7JYO
 
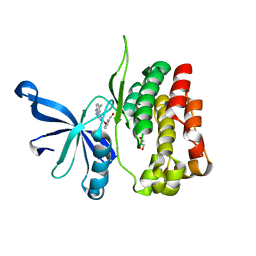 | | JAK2 JH2 in complex with JAK064 | | Descriptor: | 3-({4-amino-6-[(4-cyanophenyl)amino]-1,3,5-triazin-2-yl}oxy)benzoic acid, GLYCEROL, Tyrosine-protein kinase JAK2 | | Authors: | Puleo, D.E, Krimmer, S.G, Newton, A.S, Schlessinger, J, Jorgensen, W.L. | | Deposit date: | 2020-08-31 | | Release date: | 2021-07-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.16127229 Å) | | Cite: | Indoloxytriazines as binding molecules for the JAK2 JH2 pseudokinase domain and its V617F variant
Tetrahedron Lett., 77, 2021
|
|
7JYQ
 
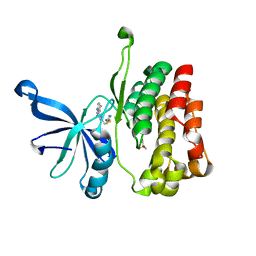 | | JAK2 JH2 in complex with JAK020 | | Descriptor: | GLYCEROL, N~2~-(4-fluorophenyl)-6-{[(5-{[(oxolan-2-yl)methyl]amino}-1,3,4-thiadiazol-2-yl)sulfanyl]methyl}-1,3,5-triazine-2,4-diamine, Tyrosine-protein kinase JAK2 | | Authors: | Puleo, D.E, Krimmer, S.G, Newton, A.S, Schlessinger, J, Jorgensen, W.L. | | Deposit date: | 2020-08-31 | | Release date: | 2021-07-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85941613 Å) | | Cite: | Indoloxytriazines as binding molecules for the JAK2 JH2 pseudokinase domain and its V617F variant
Tetrahedron Lett., 77, 2021
|
|
6DRW
 
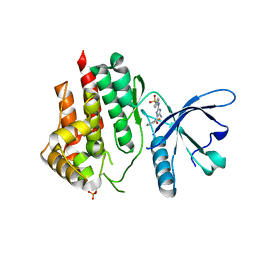 | |
1AGW
 
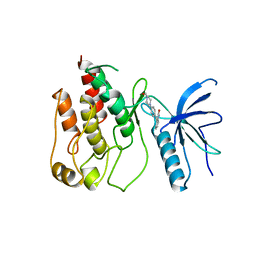 | |
3KXX
 
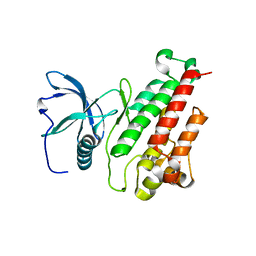 | | Structure of the mutant Fibroblast Growth Factor receptor 1 | | Descriptor: | Basic fibroblast growth factor receptor 1 | | Authors: | Bae, J.H, Boggon, T.J, Tome, F, Mandiyan, V, Lax, I, Schlessinger, J. | | Deposit date: | 2009-12-04 | | Release date: | 2010-03-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Asymmetric receptor contact is required for tyrosine autophosphorylation of fibroblast growth factor receptor in living cells.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3KY2
 
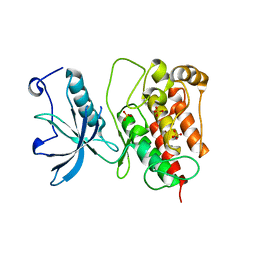 | | Crystal structure of Fibroblast Growth Factor Receptor 1 kinase domain | | Descriptor: | Basic fibroblast growth factor receptor 1, SULFATE ION | | Authors: | Bae, J.H, Boggon, T.J, Tome, F, Mandiyan, V, Lax, I, Schlessinger, J. | | Deposit date: | 2009-12-04 | | Release date: | 2010-03-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Asymmetric receptor contact is required for tyrosine autophosphorylation of fibroblast growth factor receptor in living cells.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
1AXM
 
