-Search query
-Search result
Showing 1 - 50 of 81 items for (author: hirose & t)

EMDB-61568: 
human GM-CSF with Fabs from autoantibodies, F1 and BD.
Method: single particle / : Kishikawa J, Kato T, Kurosaki T, Inoue T

EMDB-61569: 
human GM-CSF with Fabs from autoantibodies, F1 and C.
Method: single particle / : Kishikawa J, Kato T, Kurosaki T, Inoue T

EMDB-67812: 
MexBYB-Ka asymmetry
Method: single particle / : Wang J, Nakagawa A, Yamashita E

EMDB-67813: 
MexBYB-Ka symmetry-like
Method: single particle / : Wang J, Nakagawa A, Yamashita E

EMDB-67814: 
MexBYB-apo asymmetry
Method: single particle / : Wang J, Nakagawa A, Yamashita E

EMDB-67815: 
MexBYB-apo symmetry-like
Method: single particle / : Wang J, Nakagawa A, Yamashita E

PDB-22xk: 
MexBYB-Ka asymmetry
Method: single particle / : Wang J, Nakagawa A, Yamashita E

PDB-22xm: 
MexBYB-Ka symmetry-like
Method: single particle / : Wang J, Nakagawa A, Yamashita E

EMDB-68587: 
Cryo-EM consensus map of Bovine lactoferrin in water
Method: single particle / : Wada M, Yamazaki K, Wada Y, Ohga K, Ishii Y, Nakagawa A, Yoshikara H, Okuno S, Nojima T, Ochi H, Hirose M, Kato T, Katoh T

EMDB-68586: 
Cryo-EM consensus map of Bovine lactoferrin in TBS buffer
Method: single particle / : Wada M, Yamazaki K, Wada Y, Ohga K, Ishii Y, Nakagawa A, Yoshikawa H, Okuno S, Nojima T, Ochi H, Hirose M, Kato T, Katoh T

EMDB-64947: 
Cryo-EM structure of the human kappa opioid receptor signaling complex bound to compound A
Method: single particle / : Suno-Ikeda C, Sugita Y, Hirose M, Suno R

EMDB-65622: 
Cryo-EM structure of human kappa opioid receptor -G protein signaling complex bound with U-50488H
Method: single particle / : Suno-Ikeda C, Takai T, Hirose M, Inoue A, Sugita Y, Kato T, Kobayashi T, Suno R

PDB-9v6o: 
Cryo-EM structure of human kappa opioid receptor - G protein signaling complex bound with nalfurafine.
Method: single particle / : Suno-Ikeda C, Takai T, Hirose M, Inoue A, Sugita Y, Kato T, Kobayashi T, Suno R

PDB-9w49: 
Cryo-EM structure of human kappa opioid receptor -G protein signaling complex bound with U-50488H
Method: single particle / : Suno-Ikeda C, Takai T, Hirose M, Inoue A, Sugita Y, Kato T, Kobayashi T, Suno R

EMDB-39679: 
A tetrameric STAT1-DNA complex
Method: single particle / : Sugiyama A, Minami M, Sugita Y, Ose T

EMDB-39680: 
A dimeric STAT1-DNA complex
Method: single particle / : Sugiyama A, Minami M, Sugita Y, Ose T

PDB-8yyu: 
A tetrameric STAT1-DNA complex
Method: single particle / : Sugiyama A, Minami M, Sugita Y, Ose T

PDB-8yyv: 
A dimeric STAT1-DNA complex
Method: single particle / : Sugiyama A, Minami M, Sugita Y, Ose T

EMDB-61213: 
Cryo-EM structure of wild type Aquifex aeolicus RseP in complex with Fab
Method: single particle / : Asahi K, Hirose M, Aruga R, Kato T, Nogi T

EMDB-61214: 
Cryo-EM structure of Aquifex aeolicus RseP E18Q mutant in complex with Fab
Method: single particle / : Asahi K, Hirose M, Aruga R, Kato T, Nogi T

