[English] 日本語
 Yorodumi
Yorodumi- PDB-2xks: Prion-like conversion during amyloid formation at atomic resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2xks | ||||||
|---|---|---|---|---|---|---|---|
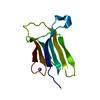
| Title | Prion-like conversion during amyloid formation at atomic resolution | ||||||
 Components Components | BETA-2-MICROGLOBULIN | ||||||
 Keywords Keywords | IMMUNE SYSTEM / AMYLOIDOSIS / IG-DOMAIN | ||||||
| Function / homology |  Function and homology information Function and homology informationearly endosome lumen / Nef mediated downregulation of MHC class I complex cell surface expression / DAP12 interactions / Endosomal/Vacuolar pathway / T cell mediated cytotoxicity / Antigen Presentation: Folding, assembly and peptide loading of class I MHC / regulation of iron ion transport / cellular response to iron(III) ion / negative regulation of iron ion transport / negative regulation of forebrain neuron differentiation ...early endosome lumen / Nef mediated downregulation of MHC class I complex cell surface expression / DAP12 interactions / Endosomal/Vacuolar pathway / T cell mediated cytotoxicity / Antigen Presentation: Folding, assembly and peptide loading of class I MHC / regulation of iron ion transport / cellular response to iron(III) ion / negative regulation of iron ion transport / negative regulation of forebrain neuron differentiation / antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent / peptide antigen assembly with MHC class I protein complex / ER to Golgi transport vesicle membrane / regulation of erythrocyte differentiation / response to molecule of bacterial origin / HFE-transferrin receptor complex / MHC class I peptide loading complex / transferrin transport / cellular response to iron ion / negative regulation of receptor-mediated endocytosis / positive regulation of T cell cytokine production / antigen processing and presentation of endogenous peptide antigen via MHC class I / MHC class I protein complex / peptide antigen assembly with MHC class II protein complex / negative regulation of neurogenesis / cellular response to nicotine / MHC class II protein complex / positive regulation of receptor-mediated endocytosis / multicellular organismal-level iron ion homeostasis / positive regulation of T cell mediated cytotoxicity / specific granule lumen / antigen processing and presentation of exogenous peptide antigen via MHC class II / positive regulation of immune response / peptide antigen binding / phagocytic vesicle membrane / recycling endosome membrane / positive regulation of T cell activation / negative regulation of epithelial cell proliferation / Interferon gamma signaling / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / Modulation by Mtb of host immune system / sensory perception of smell / tertiary granule lumen / positive regulation of cellular senescence / MHC class II protein complex binding / T cell differentiation in thymus / DAP12 signaling / late endosome membrane / negative regulation of neuron projection development / protein refolding / ER-Phagosome pathway / early endosome membrane / amyloid fibril formation / protein homotetramerization / intracellular iron ion homeostasis / learning or memory / endoplasmic reticulum lumen / Amyloid fiber formation / Golgi membrane / external side of plasma membrane / lysosomal membrane / focal adhesion / Neutrophil degranulation / SARS-CoV-2 activates/modulates innate and adaptive immune responses / structural molecule activity / endoplasmic reticulum / Golgi apparatus / protein homodimerization activity / : / extracellular exosome / extracellular region / membrane / identical protein binding / plasma membrane / cytosol Similarity search - Function | ||||||
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human) | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Eichner, T. / Kalverda, A.P. / Thompson, G.S. / Radford, S.E. / Homans, S.W. | ||||||
 Citation Citation |  Journal: Mol.Cell / Year: 2011 Journal: Mol.Cell / Year: 2011Title: Conformational Conversion During Amyloid Formation at Atomic Resolution. Authors: Eichner, T. / Kalverda, A.P. / Thompson, G.S. / Homans, S.W. / Radford, S.E. #1: Journal: J.Mol.Biol. / Year: 2009 Title: A Generic Mechanism of Beta2-Microglobulin Amyloid Assembly at Neutral Ph Involving a Specific Proline Switch. Authors: Eichner, T. / Radford, S.E. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: AUTHOR PROVIDED. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2xks.cif.gz 2xks.cif.gz | 935.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2xks.ent.gz pdb2xks.ent.gz | 786.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2xks.json.gz 2xks.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xk/2xks https://data.pdbj.org/pub/pdb/validation_reports/xk/2xks ftp://data.pdbj.org/pub/pdb/validation_reports/xk/2xks ftp://data.pdbj.org/pub/pdb/validation_reports/xk/2xks | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2xkuC  2xkt C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11748.160 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PET23A / Production host: HOMO SAPIENS (human) / Plasmid: PET23A / Production host:  |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NMR details | Text: THE STRUCTURE WAS DETERMINED USING TRIPLE-RESONANCE NMR SPECTROSCOPY ON 13C, 15N-LABELED BETA-2-MICROGLOBULIN |
- Sample preparation
Sample preparation
| Details |
| ||||||
|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 0.04 / pH: 7.5 / Pressure: 1.0 atm / Temperature: 298.0 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Software ordinal: 1 Details: THE CCPN NMR ASSIGNMENT AND DATA ANALYSIS APPLICATION. | ||||||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: LOWEST ENERGY / Conformers calculated total number: 30 / Conformers submitted total number: 30 |
 Movie
Movie Controller
Controller
































































































































































 PDBj
PDBj

 HNCA
HNCA