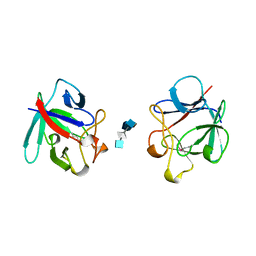 | | HEPARIN-LINKED BIOLOGICALLY-ACTIVE DIMER OF FIBROBLAST GROWTH FACTOR | | Descriptor: | 2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid, 2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, ACIDIC FIBROBLAST GROWTH FACTOR | | Authors: | DiGabriele, A.D, Lax, I, Chen, D.I, Svahn, C.M, Jaye, M, Schlessinger, J, Hendrickson, W.A. | | Deposit date: | 1997-10-16 | | Release date: | 1998-04-22 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of a heparin-linked biologically active dimer of fibroblast growth factor.
Nature, 393, 1998
|
|
4K9E
 
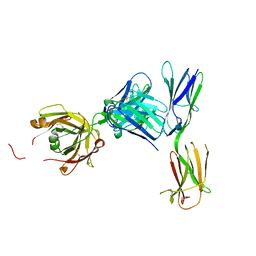 | | Crystal structure of KIT D4D5 fragment in complex with anti-Kit antibodies Fab79D | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Mast/stem cell growth factor receptor Kit, heavy chain, ... | | Authors: | Resheynyak, A.V, Boggon, T.J, Lax, I, Schlessinger, J. | | Deposit date: | 2013-04-19 | | Release date: | 2013-10-16 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural basis for KIT receptor tyrosine kinase inhibition by antibodies targeting the D4 membrane-proximal region.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
3GM2
 
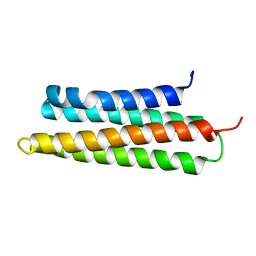 | |
3JS2
 
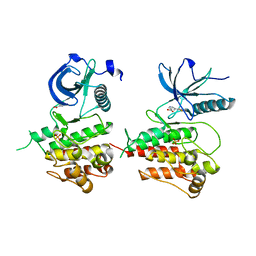 | | Crystal structure of minimal kinase domain of fibroblast growth factor receptor 1 in complex with 5-(2-thienyl)nicotinic acid | | Descriptor: | 5-(2-thienyl)nicotinic acid, Basic fibroblast growth factor receptor 1, PHOSPHATE ION | | Authors: | Bae, J.H, Ravindranathan, K.P, Mandiyan, V, Ekkati, A.R, Schlessinger, J, Jorgensen, W.L. | | Deposit date: | 2009-09-09 | | Release date: | 2010-02-23 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Discovery of novel fibroblast growth factor receptor 1 kinase inhibitors by structure-based virtual screening
J.Med.Chem., 53, 2010
|
|
2E9W
 
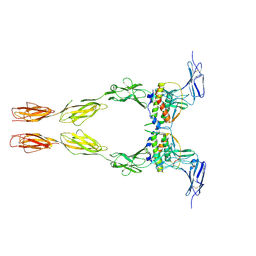 | | Crystal structure of the extracellular domain of Kit in complex with stem cell factor (SCF) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Kit ligand, Mast/stem cell growth factor receptor | | Authors: | Yuzawa, S, Opatowsky, Y, Zhang, Z, Mandiyan, V, Lax, I, Schlessinger, J. | | Deposit date: | 2007-01-27 | | Release date: | 2007-08-07 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural Basis for Activation of the Receptor Tyrosine Kinase KIT by Stem Cell Factor
Cell(Cambridge,Mass.), 130, 2007
|
|
4K94
 
