-Search query
-Search result
Showing all 43 items for (author: nannenga & b)

EMDB-44778: 
C-terminus truncated (last two residues) mutant of Human light chain ferritin reacted with Ferrous salt(3 Fe2+ per ferritin subunit) . Reconstruction of particles with one nanoparticle.
Method: single particle / : Sen S, Nannenga BL, Williams D

EMDB-44779: 
Human light chain ferritin reacted with iron (3 Fe2+ to ferritin monomer ratio). Reconstruction of particles with one nanoparticle.
Method: single particle / : Sen S, Nannenga BL, Williams D

EMDB-44780: 
Human light chain ferritin reacted with iron (3 Fe2+ to ferritin monomer ratio). Reconstruction of particles with no nanoparticle.
Method: single particle / : Sen S, Nannenga BL, Williams D

EMDB-44797: 
C-terminus truncated (last two residues) mutant of Human light chain ferritin reacted with iron (3 Fe2+ to ferritin monomer ratio). Reconstruction of particles with no nanoparticle.
Method: single particle / : Sen S, Nannenga BL, Williams D

PDB-9bpi: 
C-terminus truncated (last two residues) mutant of Human light chain ferritin reacted with Ferrous salt(3 Fe2+ per ferritin subunit) . Reconstruction of particles with one nanoparticle
Method: single particle / : Sen S, Nannenga BL, Williams D

PDB-9bpj: 
Human light chain ferritin reacted with iron (3 Fe2+ to ferritin monomer ratio). Reconstruction of particles with one nanoparticle
Method: single particle / : Sen S, Nannenga BL, Williams D

PDB-9bpk: 
Human light chain ferritin reacted with iron (3 Fe2+ to ferritin monomer ratio). Reconstruction of particles with no nanoparticle
Method: single particle / : Sen S, Nannenga BL, Williams D

PDB-9bq5: 
C-terminus truncated (last two residues) mutant of Human light chain ferritin reacted with iron (3 Fe2+ to ferritin monomer ratio). Reconstruction of particles with no nanoparticle.
Method: single particle / : Sen S, Nannenga BL, Williams D

EMDB-44453: 
MicroED structure of bovine liver catalase with missing cone solved by suspended drop
Method: electron crystallography / : Gillman C, Bu G, Gonen T

PDB-9bdj: 
MicroED structure of bovine liver catalase with missing cone eliminated by suspended drop
Method: electron crystallography / : Gillman C, Bu G, Gonen T

EMDB-43144: 
MicroED structure of SARS-CoV-2 main protease (MPro/3CLPro) with missing cone eliminated by suspended drop
Method: electron crystallography / : Bu G, Gillman C, Danelius E, Hattne J, Nannenga BL, Gonen T

PDB-8vd7: 
MicroED structure of SARS-CoV-2 main protease (MPro/3CLPro) with missing cone eliminated by suspended drop
Method: electron crystallography / : Bu G, Gillman C, Danelius E, Hattne J, Nannenga BL, Gonen T

PDB-8skw: 
MicroED structure of d(CGCGCG)2 Z-DNA
Method: electron crystallography / : Haymaker A, Bardin AA, Martynowycz MW, Nannenga BL

EMDB-26469: 
Photosynthetic assembly of Chlorobaculum tepidum (RC-FMO1)
Method: single particle / : Puskar R, Truong CD, Swain K, Li S, Cheng KW, Wang TY, Poh YP, Liu H, Chou TF, Nannenga B, Chiu PL

EMDB-26471: 
Photosynthetic assembly of Chlorobaculum tepidum (RC-FMO2)
Method: single particle / : Puskar R, Truong CD, Swain K, Li S, Cheng KW, Wang TY, Poh YP, Liu H, Chou TF, Nannenga B, Chiu PL

PDB-7uea: 
Photosynthetic assembly of Chlorobaculum tepidum (RC-FMO1)
Method: single particle / : Puskar R, Truong CD, Swain K, Li S, Cheng KW, Wang TY, Poh YP, Liu H, Chou TF, Nannenga B, Chiu PL

PDB-7ueb: 
Photosynthetic assembly of Chlorobaculum tepidum (RC-FMO2)
Method: single particle / : Puskar R, Truong CD, Swain K, Li S, Cheng KW, Wang TY, Poh YP, Liu H, Chou TF, Nannenga B, Chiu PL

EMDB-27900: 
MicroED structure of proteinase K recorded on K2
Method: electron crystallography / : Clabbers MTB, Martynowycz MW, Hattne J, Nannenga BL, Gonen T

EMDB-27901: 
MicroED structure of proteinase K recorded on K3
Method: electron crystallography / : Clabbers MTB, Martynowycz MW, Hattne J, Nannenga BL, Gonen T

EMDB-27902: 
MicroED structure of triclinic lysozyme recorded on K3
Method: electron crystallography / : Clabbers MTB, Martynowycz MW, Hattne J, Nannenga BL, Gonen T

PDB-8e52: 
MicroED structure of proteinase K recorded on K2
Method: electron crystallography / : Clabbers MTB, Martynowycz MW, Hattne J, Nannenga BL, Gonen T

PDB-8e53: 
MicroED structure of proteinase K recorded on K3
Method: electron crystallography / : Clabbers MTB, Martynowycz MW, Hattne J, Nannenga BL, Gonen T

PDB-8e54: 
MicroED structure of triclinic lysozyme recorded on K3
Method: electron crystallography / : Clabbers MTB, Martynowycz MW, Hattne J, Nannenga BL, Gonen T

