-Search query
-Search result
Showing 1 - 50 of 109 items for (author: miyata & t)

EMDB-64036: 
Cryo-EM structure of the Lhcp trimer from Ostreococcus tauri at 1.94 angstrom resolution
Method: single particle / : Seki S, Kubota M, Ishii A, Kim E, Tanaka H, Miyata T, Namba K, Kurisu G, Minagawa J, Fujii R

PDB-9uc6: 
Cryo-EM structure of the Lhcp trimer from Ostreococcus tauri at 1.94 angstrom resolution
Method: single particle / : Seki S, Kubota M, Ishii A, Kim E, Tanaka H, Miyata T, Namba K, Kurisu G, Minagawa J, Fujii R

EMDB-63799: 
Cryo-EM structure of violaxanthin-chlorophyll-a-binding protein with red shifted Chl a (rVCP) from Trachydiscus minutus at 2.4 angstrom
Method: single particle / : Seki S, Litvin R, Bina D, Tanaka H, Miyata T, Namba K, Kurisu G, Polivka T, Fujii R

PDB-9mcc: 
Cryo-EM structure of violaxanthin-chlorophyll-a-binding protein with red shifted Chl a (rVCP) from Trachydiscus minutus at 2.4 angstrom
Method: single particle / : Seki S, Litvin R, Bina D, Tanaka H, Miyata T, Namba K, Kurisu G, Polivka T, Fujii R

EMDB-62570: 
Cryo-EM structure of the TIA-1 prion-like domain amyloid fibril, WT
Method: helical / : Inaoka D, Miyata T, Makino F, Ohtani Y, Ekari M, Kobayashi R, Imamura K, Sakamoto E, Kodama ST, Yoshida N, Kato T, Namba K, Tochio H, Sekiyama N

EMDB-62571: 
Cryo-EM structure of the TIA-1 prion-like domain amyloid fibril, G355R
Method: helical / : Inaoka D, Miyata T, Makino F, Ohtani Y, Ekari M, Kobayashi R, Imamura K, Sakamoto E, Kodama ST, Yoshida N, Kato T, Namba K, Tochio H, Sekiyama N

PDB-9kty: 
Cryo-EM structure of the TIA-1 prion-like domain amyloid fibril, WT
Method: helical / : Inaoka D, Miyata T, Makino F, Ohtani Y, Ekari M, Kobayashi R, Imamura K, Sakamoto E, Kodama ST, Yoshida N, Kato T, Namba K, Tochio H, Sekiyama N

PDB-9ktz: 
Cryo-EM structure of the TIA-1 prion-like domain amyloid fibril, G355R
Method: helical / : Inaoka D, Miyata T, Makino F, Ohtani Y, Ekari M, Kobayashi R, Imamura K, Sakamoto E, Kodama ST, Yoshida N, Kato T, Namba K, Tochio H, Sekiyama N

EMDB-63050: 
Cryo-EM structure of linker-extended biparatopic antibody BA1-GP4 in complex with TNFR2
Method: single particle / : Otsuki T, Matsumoto S, Fujita J, Miyata T, Namba K, Kanada R, Okuno Y, Kamada H, Ohno H, Akiba H

PDB-9lfl: 
Cryo-EM structure of linker-extended biparatopic antibody BA1-GP4 in complex with TNFR2
Method: single particle / : Otsuki T, Matsumoto S, Fujita J, Miyata T, Namba K, Kanada R, Okuno Y, Kamada H, Ohno H, Akiba H

EMDB-61993: 
Structure of the Salmonella flagellar FliPQR complex reconstituted in the peptidisc
Method: single particle / : Kinoshita M, Miyata T, Makino F, Imada K, Namba K, Minamino T

PDB-9k29: 
Structure of the Salmonella flagellar FliPQR complex reconstituted in the peptidisc
Method: single particle / : Kinoshita M, Miyata T, Makino F, Imada K, Namba K, Minamino T

EMDB-61725: 
Cryo-EM Structure of Fructose Dehydrogenase Variant from Gluconobacter japonicus Truncating Heme 1c and C-Terminal Hydrophobic Regions
Method: single particle / : Adachi T, Ichikawa K, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Shirai O

