2AWF
 
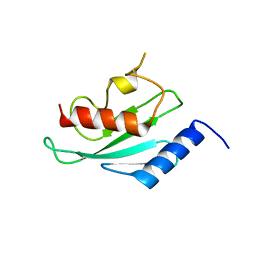 | | Structure of human Ubiquitin-conjugating enzyme E2 G1 | | Descriptor: | Ubiquitin-conjugating enzyme E2 G1 | | Authors: | Walker, J.R, Avvakumov, G.V, Xue, S, Newman, E.M, Finerty, P, Mackenzie, F, Weigelt, J, Sundstrom, M, Arrowsmith, C, Edwards, A, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2005-09-01 | | Release date: | 2005-09-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A human ubiquitin conjugating enzyme (E2)-HECT E3 ligase structure-function screen.
Mol Cell Proteomics, 11, 2012
|
|
2CLQ
 
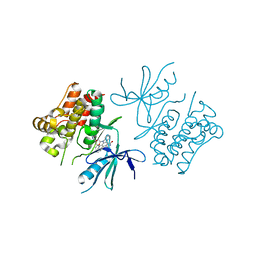 | | Structure of mitogen-activated protein kinase kinase kinase 5 | | Descriptor: | MITOGEN-ACTIVATED PROTEIN KINASE KINASE KINASE 5, STAUROSPORINE | | Authors: | Bunkoczi, G, Salah, E, Fedorov, O, Pike, A, Gileadi, O, von Delft, F, Arrowsmith, C, Edwards, A, Sundstrom, M, Weigelt, J, Knapp, S. | | Deposit date: | 2006-04-28 | | Release date: | 2006-05-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and Functional Characterization of the Human Protein Kinase Ask1.
Structure, 15, 2007
|
|
2VZ6
 
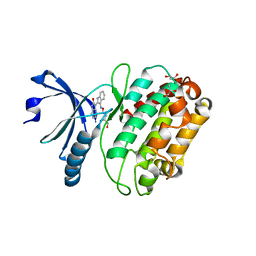 | | Structure of human calcium calmodulin dependent protein kinase type II alpha (CAMK2A) in complex with Indirubin E804 | | Descriptor: | (2Z,3E)-2,3'-BIINDOLE-2',3(1H,1'H)-DIONE 3-{O-[(3R)-3,4-DIHYDROXYBUTYL]OXIME}, CALCIUM CALMODULIN DEPENDENT PROTEIN KINASE TYPE II ALPHA CHAIN, S-1,2-PROPANEDIOL | | Authors: | Pike, A.C.W, Rellos, P, King, O, Salah, E, Parizotto, E, Fedorov, O, Shrestha, L, Burgess-Brown, N, Roos, A, Murray, J.W, von Delft, F, Edwards, A, Arrowsmith, C.H, Wikstroem, M, Bountra, C, Knapp, S. | | Deposit date: | 2008-07-30 | | Release date: | 2008-08-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the Camkiidelta/Calmodulin Complex Reveals the Molecular Mechanism of Camkii Kinase Activation.
Plos Biol., 8, 2010
|
|
2WEL
 
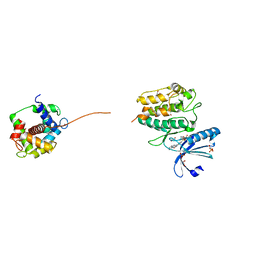 | | Crystal structure of SU6656-bound calcium/calmodulin-dependent protein kinase II delta in complex with calmodulin | | Descriptor: | (3Z)-N,N-DIMETHYL-2-OXO-3-(4,5,6,7-TETRAHYDRO-1H-INDOL-2-YLMETHYLIDENE)-2,3-DIHYDRO-1H-INDOLE-5-SULFONAMIDE, 1,2-ETHANEDIOL, CALCIUM ION, ... | | Authors: | Pike, A.C.W, Rellos, P, Salah, E, Burgess-Brown, N, Keates, T, Muniz, J, Sethi, R, Roos, A, Filippakopoulos, P, von Delft, F, Edwards, A, Weigelt, J, Arrowsmith, C.H, Bountra, C, Knapp, S. | | Deposit date: | 2009-03-31 | | Release date: | 2009-04-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the Camkiidelta/Calmodulin Complex Reveals the Molecular Mechanism of Camkii Kinase Activation.
Plos Biol., 8, 2010
|
|
2WA0
 