PDB-9j82: 
Cryo-EM structure of wild type Aquifex aeolicus RseP in complex with Fab
Method: single particle / : Asahi K, Hirose M, Aruga R, Kato T, Nogi T

PDB-9j83: 
Cryo-EM structure of Aquifex aeolicus RseP E18Q mutant in complex with Fab
Method: single particle / : Asahi K, Hirose M, Aruga R, Kato T, Nogi T

EMDB-60580: 
Bacterial flagellar sodium-driven stator PomA5PomB2 with 100 mM NaCl
Method: single particle / : Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K

EMDB-60581: 
Bacterial flagellar sodium-driven stator PomA5PomB2 with 100 mM KCl
Method: single particle / : Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K

EMDB-60584: 
Bacterial flagellar sodium-driven stator PomA5PomB2(D24N) with 100 mM NaCl
Method: single particle / : Nishikino T, Kishikiwa J, Hirose M, Kato T, Imada K

EMDB-60585: 
Bacterial flagellar sodium-driven stator PomA5PomB2(D24N) with 100 mM KCl
Method: single particle / : Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K

EMDB-60636: 
Bacterial flagellar sodium-driven stator PomA5PomB2 with 100 mM NaCl and 0.1 mM phenamil
Method: single particle / : Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K

PDB-8zyv: 
Bacterial flagellar sodium-driven stator PomA5PomB2 with 100 mM NaCl
Method: single particle / : Nishikino T, Takekawa N, Imada K

PDB-8zyw: 
Bacterial flagellar sodium-driven stator PomA5PomB2 with 100 mM KCl
Method: single particle / : Nishikino T, Takekawa N, Imada K

PDB-8zyz: 
Bacterial flagellar sodium-driven stator PomA5PomB2(D24N) with 100 mM NaCl
Method: single particle / : Nishikino T, Takekawa N, Imada K

PDB-8zz0: 
Bacterial flagellar sodium-driven stator PomA5PomB2(D24N) with 100 mM KCl
Method: single particle / : Nishikino T, Takekawa N, Imada K

PDB-9ijm: 
Bacterial flagellar sodium-driven stator PomA5PomB2 with 100 mM NaCl and 0.1 mM phenamil
Method: single particle / : Nishikino T, Takekawa N, Imada K

EMDB-60321: 
3D structure of Y-50 TCR-TMM-CD1b ternary complex
Method: single particle / : Asa M, Hirose M, Nagae M, Yamasaki S, Kato T

PDB-8zox: 
3D structure of Y-50 TCR-TMM-CD1b ternary complex
Method: single particle / : Asa M, Hirose M, Nagae M, Yamasaki S, Kato T

EMDB-39761: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 34-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-39763: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 35-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-39764: 
Homomeric 34mer of the Vibrio flagellar MS-ring protein FliF without symmetry imposition
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-39765: 
Homomeric 35mer of the Vibrio flagellar MS-ring protein FliF without symmetry imposition
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

PDB-8z4d: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 34-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M
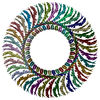
PDB-8z4g: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 35-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-37480: 
PSI-LHCI of the red alga Cyanidium caldarium RK-1 (NIES-2137)
Method: single particle / : Kato K, Hamaguchi T, Nakajima Y, Kawakami K, Yonekura K, Shen JR, Nagao R

PDB-8wey: 
PSI-LHCI of the red alga Cyanidium caldarium RK-1 (NIES-2137)
Method: single particle / : Kato K, Hamaguchi T, Nakajima Y, Kawakami K, Yonekura K, Shen JR, Nagao R

EMDB-35029: 
SARS-CoV2 spike protein with ACE2, no ACE2 binding.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35030: 
SARS-CoV2 spike protein with ACE2. 1 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35031: 
SARS-CoV2 spike protein with ACE2. 2 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35032: 
SARS-CoV2 spike protein with ACE2. 3 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35036: 
SARS-CoV2 spike protein with ACE2 decoy.no ACE2 decoy binding
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35037: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35038: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound and 2 RBD up form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35039: 
SARS-CoV2 spike protein with ACE2 decoy. 2 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
