8WC6
 
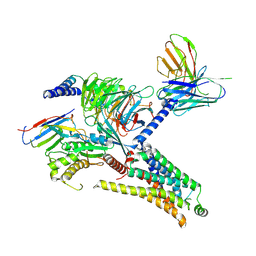 | | Cryo-EM structure of the PEA-bound mTAAR1-Gs complex | | Descriptor: | 2-PHENYLETHYLAMINE, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Rong, N.K, Guo, L.L, Zhang, M.H, Li, Q, Yang, F, Sun, J.P. | | Deposit date: | 2023-09-11 | | Release date: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structural and signaling mechanisms of TAAR1 enabled preferential agonist design.
Cell, 186, 2023
|
|
8WC5
 
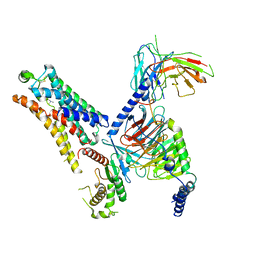 | | Cryo-EM structure of the TMA-bound mTAAR1-Gs complex | | Descriptor: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Guanine nucleotide-binding protein G(s) subunit alpha isoforms short, ... | | Authors: | Rong, N.K, Guo, L.L, Zhang, M.H, Li, Q, Yang, F, Sun, J.P. | | Deposit date: | 2023-09-11 | | Release date: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structural and signaling mechanisms of TAAR1 enabled preferential agonist design.
Cell, 186, 2023
|
|
8WCC
 
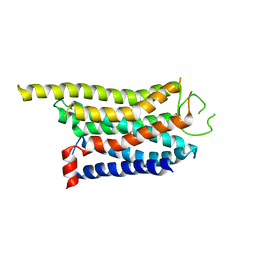 | | Cryo-EM structure of the CHA-bound mTAAR1 complex | | Descriptor: | CYCLOHEXYLAMMONIUM ION, Trace amine-associated receptor 1 | | Authors: | Rong, N.K, Guo, L.L, Zhang, M.H, Li, Q, Yang, F, Sun, J.P. | | Deposit date: | 2023-09-11 | | Release date: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (3.04 Å) | | Cite: | Structural and signaling mechanisms of TAAR1 enabled preferential agonist design.
Cell, 186, 2023
|
|
8XGY
 
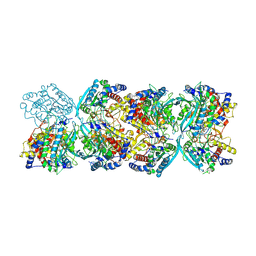 | |
8XGT
 
 | |
8XGB
 
 | |
8XGA
 
 | |
8XFV
 
 | |
6RZO
 
 | |
6S1S
 | | Crystal structure of AmpC from Pseudomonas aeruginosa in complex with [3-(2-carboxyvinyl)phenyl]boronic acid] inhibitor | | Descriptor: | (~{E})-3-[3-(dihydroxyboranyl)phenyl]prop-2-enoic acid, Beta-lactamase, PHOSPHATE ION | | Authors: | Kekez, I, Vicario, M, Bellio, P, Tosoni, E, Celenza, G, Blazquez, J, Tondi, D, Cendron, L. | | Deposit date: | 2019-06-19 | | Release date: | 2019-10-09 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Phenylboronic Acids Probing Molecular Recognition against Class A and Class C beta-lactamases.
Antibiotics, 8, 2019
|
|
6J6Y
 
 | | FGFR4 D2 - Fab complex | | Descriptor: | Fab Heavy chain, Fab light chain, Fibroblast growth factor receptor 4 | | Authors: | Takahashi, M, Hanzawa, H. | | Deposit date: | 2019-01-16 | | Release date: | 2019-08-07 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Preclinical Development of U3-1784, a Novel FGFR4 Antibody Against Cancer, and Avoidance of Its On-target Toxicity.
Mol.Cancer Ther., 18, 2019
|
|
5TW4
 