 | | Crystal structure of the human MAGEA4 | | Descriptor: | MELANOMA-ASSOCIATED ANTIGEN 4 | | Authors: | Roos, A.K, Cooper, C.D.O, Ugochukwu, E, W Yue, W, Berridge, G, Elkins, J.M, Pike, A.C.W, Bray, J, Filippakopoulos, P, Muniz, J, Chaikuad, A, Burgess-Brown, N, Arrowsmith, C.H, Weigelt, J, Edwards, A, Bountra, C, von Delft, F, Gileadi, O, Oppermann, U. | | Deposit date: | 2009-01-31 | | Release date: | 2009-03-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of Two Melanoma-Associated Antigens Suggest Allosteric Regulation of Effector Binding.
Plos One, 11, 2016
|
|
2A7L
 
 | | Structure of the human hypothetical ubiquitin-conjugating enzyme, LOC55284 | | Descriptor: | Hypothetical ubiquitin-conjugating enzyme LOC55284, SODIUM ION | | Authors: | Walker, J.R, Avvakumov, G.V, Xue, S, Newman, E.M, Mackenzie, F, Weigelt, J, Sundstrom, M, Arrowsmith, C, Edwards, A, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2005-07-05 | | Release date: | 2005-09-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | A human ubiquitin conjugating enzyme (E2)-HECT E3 ligase structure-function screen.
Mol Cell Proteomics, 11, 2012
|
|
2A4D
 
 | | Structure of the human ubiquitin-conjugating enzyme E2 variant 1 (UEV-1) | | Descriptor: | Ubiquitin-conjugating enzyme E2 variant 1 | | Authors: | Walker, J.R, Avvakumov, G.V, Xue, S, Newman, E.M, Mackenzie, F, Weigelt, J, Sundstrom, M, Arrowsmith, C, Edwards, A, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2005-06-28 | | Release date: | 2005-07-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | A human ubiquitin conjugating enzyme (E2)-HECT E3 ligase structure-function screen.
Mol Cell Proteomics, 11, 2012
|
|
2WWU
 
 | | Crystal structure of the catalytic domain of PHD finger protein 8 | | Descriptor: | ACETATE ION, NICKEL (II) ION, PHD FINGER PROTEIN 8, ... | | Authors: | Yue, W.W, Hozjan, V, Cooper, C, Tumber, A, Krojer, T, Muniz, J, Allerston, C, Salah, E, McDonough, M.A, von Delft, F, Arrowsmith, C, Weigelt, J, Edwards, A, Bountra, C, Schofield, C.J, Kavanagh, K.L, Oppermann, U. | | Deposit date: | 2009-10-29 | | Release date: | 2009-11-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structure of the PHF8 Jumonji domain, an Nepsilon-methyl lysine demethylase.
FEBS Lett., 584, 2010
|
|
5A4R
 
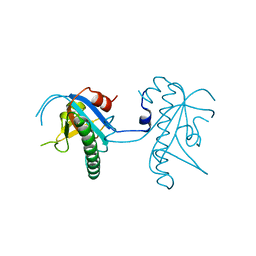 | | Crystal structure of a vitamin B12 trafficking protein | | Descriptor: | METHYLMALONIC ACIDURIA AND HOMOCYSTINURIA TYPE D HOMOLOG, MITOCHONDRIAL | | Authors: | Kopec, J, Fitzpatrick, F, Froese, D.S, Velupillai, S, Nowak, R, Chalk, R, Burgess-Brown, N, von Delft, F, Arrowsmith, C, Edwards, A, Bountra, C, Fowler, B, Baumgartner, M.R, Yue, W.W. | | Deposit date: | 2015-06-11 | | Release date: | 2015-10-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural Insights Into the Mmachc-Mmadhc Protein Complex Involved in Vitamin B12 Trafficking.
J.Biol.Chem., 290, 2015
|
|
2JNU
 