EMDB-24551: 
MicroED structure of the human adenosine receptor at 2.8A
Method: electron crystallography / : Martynowycz MW, Shiriaeva A

PDB-7rm5: 
MicroED structure of the human adenosine receptor at 2.8A
Method: electron crystallography / : Martynowycz MW, Shiriaeva A, Ge X, Hattne J, Nannenga BL, Cherezov V, Gonen T

PDB-6pq0: 
LCP-embedded Proteinase K treated with MPD
Method: electron crystallography / : Bu G, Zhu L, Jing L, Shi D, Gonen T, Liu W, Nannenga BL

PDB-6pq4: 
LCP-embedded Proteinase K treated with lipase
Method: electron crystallography / : Bu G, Zhu L, Jing L, Shi D, Gonen T, Liu W, Nannenga BL
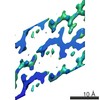
EMDB-8272: 
Human Islet Amyloid Polypeptide Segment 19-SGNNFGAILSS-29 with Early Onset S20G Mutation Determined by MicroED
Method: electron crystallography / : Krotee PAL, Rodriguez JA

EMDB-8273: 
Human Islet Amyloid Polypeptide Segment 15-FLVHSSNNFGA-25 Determined by MicroED
Method: electron crystallography / : Krotee PAL, Rodriguez JA

PDB-5knz: 
Human Islet Amyloid Polypeptide Segment 19-SGNNFGAILSS-29 with Early Onset S20G Mutation Determined by MicroED
Method: electron crystallography / : Krotee PAL, Rodriguez JA, Sawaya MR, Cascio D, Shi D, Nannenga BL, Hattne J, Reyes FE, Gonen T, Eisenberg DS

PDB-5ko0: 
Human Islet Amyloid Polypeptide Segment 15-FLVHSSNNFGA-25 Determined by MicroED
Method: electron crystallography / : Krotee PAL, Rodriguez JA, Sawaya MR, Cascio D, Shi D, Nannenga BL, Hattne J, Reyes FE, Gonen T, Eisenberg DS

EMDB-3001: 
MicroED structure of the segment, GVVHGVTTVA, from the A53T familial mutant of Parkinson's disease protein, alpha-synuclein, residues 47-56
Method: electron crystallography / : Rodriguez JA, Ivanova M, Sawaya MR, Cascio D, Reyes F, Shi D, Johnson L, Guenther E, Sangwan S, Hattne J, Nannenga B, Gonen T, Eisenberg D
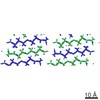
EMDB-3028: 
MicroED structure of the toxic core segment, GAVVTGVTAVA, from Parkinson's disease protein, alpha-synuclein, residues 69-78.
Method: electron crystallography / : Rodriguez JA, Ivanova M, Sawaya MR, Cascio D, Reyes F, Shi D, Johnson L, Guenther E, Zhang M, Jiang L, Arbing MA, Sangwan S, Hattne J, Whitelegge J, Brewster A, Messerschmidt M, Boutet S, Sauter NK, Nannenga B, Gonen T, Eisenberg D

PDB-4znn: 
MicroED structure of the segment, GVVHGVTTVA, from the A53T familial mutant of Parkinson's disease protein, alpha-synuclein residues 47-56
Method: electron crystallography / : Rodriguez JA, Ivanova M, Sawaya MR, Cascio D, Reyes F, Shi D, Johnson L, Guenther E, Sangwan S, Hattne J, Nannenga B, Brewster AS, Messerschmidt M, Boutet S, Sauter NK, Gonen T, Eisenberg DS

PDB-4ril: 
Structure of the amyloid forming segment, GAVVTGVTAVA, from the NAC domain of Parkinson's disease protein alpha-synuclein, residues 68-78, determined by electron diffraction
Method: electron crystallography / : Rodriguez JA, Ivanova M, Sawaya MR, Cascio D, Reyes F, Shi D, Johnson L, Guenther E, Sangwan S, Hattne J, Nannenga B, Brewster AS, Messerschmidt M, Boutet S, Sauter NK, Gonen T, Eisenberg DS

PDB-5a3e: 
2.5A structure of lysozyme determined by MicroED with data from a single crystal
Method: electron crystallography / : Nannenga BL, Shi D, Leslie AGW, Gonen T

EMDB-6342: 
2.5A structure of lysozyme determined by MicroED with data from a single crystal
Method: electron crystallography / : Nannenga BL, Shi D, Leslie AGW, Gonen T

EMDB-6314: 
Catalase solved at 3.2 Angstrom resolution by MicroED
Method: electron crystallography / : Nannenga BL, Shi D, Hattne J, Reyes FE, Gonen T

EMDB-6313: 
2.5A structure of lysozyme solved by MicroED
Method: electron crystallography / : Nannenga BL, Shi D, Leslie AGW, Gonen T

EMDB-2945: 
Structure of lysozyme solved by MicroED to 2.9 A
Method: electron crystallography / : Shi D, Nannenga BL, Iadanza MG, Gonen T

PDB-3j7b: 
Catalase solved at 3.2 Angstrom resolution by MicroED
Method: electron crystallography / : Nannenga BL, Shi D, Hattne J, Reyes FE, Gonen T

PDB-3j6k: 
2.5A structure of lysozyme solved by MicroED
Method: electron crystallography / : Nannenga BL, Shi D, Leslie AGW, Gonen T

PDB-3j4g: 
Structure of lysozyme solved by MicroED to 2.9 A
Method: electron crystallography / : Shi D, Nannenga BL, Iadanza MG, Gonen T
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
