10AD
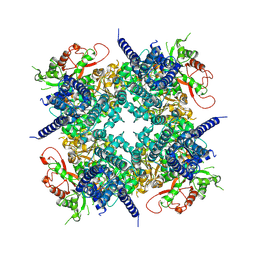 
 | | Cryo-EM structure of the human BK channel bound to the agonist NS1619 | | Descriptor: | 3-[2-oxidanyl-5-(trifluoromethyl)phenyl]-6-(trifluoromethyl)-1~{H}-benzimidazol-2-one, CALCIUM ION, Isoform 5 of Calcium-activated potassium channel subunit alpha-1, ... | | Authors: | Gonzalez-Sanabria, N, Contreras, G.F, Perozo, E, Latorre, R. | | Deposit date: | 2026-01-08 | | Release date: | 2026-02-04 | | Last modified: | 2026-02-11 | | Method: | ELECTRON MICROSCOPY (3.44 Å) | | Cite: | The BK channel-NS1619 agonist complex reveals molecular insights into allosteric activation gating.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
10AJ
 
 | | Crystal Structure of Human WRN helicase with compound 1 | | Descriptor: | 1,2-ETHANEDIOL, 4-(4-methyl-4H-1,2,4-triazol-3-yl)-1-[4-(1H-pyrrol-1-yl)benzene-1-sulfonyl]piperidine, Bifunctional 3'-5' exonuclease/ATP-dependent helicase WRN, ... | | Authors: | Toms, A.V, Caravella, J.A, Sitnikov, N, Bartels, F, Svensson, R, Jacques O'Hagan, S, Borthwick, J, Campos, S, Yin, Y, Zhao, X, Li, L, Talbot, E, Kong, H, Freund, R.R.A, Browning, B, Genung, N.E, Carreiro, S, Brennan, D, Graves, A.P, Loh, C, Tummino, P, Edmonson, S.E, Li, D. | | Deposit date: | 2026-01-08 | | Release date: | 2026-05-13 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Design of Cyclic Vinyl Sulfones as WRN Covalent Inhibitors from Noncovalent Binders.
J.Med.Chem., 2026
|
|
10AK
 
 | | Crystal Structure of Human WRN helicase with compound 4 | | Descriptor: | 1,2-ETHANEDIOL, Bifunctional 3'-5' exonuclease/ATP-dependent helicase WRN, CHLORIDE ION, ... | | Authors: | Toms, A.V, Caravella, J.A, Sitnikov, N, Bartels, F, Svensson, R, Jacques O'Hagan, S, Borthwick, J, Yin, Y, Zhoa, X, Li, L, Liu, R, Talbot, E, Kong, H, Freund, R.R.A, Browning, B, Genung, N, Carreiro, S, Brennan, D, Graves, A.P, Loh, C, Tummino, P, Edmondson, S.D, Li, D. | | Deposit date: | 2026-01-08 | | Release date: | 2026-05-13 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | Design of Cyclic Vinyl Sulfones as WRN Covalent Inhibitors from Noncovalent Binders.
J.Med.Chem., 2026
|
|
10AP
 
 | | Crystal Structure of Human WRN helicase with compound 26 | | Descriptor: | (2R)-N-[(3R)-1,1-dioxo-1lambda~6~-thiolan-3-yl]-N-{[2-(2-hydroxypropan-2-yl)pyridin-4-yl]methyl}-2-methoxy-2-[(1M)-3,3',4'-trifluoro[1,1'-biphenyl]-4-yl]acetamide, 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Toms, A.V, Caravella, J.A, Sitnikov, N, Bartels, F, Svensson, R, Jacques O'Hagan, S, Borthwick, J, Campos, S, Yin, Y, Zhao, X, Li, L, Talbot, E, Kong, H, Freund, R.R.A, Browning, B, Genung, N.E, Carreiro, S, Brennan, D, Graves, A.P, Loh, C, Tummino, P, Edmonson, S.D, Li, D. | | Deposit date: | 2026-01-08 | | Release date: | 2026-05-13 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Design of Cyclic Vinyl Sulfones as WRN Covalent Inhibitors from Noncovalent Binders.
J.Med.Chem., 2026
|
|
10AY
 