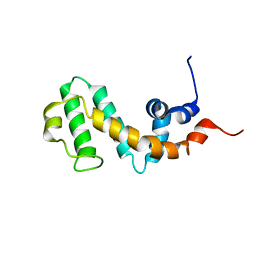 | | Solution structure of the RGS domain of human RGS14 | | Descriptor: | Regulator of G-protein signaling 14 | | Authors: | Dowler, E.F, Diehl, A, Bray, J, Elkins, J, Soundararajan, M, Doyle, D.A, Gileadi, C, Phillips, C, Schoch, G.A, Yang, X, Brockmann, C, Leidert, M, Rehbein, K, Schmieder, P, Kuhne, R, Higman, V.A, Sundstrom, M, Arrowsmith, C, Weigelt, J, Edwards, A, Oschkinat, H, Ball, L.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-02-02 | | Release date: | 2007-02-27 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structural diversity in the RGS domain and its interaction with heterotrimeric G protein alpha-subunits.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2JP2
 
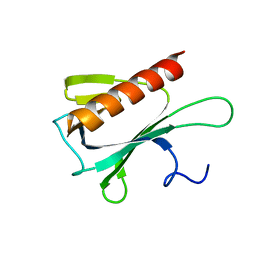 | | Solution structure and resonance assignment of the N-terminal EVH1 domain from the human Spred2 protein (Sprouty-related protein with EVH1 domain isoform 2) | | Descriptor: | Sprouty-related, EVH1 domain-containing protein 2 | | Authors: | Fossi, M, Zimmermann, J, Jarchau, T, Lemak, A, Walter, U, Wiegelt, J, Sundstrom, M, Arrowsmith, C, Edwards, A, Oschkinat, H, Ball, L.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-04-18 | | Release date: | 2007-05-15 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | 1H, 13C and 15N resonance assignment of the human Spred2 EVH1 domain
J.Biomol.NMR, 29, 2004
|
|
2JGY
 
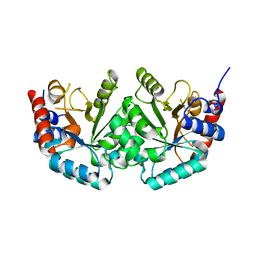 | | The crystal structure of human orotidine-5'-decarboxylase domain of human uridine monophosphate synthetase (UMPS) | | Descriptor: | UMP SYNTHASE | | Authors: | Moche, M, Ogg, D, Arrowsmith, C, Berglund, H, Busam, R, Collins, R, Dahlgren, L.G, Edwards, A, Ericsson, U.B, Flodin, S, Flores, A, Graslund, S, Hammarstrom, M, Hallberg, B.M, Holmberg-Schiavone, L, Johansson, I, Karlberg, T, Kosinska, U, Kotenyova, T, Lehtio, L, Nilsson, M.E, Nyman, T, Persson, C, Sagemark, J, Stenmark, P, Sundstrom, M, Uppenberg, J, Upsten, M, Thorsell, A.G, van den Berg, S, Weigelt, J, Nordlund, P, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-02-16 | | Release date: | 2007-03-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Crystal Structure of Human Orotidine-5'-Decarboxylase Domain of Human Uridine Monophosphate Synthetase (Umps)
To be Published
|
|
5AGA
 