PDB-9jqa: 
Cryo-EM Structure of Fructose Dehydrogenase Variant from Gluconobacter japonicus Truncating Heme 1c and C-Terminal Hydrophobic Regions
Method: single particle / : Adachi T, Ichikawa K, Miyata T, Makino F, Tanaka H, Namba K, Sowa K

EMDB-60747: 
Cryo-EM structure of Escherichia coli hibernating ribosome with RNase I mutant
Method: single particle / : Tanzawa T, Minami A, Yoshida H, Kato T, Ogawa T

EMDB-64894: 
Tomogram of a Candidatus Margulisarchaeum peptidophila strain HC1 cell
Method: electron tomography / : Imachi H, Hosogi N

EMDB-60251: 
mouse TMEM63b in LMNG-CHS micelle with YN9303-24 Fab
Method: single particle / : Miyata Y, Takahashi K, Lee Y, Sultan CS, Kuribayashi R, Takahashi M, Hata K, Bamba T, Izumi Y, Liu K, Uemura T, Nomura N, Iwata S, Nagata S, Nishizawa T, Segawa K

EMDB-60252: 
mouse TMEM63b in DDM-CHS micelle
Method: single particle / : Miyata Y, Takahashi K, Lee Y, Sultan CS, Kuribayashi R, Takahashi M, Hata K, Bamba T, Izumi Y, Liu K, Uemura T, Nomura N, Iwata S, Nagata S, Nishizawa T, Segawa K

EMDB-39676: 
Cryo-EM structure of cylindrical fiber of MyD88 TIR
Method: single particle / : Kasai K, Imamura K, Narita A, Makino F, Miyata T, Kato T, Namba K, Onishi H, Tochio H

PDB-8yym: 
Cryo-EM structure of cylindrical fiber of MyD88 TIR
Method: single particle / : Kasai K, Imamura K, Narita A, Makino F, Miyata T, Kato T, Namba K, Onishi H, Tochio H

EMDB-60718: 
Cryo-EM structure of G1-ATPase dimer from Mycoplasma mobile gliding machinery
Method: single particle / : Toyonaga T, Kato T, Kawamoto A, Miyata T, Kawakami K, Fujita J, Hamaguchi T, Namba K, Miyata M

PDB-9io5: 
Cryo-EM structure of G1-ATPase dimer from Mycoplasma mobile gliding machinery
Method: single particle / : Toyonaga T, Kato T, Kawamoto A, Miyata T, Kawakami K, Fujita J, Hamaguchi T, Namba K, Miyata M

EMDB-63267: 
Tomogram of a Candidatus Flexarchaeum multiprotrusionis strain SC1 cell
Method: electron tomography / : Murata K, Kayama Y, Imachi H

EMDB-63316: 
Tomogram of a Candidatus Margulisarchaeum peptidophila strain HC1 cell
Method: electron tomography / : Murata K, Kayama Y, Imachi H

EMDB-63317: 
Tomogram of a Candidatus Margulisarchaeum peptidophila strain HC1 cell
Method: electron tomography / : Murata K, Kayama Y, Imachi H

EMDB-63318: 
Tomogram of Candidatus Flexarchaeum multiprotrusionis strain SC1 cell
Method: electron tomography / : Murata K, Kayama Y, Imachi H

EMDB-63319: 
Tomogram of Candidatus Flexarchaeum multiprotrusionis strain SC1 cell
Method: electron tomography / : Murata K, Kayama Y, Imachi H

EMDB-63320: 
Tomogram of a Candidatus Flexarchaeum multiprotrusionis strain SC1 cell
Method: electron tomography / : Murata K, Kayama Y, Imachi H

EMDB-19402: 
Single-particle cryo-EM of Mycoplasma pneumoniae adhesin P1 complexed with the anti-adhesive Fab fragment.
Method: single particle / : Vizarraga D, Kawamoto A, Marcos-Silva M, Fita I, Miyata M, Pinyol J, Namba K, Kenri T