 | | Cryo-EM structure of CRBN-DDB1 in complex with HBS1L and TNG961 | | Descriptor: | DNA damage-binding protein 1, HBS1-like protein, N-{6-[(3R)-2,6-dioxopiperidin-3-yl]naphthalen-1-yl}-N'-{2-[6-(trifluoromethyl)-1-benzothiophen-2-yl]propan-2-yl}urea, ... | | Authors: | Whittington, D.A. | | Deposit date: | 2026-01-09 | | Release date: | 2026-04-22 | | Last modified: | 2026-05-06 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | TNG961 is a selective oral HBS1L molecular glue degrader for the treatment of FOCAD-deleted cancers.
Cancer Discov, 2026
|
|
10FZ
 
 | | 30S ribosomal subunit from E. coli missing the gene encoding for the 16S rRNA 2'-O-methyltransferase RsmI | | Descriptor: | 16S rRNA, MAGNESIUM ION, Small ribosomal subunit protein bS16, ... | | Authors: | Barmada, M.I, Nandi, S, Conn, G.L. | | Deposit date: | 2026-01-18 | | Release date: | 2026-03-18 | | Last modified: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (2.91 Å) | | Cite: | Mechanism of 30S subunit recognition and modification by the conserved bacterial ribosomal RNA methyltransferase RsmI.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
10HY
 
 | | Structure of CHK1 10-pt. mutant complex with macrocyclic LRRK2 inhibitor compound 1 ((11R)-8-chloro-3,11-dimethyl-2-(oxan-4-yl)-2,4,10,11,12,13-hexahydro-9,5-(azeno)pyrazolo[3,4-b][1,4,6,10]oxatriazacyclotridecine) | | Descriptor: | (11R)-8-chloro-3,11-dimethyl-2-(oxan-4-yl)-2,4,10,11,12,13-hexahydro-9,5-(azeno)pyrazolo[3,4-b][1,4,6,10]oxatriazacyclotridecine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L, Yu, E.C, Zhou, H. | | Deposit date: | 2026-01-21 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-22 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Discovery of Potent, Selective, CNS-Penetrant Macrocyclic LRRK2 Inhibitors for the Treatment of Parkinson's Disease.
J.Med.Chem., 69, 2026
|
|
10HZ
 
 | | Structure of CHK1 10-pt. mutant complex with macrocyclic LRRK2 inhibitor compound 7 ((10aS,13aS)-3-cyclobutyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine) | | Descriptor: | (10aS,13aS)-3-cyclobutyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L, Yu, E.C, Zhou, H. | | Deposit date: | 2026-01-21 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-22 | | Method: | X-RAY DIFFRACTION (1.666 Å) | | Cite: | Discovery of Potent, Selective, CNS-Penetrant Macrocyclic LRRK2 Inhibitors for the Treatment of Parkinson's Disease.
J.Med.Chem., 69, 2026
|
|
10IA
 
 | | Structure of CHK1 10-pt. mutant complex with macrocyclic LRRK2 inhibitor compound 12 ((10aS,13aS)-3-cyclopropyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine) | | Descriptor: | (10aS,13aS)-3-cyclopropyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L, Yu, E.C, Zhou, H. | | Deposit date: | 2026-01-21 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-22 | | Method: | X-RAY DIFFRACTION (1.739 Å) | | Cite: | Discovery of Potent, Selective, CNS-Penetrant Macrocyclic LRRK2 Inhibitors for the Treatment of Parkinson's Disease.
J.Med.Chem., 69, 2026
|
|
10IC
 
 | | Rhesus rotavirus (consensus structure at 4.7 Angstrom resolution from cryo-ET) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, CHLORIDE ION, ... | | Authors: | de Sautu, M, Leistner, C, Kirchhausen, T, Jenni, S, Harrison, S.C. | | Deposit date: | 2026-01-21 | | Release date: | 2026-03-04 | | Method: | ELECTRON MICROSCOPY (4.7 Å) | | Cite: | Mechanism of membrane perforation in rotavirus cell entry.
Biorxiv, 2026
|
|
10JT
 