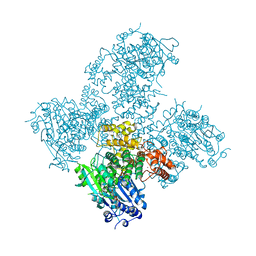 | | Crystal structure of the Helicase domain of human DNA polymerase theta in complex with AMPPNP | | Descriptor: | CITRATE ANION, DNA POLYMERASE THETA, MAGNESIUM ION, ... | | Authors: | Newman, J.A, Cooper, C.D.O, Aitkenhead, H, Pinkas, D.M, Kupinska, K, Burgess-Brown, N, von Delft, F, Arrowsmith, C.H, Edwards, A, Bountra, C, Gileadi, O. | | Deposit date: | 2015-01-29 | | Release date: | 2015-02-25 | | Last modified: | 2015-12-16 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure of the Helicase Domain of DNA Polymerase Theta Reveals a Possible Role in the Microhomology-Mediated End-Joining Pathway.
Structure, 23, 2015
|
|
2NZ6
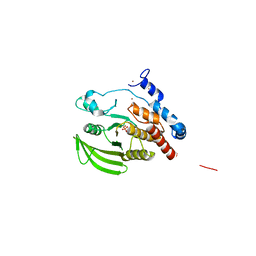 
 | | Crystal structure of the PTPRJ inactivating mutant C1239S | | Descriptor: | CHLORIDE ION, NICKEL (II) ION, PHOSPHATE ION, ... | | Authors: | Ugochukwu, E, Barr, A, Savitsky, P, Pike, A.C.W, Bunkoczi, G, Sundstrom, M, Weigelt, J, Arrowsmith, C.H, Edwards, A, von Delft, F, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-11-22 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Large-scale structural analysis of the classical human protein tyrosine phosphatome.
Cell(Cambridge,Mass.), 136, 2009
|
|
1T2A
 
 | | Crystal structure of human GDP-D-mannose 4,6-dehydratase | | Descriptor: | GDP-mannose 4,6 dehydratase, GUANOSINE-5'-DIPHOSPHATE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Walker, J.R, Vedadi, M, Sharma, S, Houston, S, Wasney, G, Loppnau, P, Sundstrom, M, Arrowsmith, C, Edwards, A, Oppermann, U. | | Deposit date: | 2004-04-20 | | Release date: | 2004-05-10 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal Structure and Biophysical Characterization of Human GDP-D-mannose 4,6-dehydratase
To be Published
|
|
2J90
 
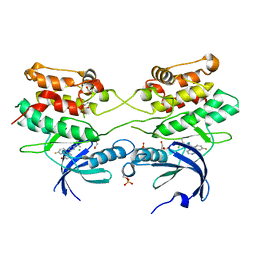 | | Crystal structure of human ZIP kinase in complex with a tetracyclic pyridone inhibitor (Pyridone 6) | | Descriptor: | 1,2-ETHANEDIOL, 2-TERT-BUTYL-9-FLUORO-3,6-DIHYDRO-7H-BENZ[H]-IMIDAZ[4,5-F]ISOQUINOLINE-7-ONE, CHLORIDE ION, ... | | Authors: | Turnbull, A.P, Berridge, G, Fedorov, O, Pike, A.C.W, Savitsky, P, Eswaran, J, Papagrigoriou, E, Ugochukwa, E, von Delft, F, Gileadi, O, Arrowsmith, C.H, Edwards, A, Weigelt, J, Sundstrom, M, Knapp, S. | | Deposit date: | 2006-10-31 | | Release date: | 2006-11-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Activation Segment Dimerization: A Mechanism for Kinase Autophosphorylation of Non-Consensus Sites.
Embo J., 27, 2008
|
|
2J7T
 
