[English] 日本語
 Yorodumi
Yorodumi- PDB-4znn: MicroED structure of the segment, GVVHGVTTVA, from the A53T famil... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4znn | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | MicroED structure of the segment, GVVHGVTTVA, from the A53T familial mutant of Parkinson's disease protein, alpha-synuclein residues 47-56 | |||||||||
 Components Components | Alpha-synuclein | |||||||||
 Keywords Keywords | LIPID BINDING PROTEIN / Amyloid / alpha-synuclein / Parkinson's Disease / Toxic Core / NACore | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / response to desipramine / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / regulation of synaptic vesicle recycling ...negative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / response to desipramine / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / regulation of synaptic vesicle recycling / negative regulation of chaperone-mediated autophagy / regulation of reactive oxygen species biosynthetic process / positive regulation of protein localization to cell periphery / mitochondrial membrane organization / negative regulation of exocytosis / regulation of glutamate secretion / dopamine biosynthetic process / response to iron(II) ion / positive regulation of neurotransmitter secretion / regulation of macrophage activation / negative regulation of dopamine metabolic process / negative regulation of platelet-derived growth factor receptor signaling pathway / SNARE complex assembly / negative regulation of thrombin-activated receptor signaling pathway / Lewy body / regulation of locomotion / negative regulation of microtubule polymerization / positive regulation of inositol phosphate biosynthetic process / synaptic vesicle priming / regulation of norepinephrine uptake / transporter regulator activity / protein kinase inhibitor activity / synaptic vesicle transport / dopamine uptake involved in synaptic transmission / regulation of dopamine secretion / positive regulation of receptor recycling / positive regulation of exocytosis / cuprous ion binding / nuclear outer membrane / mitochondrial ATP synthesis coupled electron transport / dynein complex binding / synaptic vesicle exocytosis / response to magnesium ion / positive regulation of endocytosis / negative regulation of serotonin uptake / response to type II interferon / cysteine-type endopeptidase inhibitor activity / kinesin binding / regulation of presynapse assembly / synaptic vesicle endocytosis / alpha-tubulin binding / beta-tubulin binding / phospholipase binding / phospholipid metabolic process / supramolecular fiber organization / behavioral response to cocaine / cellular response to fibroblast growth factor stimulus / cellular response to epinephrine stimulus / inclusion body / Hsp70 protein binding / enzyme inhibitor activity / response to interleukin-1 / axon terminus / cellular response to copper ion / positive regulation of release of sequestered calcium ion into cytosol / regulation of microtubule cytoskeleton organization / SNARE binding / adult locomotory behavior / glutathione metabolic process / protein tetramerization / protein sequestering activity / tubulin binding / phosphoprotein binding / excitatory postsynaptic potential / microglial cell activation / ferrous iron binding / fatty acid metabolic process / PKR-mediated signaling / phospholipid binding / synapse organization / receptor internalization / regulation of long-term neuronal synaptic plasticity / protein destabilization / tau protein binding / enzyme activator activity / terminal bouton / positive regulation of inflammatory response / long-term synaptic potentiation / synaptic vesicle membrane / actin cytoskeleton / growth cone / actin binding / cellular response to oxidative stress / neuron apoptotic process / cell cortex / histone binding / response to lipopolysaccharide / microtubule binding / amyloid fibril formation / chemical synaptic transmission Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | ELECTRON CRYSTALLOGRAPHY / electron crystallography /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / cryo EM / Resolution: 1.41 Å molecular replacement / cryo EM / Resolution: 1.41 Å | |||||||||
 Authors Authors | Rodriguez, J.A. / Ivanova, M. / Sawaya, M.R. / Cascio, D. / Reyes, F. / Shi, D. / Johnson, L. / Guenther, E. / Sangwan, S. / Hattne, J. ...Rodriguez, J.A. / Ivanova, M. / Sawaya, M.R. / Cascio, D. / Reyes, F. / Shi, D. / Johnson, L. / Guenther, E. / Sangwan, S. / Hattne, J. / Nannenga, B. / Brewster, A.S. / Messerschmidt, M. / Boutet, S. / Sauter, N.K. / Gonen, T. / Eisenberg, D.S. | |||||||||