 | | CRYSTAL STRUCTURE OF KIRSTEN RAT SARCOMA G12C COMPLEXED WITH GMPPNP AND COVALENTLY BOUND TO 1-[(2R,3R)-3-{[(7P)-7-(8-ethynyl-7-fluoronaphthalen-1-yl)-8-fluoro-2-{ [(2R,4R,7aS)-2-fluorotetrahydro-1H-pyrrolizin-7a(5H)-yl]methoxy}pyrido[4,3-d] pyrimidin-4-yl](methyl)amino}-2-methylpyrrolidin-1-yl]-3-(pyrazin-2-yl)propan-1-one | | Descriptor: | 1-[(2R,3R)-3-{[(7P)-7-(8-ethynyl-7-fluoronaphthalen-1-yl)-8-fluoro-2-{[(2R,4R,7aS)-2-fluorotetrahydro-1H-pyrrolizin-7a(5H)-yl]methoxy}pyrido[4,3-d]pyrimidin-4-yl](methyl)amino}-2-methylpyrrolidin-1-yl]-3-(pyrazin-2-yl)propan-1-one, CALCIUM ION, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Sheriff, S. | | Deposit date: | 2026-01-22 | | Release date: | 2026-03-04 | | Last modified: | 2026-03-18 | | Method: | X-RAY DIFFRACTION (1.489 Å) | | Cite: | Optimization of Covalent Warhead Trajectory for KRAS G12C Active-State Inhibition.
J.Med.Chem., 69, 2026
|
|
10KZ
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors | | Descriptor: | (3'R)-1'-(1-benzyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, Plasma kallikrein | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-26 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10LR
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable PlasmaKallikrein Inhibitors complex with Compound 4 ((3'R)-1'-(5-amino-1-benzyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-(5-amino-1-benzyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-27 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.583 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10MV
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors complex with Compound 15 ((3'R)-1'-(5-amino-1-phenyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-(5-amino-1-phenyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Ogawa, A, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-28 | | Release date: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10MW
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors compound 25 ((3'R)-1'-{(1P)-5-amino-1-[2-(trifluoromethoxy)phenyl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-{(1P)-5-amino-1-[2-(trifluoromethoxy)phenyl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, Je, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Poiou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-28 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10NU
 
 | | Structure of kRas G12C bound to Inhibitor 13ab | | Descriptor: | 1-((2R,5S)-4-((S)-6-chloro-7-(1,6-dimethyl-1H-indazol-7-yl)-8-fluoro-2-(((S)-1-methylpyrrolidin-2-yl)methoxy)quinazolin-4-yl)-2,5-dimethylpiperazin-1-yl)prop-2-en-1-one, CALCIUM ION, GLYCEROL, ... | | Authors: | Shaffer, P.L, Milligan, C, Peters, U. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Optimization of Covalent 6-Cyanoquinazoline KRAS G12C Inhibitors for the Treatment of Solid Tumors.
J.Med.Chem., 69, 2026
|
|
10NV
 
 | | Structure of kRas G12C Bound to Inhibitor 13ba | | Descriptor: | 4-((2S,5R)-4-Acryloyl-2,5-dimethylpiperazin-1-yl)-7-(1,6-dimethyl-1H-indazol-7-yl)-8-fluoro-2-(((S)-1-methylpyrrolidin-2-yl)methoxy)quinazoline-6-carbonitrile, CALCIUM ION, GTPase KRas, ... | | Authors: | Shaffer, P.L, Milligan, C. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Optimization of Covalent 6-Cyanoquinazoline KRAS G12C Inhibitors for the Treatment of Solid Tumors.
J.Med.Chem., 69, 2026
|
|
10OO
 
 | | FGFR2 mutant D650V with compound 4 (AZD3463) | | Descriptor: | (4P)-N-[4-(4-aminopiperidin-1-yl)-2-methoxyphenyl]-5-chloro-4-(1H-indol-3-yl)pyrimidin-2-amine, Fibroblast growth factor receptor 2, GLYCEROL, ... | | Authors: | Hoffman, I.D, Nelson, K.J. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure-Based Design of a Novel Covalent 4-(1-Methylindol-3-yl)pyrimidin-2-amine Series Targeting FGFR2 Resistance Mutations.
J.Med.Chem., 69, 2026
|
|
10OQ
 
 | | FGFR2 mutant D650V with compound 6 | | Descriptor: | Fibroblast growth factor receptor 2, GLYCEROL, N-[(3M)-3-{5-chloro-2-[4-(morpholin-4-yl)anilino]pyrimidin-4-yl}-1-methyl-1H-indol-6-yl]propanamide, ... | | Authors: | Hoffman, I.D, Nelson, K.J, Bensen, D.C, Rideout, M, Hudkins, R.L, Frye, C, Bailey, J.B. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure-Based Design of a Novel Covalent 4-(1-Methylindol-3-yl)pyrimidin-2-amine Series Targeting FGFR2 Resistance Mutations.
J.Med.Chem., 69, 2026
|
|
10OU
 