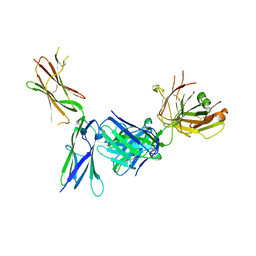 | | Crystal structure of KIT D4D5 fragment in complex with anti-Kit antibody Fab19 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Fab19 heavy chain, Fab19 light chain, ... | | Authors: | Resheynyak, A.V, Boggon, T.J, Lax, I, Schlessinger, J. | | Deposit date: | 2013-04-19 | | Release date: | 2013-10-16 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for KIT receptor tyrosine kinase inhibition by antibodies targeting the D4 membrane-proximal region.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
7MK7
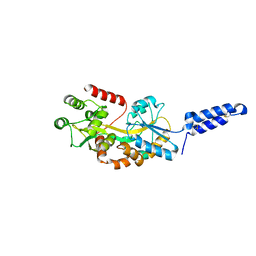 
 | | Augmentor domain of augmentor-beta | | Descriptor: | ALK and LTK ligand 1,Maltodextrin-binding protein, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Krimmer, S.G, Reshetnyak, A.V, Puleo, D.E, Schlessinger, J. | | Deposit date: | 2021-04-21 | | Release date: | 2021-11-24 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.42815185 Å) | | Cite: | Structural basis for ligand reception by anaplastic lymphoma kinase.
Nature, 600, 2021
|
|
7T0P
 
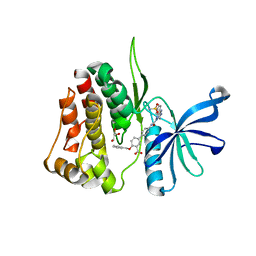 | | JAK2 JH2 IN COMPLEX WITH JAK315 | | Descriptor: | 4'-{[5-amino-3-(4-sulfamoylanilino)-1H-1,2,4-triazole-1-carbonyl]amino}-4-(benzyloxy)[1,1'-biphenyl]-3-carboxylic acid, GLYCEROL, Tyrosine-protein kinase JAK2 | | Authors: | Ippolito, J.A, Liosi, M.-E, Krimmer, S.G, Schlessinger, J, Jorgensen, W.L. | | Deposit date: | 2021-11-30 | | Release date: | 2022-06-15 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Insights on JAK2 Modulation by Potent, Selective, and Cell-Permeable Pseudokinase-Domain Ligands.
J.Med.Chem., 65, 2022
|
|
6XJK
 
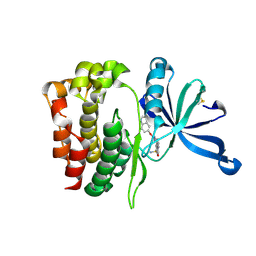 | | JAK2 JH2 in complex with JAK067 | | Descriptor: | 4-({4-amino-6-[(1H-indol-5-yl)oxy]-1,3,5-triazin-2-yl}amino)benzene-1-sulfonamide, Tyrosine-protein kinase JAK2 | | Authors: | Puleo, D.E, Krimmer, S.G, Newton, A.S, Schlessinger, J, Jorgensen, W.L. | | Deposit date: | 2020-06-24 | | Release date: | 2020-11-25 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.023508 Å) | | Cite: | Explicit Representation of Cation-pi Interactions in Force Fields with 1/r4 Nonbonded Terms.
J Chem Theory Comput, 16, 2020
|
|
3GM1
 
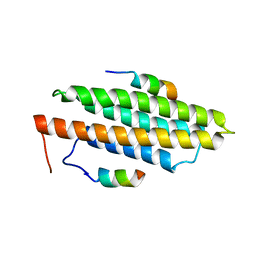 | |
7T1T
 
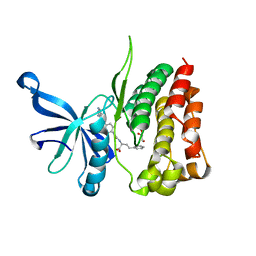 | | JAK2 JH2 IN COMPLEX WITH JAK292 | | Descriptor: | (2S)-2-[({4-[(2-amino-7H-pyrrolo[2,3-d]pyrimidin-4-yl)oxy]phenyl}carbamoyl)amino]-4-phenylbutanoic acid, GLYCEROL, Tyrosine-protein kinase JAK2 | | Authors: | Ippolito, J.A, Henry, S, Krimmer, S.G, Schlessinger, J, Jorgensen, W.L. | | Deposit date: | 2021-12-02 | | Release date: | 2022-05-25 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Conversion of a False Virtual Screen Hit into Selective JAK2 JH2 Domain Binders Using Convergent Design Strategies
Acs Med.Chem.Lett., 13, 2022
|
|
2EC8
 