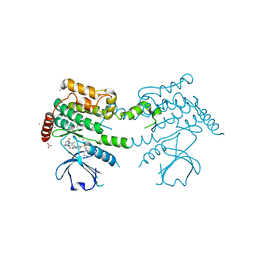 | | Crystal structure of human serine threonine kinase-10 bound to SU11274 | | Descriptor: | (3Z)-N-(3-CHLOROPHENYL)-3-({3,5-DIMETHYL-4-[(4-METHYLPIPERAZIN-1-YL)CARBONYL]-1H-PYRROL-2-YL}METHYLENE)-N-METHYL-2-OXOINDOLINE-5-SULFONAMIDE, ACETATE ION, CALCIUM ION, ... | | Authors: | Pike, A.C.W, Rellos, P, Fedorov, O, Das, S, Debreczeni, J, Sobott, F, Watt, S, Savitsky, P, Eswaran, J, Turnbull, A.P, Papagrigoriou, E, Ugochukwa, E, Gorrec, F, Umeano, C.C, von Delft, F, Arrowsmith, C.H, Edwards, A, Weigelt, J, Sundstrom, M, Knapp, S. | | Deposit date: | 2006-10-17 | | Release date: | 2006-11-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Activation Segment Dimerization: A Mechanism for Kinase Autophosphorylation of Non-Consensus Sites.
Embo J., 27, 2008
|
|
2J6L
 
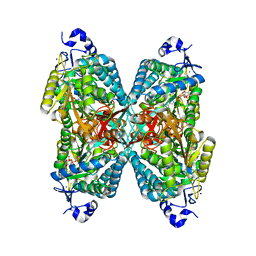 | | Structure of aminoadipate-semialdehyde dehydrogenase | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, ALDEHYDE DEHYDROGENASE FAMILY 7 MEMBER A1, BROMIDE ION, ... | | Authors: | Bunkoczi, G, Guo, K, Debreczeni, J.E, Smee, C, Arrowsmith, C, Edwards, A, Sundstrom, M, Weigelt, J, von Delft, F, Oppermann, U. | | Deposit date: | 2006-09-29 | | Release date: | 2006-10-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Aldehyde Dehydrogenase 7A1 (Aldh7A1) is a Novel Enzyme Involved in Cellular Defense Against Hyperosmotic Stress.
J.Biol.Chem., 285, 2010
|
|
2J51
 
 | | Crystal structure of Human STE20-like kinase bound to 5-Amino-3-((4-(aminosulfonyl)phenyl)amino) -N-(2,6-difluorophenyl)-1H-1,2,4-triazole- 1-carbothioamide | | Descriptor: | 1,2-ETHANEDIOL, 5-AMINO-3-{[4-(AMINOSULFONYL)PHENYL]AMINO}-N-(2,6-DIFLUOROPHENYL)-1H-1,2,4-TRIAZOLE-1-CARBOTHIOAMIDE, STE20-LIKE SERINE/THREONINE-PROTEIN KINASE, ... | | Authors: | Pike, A.C.W, Rellos, P, Fedorov, O, Keates, T, Salah, E, Savitsky, P, Papagrigoriou, E, Turnbull, A.P, Debreczeni, J.E, von Delft, F, Arrowsmith, C, Edwards, A, Weigelt, J, Sundstrom, M, Knapp, S. | | Deposit date: | 2006-09-08 | | Release date: | 2006-10-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Activation Segment Dimerization: A Mechanism for Kinase Autophosphorylation of Non-Consensus Sites.
Embo J., 27, 2008
|
|
2JFM
 
 | | CRYSTAL STRUCTURE OF HUMAN STE20-LIKE KINASE (UNLIGANDED FORM) | | Descriptor: | 1,2-ETHANEDIOL, STE20-LIKE SERINE-THREONINE KINASE | | Authors: | Pike, A.C.W, Rellos, P, Fedorov, O, Keates, T, Salah, E, Savitsky, P, Papagrigoriou, E, Bunkoczi, G, von Delft, F, Arrowsmith, C.H, Edwards, A, Weigelt, J, Sundstrom, M, Knapp, S. | | Deposit date: | 2007-02-02 | | Release date: | 2007-02-27 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Activation Segment Dimerization: A Mechanism for Kinase Autophosphorylation of Non-Consensus Sites.
Embo J., 27, 2008
|
|
2OO1
 