 | |
5TXI
 
 | | Crystal structure of wild-type S. aureus penicillin binding protein 4 (PBP4) in complex with ceftobiprole | | Descriptor: | (2R)-2-[(1R)-1-{[(2Z)-2-(5-amino-1,2,4-thiadiazol-3-yl)-2-(hydroxyimino)acetyl]amino}-2-oxoethyl]-5-({2-oxo-1-[(3R)-pyr rolidin-3-yl]-2,5-dihydro-1H-pyrrol-3-yl}methyl)-3,6-dihydro-2H-1,3-thiazine-4-carboxylic acid, CHLORIDE ION, GLYCEROL, ... | | Authors: | Alexander, J.A.N, Strynadka, N.C.J. | | Deposit date: | 2016-11-16 | | Release date: | 2018-05-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and kinetic analysis of penicillin-binding protein 4 (PBP4)-mediated antibiotic resistance inStaphylococcus aureus.
J. Biol. Chem., 2018
|
|
5TZB
 | | Burkholderia sp. beta-aminopeptidase | | Descriptor: | CALCIUM ION, D-aminopeptidase | | Authors: | McGowan, S, Drinkwater, N, John, M, Dumsday, G. | | Deposit date: | 2016-11-21 | | Release date: | 2017-07-12 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.977 Å) | | Cite: | Crystal structure of a beta-aminopeptidase from an Australian Burkholderia sp.
Acta Crystallogr F Struct Biol Commun, 73, 2017
|
|
5TY2
 
 | | Crystal structure of S. aureus penicillin binding protein 4 (PBP4) mutant (E183A, F241R) in complex with nafcillin | | Descriptor: | (2R,4S)-2-[(1R)-1-{[(2-ethoxynaphthalen-1-yl)carbonyl]amino}-2-oxoethyl]-5,5-dimethyl-1,3-thiazolidine-4-carboxylic acid, CHLORIDE ION, Penicillin-binding protein 4, ... | | Authors: | Alexander, J.A.N, Strynadka, N.C.J. | | Deposit date: | 2016-11-18 | | Release date: | 2018-06-13 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and kinetic analysis of penicillin-binding protein 4 (PBP4)-mediated antibiotic resistance inStaphylococcus aureus.
J. Biol. Chem., 2018
|
|
5TC4
 
 | | Crystal structure of human mitochondrial methylenetetrahydrofolate dehydrogenase-cyclohydrolase (MTHFD2) in complex with LY345899 and cofactors | | Descriptor: | 4-(7-AMINO-9-HYDROXY-1-OXO-3,3A,4,5-TETRAHYDRO-2,5,6,8,9B-PENTAAZA-CYCLOPENTA[A]NAPHTHALEN-2-YL)-PHENYLCARBONYL-GLUTAMI C ACID, Bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase, mitochondrial, ... | | Authors: | Gustafsson, R, Jemth, A.-S, Gustafsson Sheppard, N, Farnegardh, K, Loseva, O, Wiita, E, Bonagas, N, Dahllund, L, Llona-Minguez, S, Haggblad, M, Henriksson, M, Andersson, Y, Homan, E, Helleday, T, Stenmark, P. | | Deposit date: | 2016-09-14 | | Release date: | 2016-12-14 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Crystal Structure of the Emerging Cancer Target MTHFD2 in Complex with a Substrate-Based Inhibitor.
Cancer Res., 77, 2017
|
|
5URF
 
 | | The structure of human bocavirus 1 | | Descriptor: | viral protein 3 | | Authors: | Mietzsch, M, Kailasan, S, Garrison, J, Ilyas, M, Chipman, P, Kandola, K, Jansen, M, Spear, J, Sousa, D, McKenna, R, Soderlund-Venermo, M, Baker, T, Agbandje-McKenna, M. | | Deposit date: | 2017-02-10 | | Release date: | 2017-03-29 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural Insights into Human Bocaparvoviruses.
J. Virol., 91, 2017
|
|
5VAA
 
