-Search query
-Search result
Showing all 38 items for (author: stoddard & d)

EMDB-40260: 
CryoEM map of a de novo designed T=4 icosahedral nanocage hierarchically built from pseudosymmetric trimers; design Ico(T=4)-4

EMDB-40267: 
CryoEM map of a T=1 off-target state of design Ico(T=4)-4

EMDB-40268: 
CryoEM map of a de novo designed T=4 octahedral nanocage hierarchically built from pseudosymmetric trimers; design Oct(T=4)-3

EMDB-40269: 
CryoEM map of a T=1 off-target state of design Oct(T=4)-3

EMDB-40208: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15

EMDB-40209: 
Chlorophyll-binding region of de novo-designed nanocage O32-15

PDB-8glt: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15

EMDB-28534: 
Type IIS Restriction Endonuclease PaqCI, DNA bound

PDB-8epx: 
Type IIS Restriction Endonuclease PaqCI, DNA bound

EMDB-28244: 
CryoEM characterization of BrxL -- a unique AAA+ phage restriction Factor.
EMDB-28248: 
CryoEM characterization of a unique AAA+ BrxL phage restriction factor

PDB-8emc: 
CryoEM characterization of BrxL -- a unique AAA+ phage restriction Factor.

PDB-8emh: 
CryoEM characterization of a unique AAA+ BrxL phage restriction factor

EMDB-29052: 
SARS-CoV-2 Spike Hexapro - C68.59 Fab (Class 3 - disordered)

EMDB-29053: 
SARS-CoV-2 Spike Hexapro - C59.68 Fab (Class 1 - No Fab bound)

EMDB-29054: 
SARS-CoV-2 Spike Hexapro - C68.59 Fab (Class 2 - Fab bound)
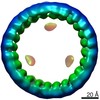
EMDB-24425: 
Circular tandem repeat protein with novel repeat topology and enhanced subunit contact surfaces

PDB-7rdr: 
Circular tandem repeat protein with novel repeat topology and enhanced subunit contact surfaces

EMDB-22712: 
Subtomogram average of flagellar axoneme doublet from the wild type Tetrahymena thermophila

EMDB-22713: 
Subtomogram average of flagellar axoneme doublet from the Tetrahymena thermophila Fap115 knockout mutant

EMDB-23461: 
cryoEM structure DrdV-DNA complex

EMDB-23543: 
cryoEM structure DrdV-DNA complex

PDB-7lo5: 
cryoEM structure DrdV-DNA complex

PDB-7lvv: 
cryoEM structure DrdV-DNA complex

EMDB-21955: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Upright conformation

EMDB-21956: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Intermediate conformation

EMDB-21957: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Reversed conformation

PDB-6wxe: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Upright conformation

PDB-6wxf: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Intermediate conformation

PDB-6wxg: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Reversed conformation

EMDB-20309: 
cryoEM structure of yeast glucokinase filament

PDB-6pdt: 
cryoEM structure of yeast glucokinase filament

EMDB-9127: 
A nucleosome bridging mechanism for activation of a maintenance DNA methyltransferase

EMDB-7805: 
Subtomogram average (699 axonemal repeats) of isolated Rib72A/B double knock out Tetrahymena cilia showing the lumen of the ciliary doublet microtubule (DMT)

EMDB-7806: 
Subtomogram average (758 axonemal repeats) of isolated Rib72A knock out Tetrahymena cilia showing the lumen of the ciliary doublet microtubule (DMT)

EMDB-7807: 
Subtomogram average (735 axonemal repeats) of isolated Rib72B knock out Tetrahymena cilia Rescued with Rib72B-GFP, showing the lumen of the ciliary doublet microtubule (DMT)

EMDB-7811: 
Subtomogram average (702 axonemal repeats) of isolated Rib72B knock out Tetrahymena cilia showing the lumen of the ciliary doublet microtubule (DMT)

EMDB-8696: 
Regulation of Rvb1/Rvb2 by a domain within the INO80 chromatin remodeling complex implicates the yeast Rvbs as protein assembly chaperones
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