PDB-8ror: 
Single-particle cryo-EM of Mycoplasma pneumoniae adhesin P1 complexed with the anti-adhesive Fab fragment.
Method: single particle / : Vizarraga D, Kawamoto A, Marcos-Silva M, Fita I, Miyata M, Pinyol J, Namba K, Kenri T

EMDB-19403: 
Single-particle cryo-EM of the P1-P40P90 heterodimer from Mycoplasma pneumoniae below 10 Angstrom resolution.
Method: single particle / : Kawamoto A, Vizarraga D, Marcos-Silva M, Fita I, Miyata M, Pinyol J, Namba K, Kenri T

EMDB-60007: 
Structure of the Salmonella flagellar MS-ring with C11 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-60008: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C33 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-60009: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C34 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-62210: 
Structure of the 34-mer RBM3 ring of Salmonella flagellar MS-ring protein FliF with C1 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-62211: 
Structure of the 33-mer RBM3 ring of Salmonella flagellar MS-ring protein FliF with C1 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

PDB-8zds: 
Structure of the Salmonella flagellar MS-ring with C11 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K
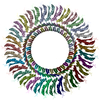
PDB-8zdt: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C33 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K
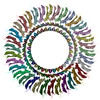
PDB-8zdu: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C34 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-38825: 
Cryo-EM structure of Light-harvesting complex from Spinacia oleracea
Method: single particle / : Seki S, Miyata T, Tanaka H, Namba K, Kurisu G, Fujii R

EMDB-39192: 
Cryo-EM structure of in vitro reconstituted Light-harvesting complex II trimer
Method: single particle / : Seki S, Miyata T, Tanaka H, Namba K, Kurisu G, Fujii R

EMDB-39304: 
Cryo-EM structure of light-harvesting complex II with C1 symmetry
Method: single particle / : Seki S, Miyata T, Tanaka H, Namba K, Kurisu G, Fujii R

EMDB-39305: 
Cryo-EM structure of in vitro reconstituted LHCII with C1 symmetry
Method: single particle / : Seki S, Miyata T, Tanaka H, Namba K, Kurisu G, Fujii R

PDB-8y15: 
Cryo-EM structure of Light-harvesting complex from Spinacia oleracea
Method: single particle / : Seki S, Miyata T, Tanaka H, Namba K, Kurisu G, Fujii R

PDB-8yee: 
Cryo-EM structure of in vitro reconstituted Light-harvesting complex II trimer
Method: single particle / : Seki S, Miyata T, Tanaka H, Namba K, Kurisu G, Fujii R

EMDB-37501: 
mouse TMEM63b in LMNG-CHS micelle
Method: single particle / : Miyata Y, Takahashi K, Lee Y, Sultan CS, Kuribayashi R, Takahashi M, Hata K, Bamba T, Izumi Y, Liu K, Uemura T, Nomura N, Iwata S, Nagata S, Nishizawa T, Segawa K

EMDB-37502: 
mouse TMEM63b in DDM-CHS micelle with YN9303-24 Fab
Method: single particle / : Miyata Y, Takahashi K, Lee Y, Sultan CS, Kuribayashi R, Takahashi M, Hata K, Bamba T, Izumi Y, Liu K, Uemura T, Nomura N, Iwata S, Nagata S, Nishizawa T, Segawa K

PDB-8wg3: 
mouse TMEM63b in LMNG-CHS micelle
Method: single particle / : Miyata Y, Takahashi K, Lee Y, Sultan CS, Kuribayashi R, Takahashi M, Hata K, Bamba T, Izumi Y, Liu K, Uemura T, Nomura N, Iwata S, Nagata S, Nishizawa T, Segawa K

PDB-8wg4: 
mouse TMEM63b in DDM-CHS micelle with YN9303-24 Fab
Method: single particle / : Miyata Y, Takahashi K, Lee Y, Sultan CS, Kuribayashi R, Takahashi M, Hata K, Bamba T, Izumi Y, Liu K, Uemura T, Nomura N, Iwata S, Nagata S, Nishizawa T, Segawa K

EMDB-37355: 
Cryo-EM structure of helical filament of MyD88 TIR
Method: single particle / : Kasai K, Imamura K, Narita A, Makino F, Miyata T, Kato T, Namba K, Onishi H, Tochio H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
