-Search query
-Search result
Showing 1 - 50 of 186 items for (author: miki & k)

EMDB-62427: 
Cryo-EM structure of the heterotrimeric interleukin-2 receptor in complex with interleukin-2 and anti-CD25 Fab S417
Method: single particle / : Katsura K, Matsumoto T, Shirouzu M

PDB-9kmc: 
Cryo-EM structure of the heterotrimeric interleukin-2 receptor in complex with interleukin-2 and anti-CD25 Fab S417
Method: single particle / : Katsura K, Matsumoto T, Shirouzu M

EMDB-61993: 
Structure of the Salmonella flagellar FliPQR complex reconstituted in the peptidisc
Method: single particle / : Kinoshita M, Miyata T, Makino F, Imada K, Namba K, Minamino T

PDB-9k29: 
Structure of the Salmonella flagellar FliPQR complex reconstituted in the peptidisc
Method: single particle / : Kinoshita M, Miyata T, Makino F, Imada K, Namba K, Minamino T

EMDB-62659: 
Cryo-EM structure of the LGI1 LRR-LGI1 EPTP-ADAM22 ECD complex
Method: single particle / : Yamaguchi T, Okatsu K, Kubota M, Mitsumori A, Yamagata A, Fukai S

EMDB-62668: 
Cryo-EM structure of the 3:3 LGI1-ADAM22 complex
Method: single particle / : Yamaguchi T, Okatsu K, Kubota M, Mistumori A, Yamagata A, Fukai S

PDB-9kzc: 
Cryo-EM structure of the LGI1 LRR-LGI1 EPTP-ADAM22 ECD complex
Method: single particle / : Yamaguchi T, Okatsu K, Kubota M, Mitsumori A, Yamagata A, Fukai S

PDB-9kzt: 
Cryo-EM structure of the 3:3 LGI1-ADAM22 complex
Method: single particle / : Yamaguchi T, Okatsu K, Kubota M, Mistumori A, Yamagata A, Fukai S

EMDB-61174: 
HBx fused DDB1 4M mutant
Method: single particle / : Tanaka H, Kita S, Sasaki M, Maenaka K, Machida S

EMDB-61175: 
HBx complexed with DDB1
Method: single particle / : Tanaka H, Kita S, Sasaki M, Maenaka K, Machida S

EMDB-63366: 
HBx R96E fused DDB1 4M mutant
Method: single particle / : Tanaka H, Kita S, Sasaki M, Maenaka K, Machida S

PDB-9j6j: 
HBx fused DDB1 4M mutant
Method: single particle / : Tanaka H, Kita S, Sasaki M, Maenaka K, Machida S

PDB-9j6k: 
HBx complexed with DDB1
Method: single particle / : Tanaka H, Kita S, Sasaki M, Maenaka K, Machida S

EMDB-63392: 
The chimeric flagellar motor complex between MotA1B1 from Paenibacillus sp. TCA20 and MotAB from E.coli, state 1
Method: single particle / : Onoe S, Nishikino T, Kinoshita M, Kishikawa J, Kato T

EMDB-63393: 
The chimeric flagellar motor complex between MotA1B1 from Paenibacillus sp. TCA20 and MotAB from E.coli, state 2
Method: single particle / : Onoe S, Nishikino T, Kinoshita M, Kishikawa J, Kato T

EMDB-63394: 
The chimeric flagellar motor complex between MotA1B1 from Paenibacillus sp. TCA20 and MotAB from E.coli, state 3
Method: single particle / : Onoe S, Nishikino T, Kinoshita M, Kishikawa J, Kato T

PDB-9lu9: 
The chimeric flagellar motor complex between MotA1B1 from Paenibacillus sp. TCA20 and MotAB from E.coli, state 1
Method: single particle / : Onoe S, Nishikino T, Kinoshita M, Kishikawa J, Kato T

PDB-9lub: 
The chimeric flagellar motor complex between MotA1B1 from Paenibacillus sp. TCA20 and MotAB from E.coli, state 2
Method: single particle / : Onoe S, Nishikino T, Kinoshita M, Kishikawa J, Kato T

PDB-9luc: 
The chimeric flagellar motor complex between MotA1B1 from Paenibacillus sp. TCA20 and MotAB from E.coli, state 3
Method: single particle / : Onoe S, Nishikino T, Kinoshita M, Kishikawa J, Kato T

EMDB-60007: 
Structure of the Salmonella flagellar MS-ring with C11 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-60008: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C33 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-60009: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C34 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-62210: 
Structure of the 34-mer RBM3 ring of Salmonella flagellar MS-ring protein FliF with C1 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-62211: 
Structure of the 33-mer RBM3 ring of Salmonella flagellar MS-ring protein FliF with C1 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

PDB-8zds: 
Structure of the Salmonella flagellar MS-ring with C11 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K
PDB-8zdt: 
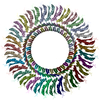
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C33 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K
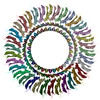
PDB-8zdu: 
Structure of the RBM3 ring of Salmonella flagellar MS-ring protein FliF with C34 symmetry applied
Method: single particle / : Kinoshita M, Makino F, Miyata T, Imada K, Minamino T, Namba K

EMDB-60274: 
SARS-CoV-2 XBB.1.5 spike glycoprotein trimer in complex with antigen-binding fragments (Fabs)
Method: single particle / : Sugita Y, Kimura K, Noda T, Hashiguchi T

EMDB-39761: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 34-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-39763: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 35-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-39764: 
Homomeric 34mer of the Vibrio flagellar MS-ring protein FliF without symmetry imposition
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-39765: 
Homomeric 35mer of the Vibrio flagellar MS-ring protein FliF without symmetry imposition
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

PDB-8z4d: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 34-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M
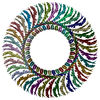
PDB-8z4g: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 35-fold symmetry applied
Method: single particle / : Takekawa N, Nishikino T, Kishikawa J, Hirose M, Kato T, Imada K, Homma M

EMDB-19599: 
Structural characterization of Thogoto Virus nucleoprotein provides insights into RNA encapsidation and assembly
Method: subtomogram averaging / : Roske Y, Mikirtumov V, Daumke O, Kudryashev M, Dick A

PDB-8ryt: 
Structural characterization of Thogoto Virus nucleoprotein provides insights into RNA encapsidation and assembly
Method: subtomogram averaging / : Roske Y, Mikirtumov V, Daumke O, Kudryashev M, Dick A

EMDB-38453: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

EMDB-38454: 
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

PDB-8xlm: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

PDB-8xln: 
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

EMDB-37648: 
SARS-CoV-2 EG.5.1 spike glycoprotein (1-up state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

EMDB-37650: 
SARS-CoV-2 EG.5.1 spike glycoprotein (closed-2 state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

EMDB-37651: 
SARS-CoV-2 EG.5.1 spike glycoprotein (closed-1 state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

PDB-8wmd: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein (closed-2 state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K

PDB-8wmf: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein (closed-1 state)
Method: single particle / : Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K
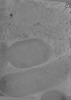
EMDB-18953: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells
Method: electron tomography / : Feldmueller M, Pilhofer M

EMDB-18954: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells
Method: electron tomography / : Feldmueller M, Pilhofer M
EMDB-18955: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells
Method: electron tomography / : Feldmueller M, Pilhofer M

EMDB-18957: 
Cryo-electron tomogram of Yersinia entomophaga MH96 cells
Method: electron tomography / : Feldmueller M, Pilhofer M
EMDB-18958: 
Cryo-electron tomogram of Yersinia entomophaga MH96 cells
Method: electron tomography / : Feldmueller M, Pilhofer M
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
