuppressant Peptolide in a Single Crystal and in a Crystal Complex with Human Cyclophilin A.
J.Mol.Biol., 283, 1998
|
|
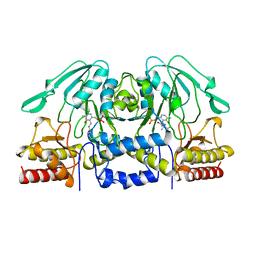
1BJO
 
 | |
1BJN
 
 | |
1CWC
 | |
2SOC
 
 | |
1MIL
 
 | | TRANSFORMING PROTEIN | | Descriptor: | SHC ADAPTOR PROTEIN | | Authors: | Mikol, V. | | Deposit date: | 1995-09-20 | | Release date: | 1996-11-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the SH2 domain from the adaptor protein SHC: a model for peptide binding based on X-ray and NMR data.
J.Mol.Biol., 254, 1995
|
|