| Funding support |  United States, 2items United States, 2items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2015 Journal: Nature / Year: 2015Title: Structure of the toxic core of α-synuclein from invisible crystals. Authors: Jose A Rodriguez / Magdalena I Ivanova / Michael R Sawaya / Duilio Cascio / Francis E Reyes / Dan Shi / Smriti Sangwan / Elizabeth L Guenther / Lisa M Johnson / Meng Zhang / Lin Jiang / Mark ...Authors: Jose A Rodriguez / Magdalena I Ivanova / Michael R Sawaya / Duilio Cascio / Francis E Reyes / Dan Shi / Smriti Sangwan / Elizabeth L Guenther / Lisa M Johnson / Meng Zhang / Lin Jiang / Mark A Arbing / Brent L Nannenga / Johan Hattne / Julian Whitelegge / Aaron S Brewster / Marc Messerschmidt / Sébastien Boutet / Nicholas K Sauter / Tamir Gonen / David S Eisenberg /  Abstract: The protein α-synuclein is the main component of Lewy bodies, the neuron-associated aggregates seen in Parkinson disease and other neurodegenerative pathologies. An 11-residue segment, which we term ...The protein α-synuclein is the main component of Lewy bodies, the neuron-associated aggregates seen in Parkinson disease and other neurodegenerative pathologies. An 11-residue segment, which we term NACore, appears to be responsible for amyloid formation and cytotoxicity of human α-synuclein. Here we describe crystals of NACore that have dimensions smaller than the wavelength of visible light and thus are invisible by optical microscopy. As the crystals are thousands of times too small for structure determination by synchrotron X-ray diffraction, we use micro-electron diffraction to determine the structure at atomic resolution. The 1.4 Å resolution structure demonstrates that this method can determine previously unknown protein structures and here yields, to our knowledge, the highest resolution achieved by any cryo-electron microscopy method to date. The structure exhibits protofibrils built of pairs of face-to-face β-sheets. X-ray fibre diffraction patterns show the similarity of NACore to toxic fibrils of full-length α-synuclein. The NACore structure, together with that of a second segment, inspires a model for most of the ordered portion of the toxic, full-length α-synuclein fibril, presenting opportunities for the design of inhibitors of α-synuclein fibrils. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4znn.cif.gz 4znn.cif.gz | 12.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4znn.ent.gz pdb4znn.ent.gz | 4.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4znn.json.gz 4znn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zn/4znn https://data.pdbj.org/pub/pdb/validation_reports/zn/4znn ftp://data.pdbj.org/pub/pdb/validation_reports/zn/4znn ftp://data.pdbj.org/pub/pdb/validation_reports/zn/4znn | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3001MC 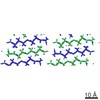 3028C  4rikC  4rilC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 940.074 Da / Num. of mol.: 1 / Mutation: A53T / Source method: obtained synthetically Details: Synthetic peptide GVVHGVTTVA corresponding to segment 47-56 of human alpha-synuclein Source: (synth.)  Homo sapiens (human) / References: UniProt: P37840 Homo sapiens (human) / References: UniProt: P37840 |
|---|---|
| #2: Water | ChemComp-HOH / |
| Has protein modification | N |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON CRYSTALLOGRAPHY / Number of used crystals: 1 |
|---|---|
| EM experiment | Aggregation state: 3D ARRAY / 3D reconstruction method: electron crystallography |
- Sample preparation
Sample preparation
| Component | Name: Human alpha-synuclein / Type: COMPLEX / Entity ID: #1 / Source: MULTIPLE SOURCES |
|---|---|
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES / Details: crystal |
| Crystal grow | Temperature: 310 K / Method: batch mode / pH: 7 Details: 1 mg of synthetic peptide GVVHGVTTVA was dissolved in 200 microliters of 50 mM phosphate buffer pH 7.0 and 0.1% w/v DMSO and shaken overnight in an orbital mixing plate PH range: 7 |
-Data collection
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Microscopy | Model: FEI TECNAI F20 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Electron lens | Mode: DIFFRACTION | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Specimen holder | Cryogen: NITROGEN | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diffraction source | Source: ELECTRON MICROSCOPE / Type: OTHER / Wavelength: 0.0251 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Detector: CMOS / Date: Apr 20, 2015 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: electron | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.0251 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.41→16.5 Å / Num. obs: 1120 / % possible obs: 86.9 % / Observed criterion σ(I): -3 / Redundancy: 3.7 % / Biso Wilson estimate: 22.483 Å2 / Rmerge F obs: 0.967 / Rmerge(I) obs: 0.236 / Rrim(I) all: 0.264 / Χ2: 0.831 / Net I/σ(I): 4.62 / Num. measured all: 4110 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1 / Rejects: _
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement |
|---|---|
| Phasing MR | Model details: Phaser MODE: MR_AUTO |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: Ideal model Resolution: 1.41→16.47 Å / Cor.coef. Fo:Fc: 0.953 / Cor.coef. Fo:Fc free: 0.958 / WRfactor Rfree: 0.2879 / WRfactor Rwork: 0.2427 / FOM work R set: 0.6203 / SU B: 3.491 / SU ML: 0.111 / SU R Cruickshank DPI: 0.1115 / SU Rfree: 0.1137 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.112 / ESU R Free: 0.114 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 50.34 Å2 / Biso mean: 18.371 Å2 / Biso min: 11.23 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.41→16.47 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.409→1.574 Å / Total num. of bins used: 5
|
 Movie
Movie Controller
Controller











 PDBj
PDBj