 | | Crystal structure of mouse IgG2a Fc T370K mutant | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLYCEROL, Ig gamma-2A chain C region, ... | | Authors: | Armstrong, A.A, Gilliland, G.L. | | Deposit date: | 2017-03-24 | | Release date: | 2017-06-07 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Efficient Generation of Bispecific Murine Antibodies for Pre-Clinical Investigations in Syngeneic Rodent Models.
Sci Rep, 7, 2017
|
|
3G1M
 
 | | EthR from Mycobacterium tuberculosis in complex with compound BDM31381 | | Descriptor: | 1-(thiophen-2-ylacetyl)-4-(3-thiophen-2-yl-1,2,4-oxadiazol-5-yl)piperidine, TRANSCRIPTIONAL REGULATORY REPRESSOR PROTEIN (TETR-FAMILY) ETHR | | Authors: | Willand, N, Dirie, B, Carette, X, Bifani, P, Singhal, A, Desroses, M, Leroux, F, Willery, E, Mathys, V, Deprez-Poulain, R, Delcroix, G, Frenois, F, Aumercier, M, Locht, C, Villeret, V, Deprez, B, Baulard, A.R. | | Deposit date: | 2009-01-30 | | Release date: | 2009-07-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Synthetic EthR inhibitors boost antituberculous activity of ethionamide.
Nat.Med. (N.Y.), 15, 2009
|
|
5TW8
 | |
3G1O
 
 | | EthR from Mycobacterium tuberculosis in complex with compound BDM14500 | | Descriptor: | TRANSCRIPTIONAL REGULATORY REPRESSOR PROTEIN (TETR-FAMILY) ETHR, tert-butyl 4-(3-thiophen-2-yl-1,2,4-oxadiazol-5-yl)piperidine-1-carboxylate | | Authors: | Willand, N, Dirie, B, Carette, X, Bifani, P, Singhal, A, Desroses, M, Leroux, F, Willery, E, Mathys, V, Deprez-Poulain, R, Delcroix, G, Frenois, F, Aumercier, M, Locht, C, Villeret, V, Deprez, B, Baulard, A.R. | | Deposit date: | 2009-01-30 | | Release date: | 2009-07-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Synthetic EthR inhibitors boost antituberculous activity of ethionamide.
NAT.MED. (N.Y.), 15, 2009
|
|
3G1L
 
 | | EthR from Mycobacterium tuberculosis in complex with compound BDM14744 | | Descriptor: | 3-(4-fluorophenyl)-5-phenyl-4H-1,2,4-triazole, TRANSCRIPTIONAL REGULATORY REPRESSOR PROTEIN (TETR-FAMILY) ETHR | | Authors: | Willand, N, Dirie, B, Carette, X, Bifani, P, Singhal, A, Desroses, M, Leroux, F, Willery, E, Mathys, V, Deprez-Poulain, R, Delcroix, G, Frenois, F, Aumercier, M, Locht, C, Villeret, V, Deprez, B, Baulard, A.R. | | Deposit date: | 2009-01-30 | | Release date: | 2009-07-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Synthetic EthR inhibitors boost antituberculous activity of ethionamide.
Nat.Med. (N.Y.), 15, 2009
|
|
6RZN
 
 | |
2UZH
 
 | | Mycobacterium smegmatis 2C-methyl-D-erythritol-2,4-cyclodiphosphate synthase (IspF) | | Descriptor: | 1,2-ETHANEDIOL, 2C-METHYL-D-ERYTHRITOL 2,4-CYCLODIPHOSPHATE SYNTHASE, 3-METHYLBUT-3-ENYL TRIHYDROGEN DIPHOSPHATE, ... | | Authors: | Buetow, L, Brown, A.C, Parish, T, Hunter, W.N. | | Deposit date: | 2007-04-27 | | Release date: | 2007-11-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Structure of Mycobacteria 2C-Methyl-D-Erythritol-2,4-Cyclodiphosphate Synthase, an Essential Enzyme, Provides a Platform for Drug Discovery.
Bmc Struct.Biol., 7, 2007
|
|
6U4O
 
 | |