 | | Crystal structure of the Bromo domain 2 of human Bromodomain containing protein 3 (BRD3) | | Descriptor: | 1,2-ETHANEDIOL, 2-(2-(2-(2-(2-(2-ETHOXYETHOXY)ETHOXY)ETHOXY)ETHOXY)ETHOXY)ETHANOL, Bromodomain-containing protein 3, ... | | Authors: | Filippakopoulos, P, Bullock, A, Papagrigoriou, E, Keates, T, Cooper, C, Smee, C, Ugochukwu, E, Debreczeni, J, von Delft, F, Arrowsmith, C, Edwards, A, Weigelt, J, Sundstrom, M, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-01-25 | | Release date: | 2007-02-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Histone recognition and large-scale structural analysis of the human bromodomain family.
Cell(Cambridge,Mass.), 149, 2012
|
|
2OOQ
 
 | | Crystal Structure of the Human Receptor Phosphatase PTPRT | | Descriptor: | 1,2-ETHANEDIOL, 2-[3-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-PROPYLAMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Receptor-type tyrosine-protein phosphatase T, ... | | Authors: | Ugochukwu, E, Alfano, I, Barr, A, Keates, T, Eswaran, J, Salah, E, Savitsky, P, Bunkoczi, G, Edwards, A, Arrowsmith, C.H, Weigelt, J, Sundstrom, M, von Delft, F, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-01-26 | | Release date: | 2007-02-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Large-scale structural analysis of the classical human protein tyrosine phosphatome.
Cell(Cambridge,Mass.), 136, 2009
|
|
2P6N
 
 | | Human DEAD-box RNA helicase DDX41, helicase domain | | Descriptor: | ATP-dependent RNA helicase DDX41 | | Authors: | Karlberg, T, Ogg, D, Arrowsmith, C.H, Berglund, H, Busam, R.D, Collins, R, Dahlgren, L.G, Edwards, A, Flodin, S, Flores, A, Graslund, S, Hallberg, B.M, Hammarstrom, M, Johansson, I, Kotenyova, T, Lehtio, L, Moche, M, Nordlund, P, Nyman, T, Persson, C, Sagemark, J, Stenmark, P, Sundstrom, M, Thorsell, A.G, Van Den Berg, S, Weigelt, J, Holmberg-Schiavone, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-03-19 | | Release date: | 2007-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Comparative Structural Analysis of Human DEAD-Box RNA Helicases.
Plos One, 5, 2010
|
|
5RLJ
 
 | | PanDDA analysis group deposition -- Crystal Structure of SARS-CoV-2 helicase in complex with Z1407673036 | | Descriptor: | (2S)-2-phenylpropane-1-sulfonamide, Helicase, PHOSPHATE ION, ... | | Authors: | Newman, J.A, Yosaatmadja, Y, Douangamath, A, Aimon, A, Powell, A.J, Dias, A, Fearon, D, Dunnett, L, Brandao-Neto, J, Krojer, T, Skyner, R, Gorrie-Stone, T, Thompson, W, von Delft, F, Arrowsmith, C.H, Edwards, A, Bountra, C, Gileadi, O. | | Deposit date: | 2020-09-16 | | Release date: | 2020-09-30 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.879 Å) | | Cite: | Structure, mechanism and crystallographic fragment screening of the SARS-CoV-2 NSP13 helicase.
Nat Commun, 12, 2021
|
|
5RM2
 
 | | PanDDA analysis group deposition -- Crystal Structure of SARS-CoV-2 helicase in complex with Z1741964527 | | Descriptor: | 1-(diphenylmethyl)azetidin-3-ol, Helicase, PHOSPHATE ION, ... | | Authors: | Newman, J.A, Yosaatmadja, Y, Douangamath, A, Aimon, A, Powell, A.J, Dias, A, Fearon, D, Dunnett, L, Brandao-Neto, J, Krojer, T, Skyner, R, Gorrie-Stone, T, Thompson, W, von Delft, F, Arrowsmith, C.H, Edwards, A, Bountra, C, Gileadi, O. | | Deposit date: | 2020-09-16 | | Release date: | 2020-09-30 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structure, mechanism and crystallographic fragment screening of the SARS-CoV-2 NSP13 helicase.
Nat Commun, 12, 2021
|
|