 | | FGFR2 mutant D650V with compound 12 | | Descriptor: | 1,2-ETHANEDIOL, Fibroblast growth factor receptor 2, GLYCEROL, ... | | Authors: | Hoffman, I.D, Nelson, K.J, Bensen, D.C, Rideout, M, Hudkins, R.L, Frye, C, Bailey, J.B. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structure-Based Design of a Novel Covalent 4-(1-Methylindol-3-yl)pyrimidin-2-amine Series Targeting FGFR2 Resistance Mutations.
J.Med.Chem., 69, 2026
|
|
10PX
 
 | | Crystal structure of the wild-type Thermus thermophilus 70S ribosome in complex with benzoxaborole derivative of azithromycin (AZI-BB2), mRNA, aminoacylated A-site Phe-tRNAphe, aminoacylated P-site fMet-tRNAmet, and deacylated E-site tRNAphe at 2.45A resolution | | Descriptor: | 16S Ribosomal RNA, 23S Ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Chen, C.-W, Volynkina, I.A, Bortyazh, M.O, Tereshchenkov, A.G, Karakchieva, A.O, Lukianov, D.A, Komarova, E.S, Tupikin, A.E, Skvortsov, D.A, Tevyashova, A.N, Tikhomirov, A.S, Tashlitsky, V.N, Kabilov, M.R, Shchekotikhin, A.E, Dontsova, O.A, Sergiev, P.V, Polikanov, Y.S. | | Deposit date: | 2026-02-01 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-08 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Benzoxaborole-modified azithromycins inhibit translation without inducing ermC expression.
Antimicrob.Agents Chemother., 2026
|
|
10QS
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors Compound 13 ((3'R)-1'-{5-amino-1-[(2S)-1,1,1-trifluorobutan-2-yl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-{5-amino-1-[(2S)-1,1,1-trifluorobutan-2-yl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Plasma kallikrein | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yany, S, Cheng, A, Ellsworth, K, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogowa, A. | | Deposit date: | 2026-02-02 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.733 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10RA
 
 | | Human SARM1 TIR domain bound to compound 6 | | Descriptor: | 1-[(2R,3R,4S,5R)-5-({[(S)-{[(R)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-3,4-dihydroxyoxolan-2-yl]-4-[(4S,6P)-6-(5-fluoro-1H-indol-2-yl)imidazo[1,2-a]pyrazin-2-yl]pyridin-1-ium (non-preferred name), Sterile alpha and TIR motif-containing protein 1 | | Authors: | Olland, A.M, Johnson, Z.L, Nomme, J, Ciesielski, F, Ronin, C. | | Deposit date: | 2026-02-02 | | Release date: | 2026-04-22 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structure-Based Discovery of Imidazo[4,5- c ]pyridine SARM1 Modulators Showing Paradoxical Activation.
J.Med.Chem., 69, 2026
|
|
10RB
 
 | | Human SARM1 TIR domain bound to compound 14 | | Descriptor: | 1-[(2R,3R,4S,5R)-5-({[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-3,4-dihydroxyoxolan-2-yl]-4-[(6P)-7-{[2-(dimethylamino)ethyl]amino}-6-(5-fluoro-1H-indol-2-yl)-1-methyl-1H-imidazo[4,5-c]pyridin-2-yl]pyridin-1-ium (non-preferred name), NAD(+) hydrolase SARM1 | | Authors: | Dementiev, A, Olland, A.M, Johnson, Z.L. | | Deposit date: | 2026-02-02 | | Release date: | 2026-04-22 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure-Based Discovery of Imidazo[4,5- c ]pyridine SARM1 Modulators Showing Paradoxical Activation.
J.Med.Chem., 69, 2026
|
|
10RC
 
 | | Human SARM1 TIR domain bound to compound 22 | | Descriptor: | 1-[(2R,3R,4S,5R)-5-({[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-3,4-dihydroxyoxolan-2-yl]-4-{(6P)-7-{[2-(dimethylamino)ethyl]amino}-6-[6-fluoro-4-(pyrimidin-2-yl)-1H-indol-2-yl]-1-methyl-1H-imidazo[4,5-c]pyridin-2-yl}pyridin-1-ium (non-preferred name), GLYCEROL, NAD(+) hydrolase SARM1 | | Authors: | Dementiev, A, Johnson, Z.L, Olland, A.M. | | Deposit date: | 2026-02-02 | | Release date: | 2026-04-22 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-Based Discovery of Imidazo[4,5- c ]pyridine SARM1 Modulators Showing Paradoxical Activation.
J.Med.Chem., 69, 2026
|
|







