8UF7
 
 | | Cryo-EM structure of POmAb, a Type-I anti-prothrombin antiphospholipid antibody, bound to kringle-1 of human prothrombin | | Descriptor: | POmAb Heavy Chain, POmAb Light Chain, Prothrombin | | Authors: | Kumar, S, Summers, B, Basore, K, Pozzi, N. | | Deposit date: | 2023-10-03 | | Release date: | 2024-02-14 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Cryo-EM structure and functional basis of prothrombin recognition by a type I antiprothrombin antiphospholipid antibody.
Blood, 143, 2024
|
|
5TO3
 
 | | Crystal structure of thrombin mutant W215A/E217A fused to EGF456 of thrombomodulin via a 31-residue linker and bound to PPACK | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-beta-D-mannopyranose-(1-4)-beta-D-mannopyranose-(1-4)-alpha-D-mannopyranose-(1-4)-[beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, ... | | Authors: | Barranco-Medina, S, Murphy, M, Pelc, L, Chen, Z, Di Cera, E, Pozzi, N. | | Deposit date: | 2016-10-16 | | Release date: | 2017-03-29 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Rational Design of Protein C Activators.
Sci Rep, 7, 2017
|
|
6KKB
 
 | | A novel agonist of THRb | | Descriptor: | 2-[(1-methyl-4-oxidanyl-7-phenoxy-isoquinolin-3-yl)carbonylamino]ethanoic acid, SRC2-3, Thyroid hormone receptor beta | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2019-07-24 | | Release date: | 2019-10-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Revealing a Mutant-Induced Receptor Allosteric Mechanism for the Thyroid Hormone Resistance.
Iscience, 20, 2019
|
|
3D49
 
 | | Thrombin Inhibition | | Descriptor: | BENZAMIDINE, Hirudin variant-1, SODIUM ION, ... | | Authors: | Baum, B, Heine, A, Klebe, G. | | Deposit date: | 2008-05-14 | | Release date: | 2009-04-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Understanding Thrombin Inhibition
To be Published
|
|
3DA9
 
 | |
7WMO
 
 | | A novel chemical derivative(92) of THRB agonist | | Descriptor: | 2-[[1-ethoxy-7-[4-(3-fluoranyl-5-methoxy-phenyl)carbonyl-2,6-dimethyl-phenoxy]-4-oxidanyl-isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-15 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7WML
 
 | | A novel chemical derivative(85) of THRB agonist | | Descriptor: | 2-[[7-[4-(3,5-dimethylphenyl)carbonyl-2,6-dimethyl-phenoxy]-1-ethoxy-4-oxidanyl-isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-15 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7WMJ
 
 | | A novel chemical derivative(71) of THRB agonist | | Descriptor: | 2-[[7-[2,6-dimethyl-4-(phenylcarbonyl)phenoxy]-1-methoxy-4-oxidanyl-isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7WMH
 
 | | A novel chemical derivative(56) of THRB agonist | | Descriptor: | 2-[[1-methoxy-4-oxidanyl-7-(4-phenylphenoxy)isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7WMG
 
 | | A novel chemical derivative(52) of THRB agonist | | Descriptor: | 2-[[1-methoxy-4-oxidanyl-7-(4-phenoxyphenoxy)isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.93 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7WLX
 
 | | A novel chemical derivative(53) of THRB agonist | | Descriptor: | 2-[[1-methoxy-4-oxidanyl-7-[4-(phenylmethyl)phenoxy]isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-14 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7WMN
 
 | | A novel chemical derivative(89) of THRB agonist | | Descriptor: | 2-[[7-[2,6-dimethyl-4-(3-methylphenyl)carbonyl-phenoxy]-1-methoxy-4-oxidanyl-isoquinolin-3-yl]carbonylamino]ethanoic acid, Isoform Beta-2 of Thyroid hormone receptor beta, Nuclear receptor coactivator 2 | | Authors: | Yao, B.Q, Li, Y. | | Deposit date: | 2022-01-15 | | Release date: | 2022-05-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Discovery of a Highly Selective and H435R-Sensitive Thyroid Hormone Receptor beta Agonist.
J.Med.Chem., 65, 2022
|
|
7ZKO
 
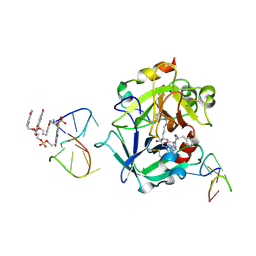 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form delta | | Descriptor: | 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
7ZKN
 
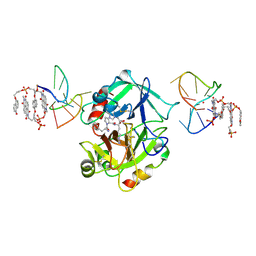 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form gamma | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.03 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
7ZKM
 
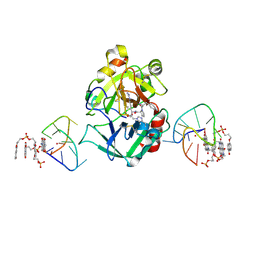 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form beta | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
7ZKL
 
