[English] 日本語
 Yorodumi
Yorodumi- EMDB-60959: Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292)... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | flagellin / tail / flagella / filament / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationTLR5 cascade / MyD88 cascade initiated on plasma membrane / NFkB and MAPK activation mediated by TRAF6 / The IPAF inflammasome / bacterial-type flagellum / bacterial-type flagellum-dependent cell motility / receptor ligand activity / structural molecule activity / : Similarity search - Function | |||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) | |||||||||
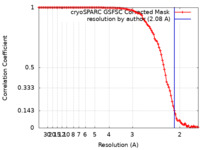
| Method | helical reconstruction / cryo EM / Resolution: 2.08 Å | |||||||||
 Authors Authors | Waraich K / Makino F / Miyata T / Kinoshita M / Minamino T / Namba K | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament Authors: Waraich K / Makino F / Miyata T / Kinoshita M / Minamino T / Namba K | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60959.map.gz emd_60959.map.gz | 37.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60959-v30.xml emd-60959-v30.xml emd-60959.xml emd-60959.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_60959_fsc.xml emd_60959_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_60959.png emd_60959.png | 73.3 KB | ||
| Filedesc metadata |  emd-60959.cif.gz emd-60959.cif.gz | 6.1 KB | ||
| Others |  emd_60959_half_map_1.map.gz emd_60959_half_map_1.map.gz emd_60959_half_map_2.map.gz emd_60959_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60959 http://ftp.pdbj.org/pub/emdb/structures/EMD-60959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60959 | HTTPS FTP |
-Related structure data
| Related structure data |  9iwqMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60959.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60959.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.87 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_60959_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
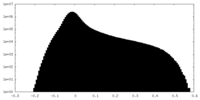
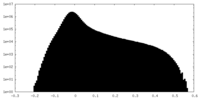
| Projections & Slices |
| ||||||||||||
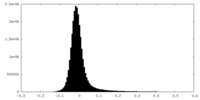
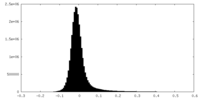
| Density Histograms |
-Half map: #1
| File | emd_60959_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292)...
| Entire | Name: Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament |
|---|---|
| Components |
|
-Supramolecule #1: Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292)...
| Supramolecule | Name: Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) |
-Macromolecule #1: Flagellin
| Macromolecule | Name: Flagellin / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) |
| Molecular weight | Theoretical: 42.587551 KDa |
| Recombinant expression | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 (bacteria) |
| Sequence | String: AQVINTNSLS LLTQNNLNKS QSALGTAIER LSSGLRINSA KDDAAGQAIA NRFTANIKGL TQASRNANDG ISIAQTTEGA LNEINNNLQ RVRELAVQSA NSTNSQSDLD SIQAEITQRL NEIDRVSGQT QFNGVKVLAQ DNTLTIQVGA NDGETIDIDL K QINSQTLG ...String: AQVINTNSLS LLTQNNLNKS QSALGTAIER LSSGLRINSA KDDAAGQAIA NRFTANIKGL TQASRNANDG ISIAQTTEGA LNEINNNLQ RVRELAVQSA NSTNSQSDLD SIQAEITQRL NEIDRVSGQT QFNGVKVLAQ DNTLTIQVGA NDGETIDIDL K QINSQTLG LDTLNVQQKY KVSDTAATVT GYADTTIALD NSTFKAALTA AGVTGTASVV KMSYTDNNGK TIDGGLAVKV GD DYYSATQ NKDGSISINT TKYTADDGTS KTALNKLGGA DGKTEVVSIG GKTYAASKAE GHNFKAQPDL AEAAATTTEN PLQ KIDAAL AQVDTLRSDL AAVQNRFNSA ITNLGNTVNN LTSARSRIED SDYATEVSNM SRAQILQQAG TSVLAQANQV PQNV LSLLR UniProtKB: Flagellin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)