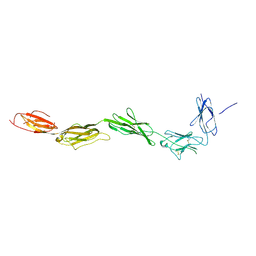 | | Crystal structure of the exctracellular domain of the receptor tyrosine kinase, Kit | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Mast/stem cell growth factor receptor | | Authors: | Yuzawa, S, Opatowsky, Y, Zhang, Z, Mandiyan, V, Lax, I, Schlessinger, J. | | Deposit date: | 2007-02-11 | | Release date: | 2007-08-07 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural Basis for Activation of the Receptor Tyrosine Kinase KIT by Stem Cell Factor
Cell(Cambridge,Mass.), 130, 2007
|
|
3GQI
 
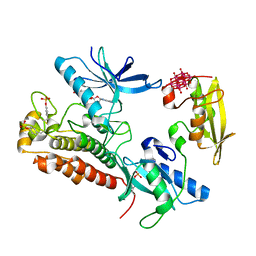 | | Crystal Structure of activated receptor tyrosine kinase in complex with substrates | | Descriptor: | Basic fibroblast growth factor receptor 1, DECAVANADATE, MAGNESIUM ION, ... | | Authors: | Bae, J.H, Lew, E.D, Yuzawa, S, Tome, F, Lax, I, Schlessinger, J. | | Deposit date: | 2009-03-24 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The selectivity of receptor tyrosine kinase signaling is controlled by a secondary SH2 domain binding site.
Cell(Cambridge,Mass.), 138, 2009
|
|
3GQL
 
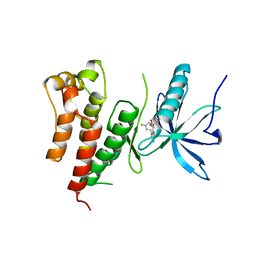 | | Crystal Structure of activated receptor tyrosine kinase in complex with substrates | | Descriptor: | (E)-[4-(3,5-difluorophenyl)-3H-pyrrolo[2,3-b]pyridin-3-ylidene](3-methoxyphenyl)methanol, Basic fibroblast growth factor receptor 1 | | Authors: | Bae, J.H, Lew, E.D, Yuzawa, S, Tome, F, Lax, I, Schlessinger, J. | | Deposit date: | 2009-03-24 | | Release date: | 2009-08-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The selectivity of receptor tyrosine kinase signaling is controlled by a secondary SH2 domain binding site.
Cell(Cambridge,Mass.), 138, 2009
|
|
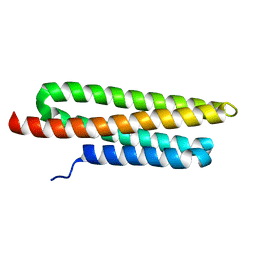
3GM3
 
 | |
1FGI
 
 | |
3KVQ
 
 | |
5VAN
 
 | | Crystal Structure of Beta-Klotho | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lee, S, Schlessinger, J. | | Deposit date: | 2017-03-27 | | Release date: | 2018-01-31 | | Last modified: | 2021-03-24 | | Method: | X-RAY DIFFRACTION (2.202 Å) | | Cite: | Structures of beta-klotho reveal a 'zip code'-like mechanism for endocrine FGF signalling.
Nature, 553, 2018
|
|
5VAK
 
 | | Crystal Structure of Beta-Klotho, Domain 1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Beta-klotho, ... | | Authors: | Lee, S, Schlessinger, J. | | Deposit date: | 2017-03-27 | | Release date: | 2018-01-31 | | Last modified: | 2021-03-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of beta-klotho reveal a 'zip code'-like mechanism for endocrine FGF signalling.
Nature, 553, 2018
|
|
