-検索条件
-検索結果
検索 (著者・登録者: vasishtan & d)の結果65件中、1から50件目までを表示しています
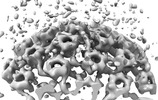
EMDB-18482: 
Herpes simplex virus 1 capsid (WT) vertices in perinuclear NEC-coated vesicles determined in situ

EMDB-18484: 
Herpes simplex virus 1 nuclear egress complex (WT) determined in situ from perinuclear vesicles

EMDB-17974: 
Pseudorabies virus cytosolic C-capsid (US3 KO) vertices determined in situ

EMDB-17975: 
Pseudorabies virus primary enveloped (perinuclear) C-capsid (US3 KO) vertices determined in situ

EMDB-17976: 
Pseudorabies nuclear C-capsids (US3 KO) vertices determined in situ

EMDB-18473: 
Subtomogram average of pseudorabies virus nuclear egress complex helical form (UL31/34) determined in situ

EMDB-18474: 
Subtomogram average of pseudorabies virus nuclear egress complex (UL31/34) determined in situ
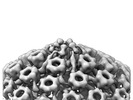

EMDB-18479: 
Pseudorabies virus cytosolic C-capsid (WT) vertices determined in situ

EMDB-18480: 
Pseudorabies virus nuclear C-capsid (WT) vertices determined in situ
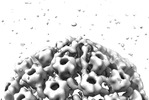

EMDB-18481: 
Herpes simplex virus 1 cytosolic C-capsid (WT) vertices determined in situ
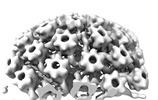
EMDB-18483: 
Herpes simplex virus 1 nuclear C-capsid (WT) vertices determined in situ
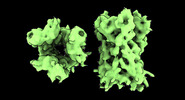
EMDB-17704: 
Subtomogram average of Vaccinia A10 trimer with open center from in vitro cores
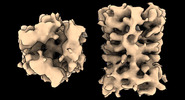
EMDB-17708: 
Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores

EMDB-17753: 
Subtomogram average of Vaccinia A10 trimer from in situ cores

EMDB-15532: 
Plasmodium falciparum sporozoite subpellicular microtubule with interrupted luminal helix determined in situ

EMDB-15534: 
Plasmodium falciparum gametocyte subpellicular microtubule with 13 protofilaments determined in situ

EMDB-15535: 
Plasmodium falciparum gametocyte subpellicular microtubule with 14 protofilaments determined in situ

EMDB-15536: 
Plasmodium falciparum gametocyte subpellicular microtubule with 15 protofilaments determined in situ

EMDB-15537: 
Plasmodium falciparum gametocyte subpellicular microtubule with 16 protofilaments determined in situ

EMDB-15538: 
Plasmodium falciparum gametocyte subpellicular microtubule with 17 protofilaments determined in situ

EMDB-15539: 
Plasmodium falciparum gametocyte subpellicular microtubule with 18 protofilaments determined in situ

EMDB-15627: 
Tomogram of the Mimivirus genomic fiber

EMDB-15628: 
Tomogram of the Mimivirus genomic fiber

EMDB-15629: 
Tomogram of the Mimivirus genomic fiber

EMDB-15630: 
Tomogram of the Mimivirus genomic fiber

EMDB-13641: 
Structure of the Mimivirus genomic fibre asymmetric unit

EMDB-14353: 
Structure of the Mimivirus genomic fibre in its compact 6-start helix form

EMDB-14354: 
Structure of the Mimivirus genomic fibre in its compact 5-start helix form

EMDB-14355: 
Structure of the Mimivirus genomic fibre in its relaxed 5-start helix form

PDB-7ptv: 
Structure of the Mimivirus genomic fibre asymmetric unit

PDB-7yx3: 
Structure of the Mimivirus genomic fibre in its compact 6-start helix form

PDB-7yx4: 
Structure of the Mimivirus genomic fibre in its compact 5-start helix form

PDB-7yx5: 
Structure of the Mimivirus genomic fibre in its relaxed 5-start helix form

EMDB-12188: 
Structure of DNA origami signpost from cryo-ET and subvolume averaging

PDB-7bho: 
DNA origami signpost designed model

EMDB-11123: 
Pre-fusion conformation of glycoprotein B of Herpes simplex virus 1

PDB-6z9m: 
Pseudoatomic model of the pre-fusion conformation of glycoprotein B of Herpes simplex virus 1

EMDB-4995: 
Subtomogram average of a part of the rabies lyssavirus ribonucleoprotein

EMDB-3357: 
electron density map of murine leukaemia virus envelope glycoprotein tagged in the proline rich region with YFP as reconstructed by subtomogram averaging on viruses produced in DFJ8 cells

EMDB-3363: 
electron density map of murine leukaemia virus envelope glycoprotein tagged in the proline rich region with YFP as reconstructed by subtomogram averaging on murine leukemia virus virus like particles (3 plasmid system) produced in COS1 cells

EMDB-3365: 
electron density map of murine leukaemia virus envelope glycoprotein as reconstructed by subtomogram averaging on murine leukemia virus virus like particles (3 plasmid system) produced in COS1 cells

EMDB-3373: 
electron density map of murine leukaemia virus envelope glycoprotein as reconstructed by subtomogram averaging applying a mask on 1 protomer on murine leukemia virus particles and virus like particles and applying 3fold symmetry to the single protomer afterwards

EMDB-4043: 
13-protofilament microtubule structure determined in situ from U87MG cell

EMDB-4045: 
13-protofilament microtubule structure determined in situ from U2OS cell

EMDB-3362: 
Natively membrane-anchored full-length Herpes simplex virus 1 glycoprotein B

PDB-5fki: 
Pseudorabies virus (PrV) nuclear egress complex proteins fitted as a hexameric lattice into a sub-tomogram average derived from focused- ion beam milled lamellae electron cryo-microscopic data

EMDB-3197: 
Sub-tomogram averaging of electron cryo-microscopic data taken from focused-ion beam milled lamellae of nuclei of Pseudorabies virus (PrV) nuclear egress complex-expressing cells

EMDB-3215: 
Sub-tomogram averaging of electron cryo-microscopic data taken from focused-ion beam milled lamellae of nuclei of Pseudorabies virus (PrV) nuclear egress complex-expressing cells

EMDB-2530: 
Tomographic subvolume average of EFF-1 fusogen on extracellular vesicles

EMDB-2531: 
Tomographic subvolume average of EFF-1 fusogen on extracellular vesicles
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