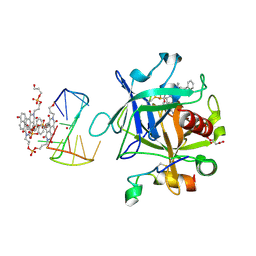 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form alpha | | Descriptor: | (2S)-2-hydroxybutanedioic acid, 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.18 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
2BVX
 
 | | Design and Discovery of Novel, Potent Thrombin Inhibitors with a Solubilizing Cationic P1-P2-Linker | | Descriptor: | ALPHA THROMBIN, HIRUDIN VARIANT-2, N-(5-CHLORO-BENZO[B]THIOPHEN-3-YLMETHYL)-2-[6-CHLORO-OXO-3-(2-PYRIDIN-2-YL-ETHYLAMINO)-2H-PYRAZIN-1-YL]-ACETAMIDE | | Authors: | Bulat, S, Bosio, S, Grabowski, E, Papadopoulos, M.A, Cerezo-Galvez, S, Rosenbaum, C, Matassa, V.G, Ott, I, Metz, G, Schamberger, J, Sekul, R, Feurer, A. | | Deposit date: | 2005-07-04 | | Release date: | 2006-10-25 | | Last modified: | 2016-12-21 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Design and Discovery of Novel, Potent Pyrazinone-Based Thrombin Inhibitors with a Solubilizing P1-P2-Linker
Lett.Drug Des.Discovery, 3, 2006
|
|
1ETT
 
 | |
1EB1
 
 | | Complex structure of human thrombin with N-methyl-arginine inhibitor | | Descriptor: | 3-CYCLOHEXYL-D-ALANYL-L-PROLYL-N~2~-METHYL-L-ARGININE, PEPTIDE INHIBITOR, THROMBIN HEAVY CHAIN, ... | | Authors: | Friedrich, R, Steinmetzer, T, Bode, W. | | Deposit date: | 2001-07-18 | | Release date: | 2002-01-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Methyl Group of N(Alpha)(Me)Arg-Containing Peptides Disturbs the Active-Site Geometry of Thrombin, Impairing Efficient Cleavage
J.Mol.Biol., 316, 2002
|
|
2BVR
 
 | | Human thrombin complexed with fragment-based small molecules occupying the S1 pocket | | Descriptor: | 2-[2-(4-CHLORO-PHENYLSULFANYL)-ACETYLAMINO]-3-(4-GUANIDINO-PHENYL)-PROPIONAMIDE, ALPHA THROMBIN, HIRUDIN VARIANT-2 | | Authors: | Neumann, T, Junker, H.-D, Keil, O, Burkert, K, Ottleben, H, Gamer, J, Sekul, R, Deppe, H, Feurer, A, Tomandl, D, Metz, G. | | Deposit date: | 2005-07-04 | | Release date: | 2006-10-25 | | Last modified: | 2016-12-21 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Discovery of Thrombin Inhibitor Fragments from Chemical Microarray Screening
Lett.Drug Des.Discovery, 2, 2005
|
|
2BVS
 
 | | Human thrombin complexed with fragment-based small molecules occupying the S1 pocket | | Descriptor: | ALPHA THROMBIN, HIRUDIN VARIANT-2, N-[2-(2-CARBAMOYLMETHOXY-ETHOXY)-ETHYL]-2-[2-(4-CHLORO-PHENYLSULFANYL)-ACETYLAMINO]-3-(4-GUANIDINO-PHENYL)-PROPIONAMIDE | | Authors: | Neumann, T, Junker, H.-D, Keil, O, Burkert, K, Ottleben, H, Gamer, J, Sekul, R, Deppe, H, Feurer, A, Tomandl, D, Metz, G. | | Deposit date: | 2005-07-04 | | Release date: | 2006-10-25 | | Last modified: | 2016-12-21 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Discovery of Thrombin Inhibitor Fragments from Chemical Microarray Screening
Lett.Drug Des.Discovery, 2, 2005
|
|
2BXT
 
 | | Design and Discovery of Novel, Potent Thrombin Inhibitors with a Solubilizing Cationic P1-P2-Linker | | Descriptor: | 6-CHLORO-1-(2-{[(5-CHLORO-1-BENZOTHIEN-3-YL)METHYL]AMINO}ETHYL)-3-[(2-PYRIDIN-2-YLETHYL)AMINO]-1,4-DIHYDROPYRAZIN-2-OL, ALPHA THROMBIN, HIRUDIN VARIANT-2 | | Authors: | Bulat, S, Bosio, S, Grabowski, E, Papadopoulos, M.A, Cerezo-Galvez, S, Rosenbaum, C, Matassa, V.G, Ott, I, Metz, G, Schamberger, J, Sekul, R, Feurer, A. | | Deposit date: | 2005-07-27 | | Release date: | 2006-10-26 | | Last modified: | 2016-12-21 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Design and Discovery of Novel, Potent Pyrazinone-Based Thrombin Inhibitors with a Solubilizing P1-P2-Linker
Lett.Drug Des.Discovery, 3, 2006
|
|
1EOL
 
 | | Design of P1' and P3' residues of trivalent thrombin inhibitors and their crystal structures | | Descriptor: | ALPHA THROMBIN, THROMBIN INHIBITOR P628 | | Authors: | Slon-Usakiewicz, J.J, Sivaraman, J, Li, Y, Cygler, M, Konishi, Y. | | Deposit date: | 2000-03-23 | | Release date: | 2000-05-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Design of P1' and P3' residues of trivalent thrombin inhibitors and their crystal structures.
Biochemistry, 39, 2000
|
|
1ETS
 
 | |
6PXJ
 
 | | Crystal structure of human thrombin mutant I16T | | Descriptor: | GLYCEROL, MAGNESIUM ION, Thrombin heavy chain, ... | | Authors: | Stojanovski, B, Chen, Z, Koester, S.K, Pelc, L.A, Di Cera, E. | | Deposit date: | 2019-07-26 | | Release date: | 2019-12-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Role of the I16-D194 ionic interaction in the trypsin fold.
Sci Rep, 9, 2019
|
|
