6YY4
 
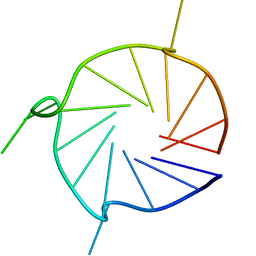 | | Parallel 17-mer DNA G-quadruplex | | Descriptor: | DNA (5'-D(*GP*GP*GP*TP*GP*GP*GP*AP*AP*GP*GP*GP*TP*GP*GP*GP*A)-3') | | Authors: | Srb, P, Curtis, C, Veverka, V. | | Deposit date: | 2020-05-04 | | Release date: | 2021-01-20 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Overlapping but distinct: a new model for G-quadruplex biochemical specificity.
Nucleic Acids Res., 49, 2021
|
|
2MBD
 
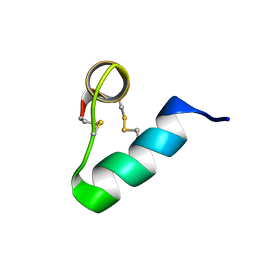 | | Lasiocepsin | | Descriptor: | lasiocepsin | | Authors: | Monincova, L, Budesinsky, M, Cujova, S, Cerovsky, V, Veverka, V. | | Deposit date: | 2013-07-30 | | Release date: | 2014-01-15 | | Last modified: | 2014-01-29 | | Method: | SOLUTION NMR | | Cite: | Structural basis for antimicrobial activity of lasiocepsin.
Chembiochem, 15, 2014
|
|
5O2P
 
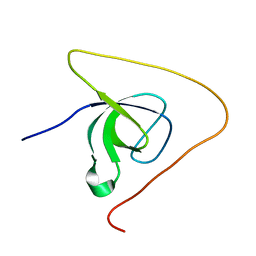 | | p130Cas SH3 domain PTP-PEST peptide chimera | | Descriptor: | Breast cancer anti-estrogen resistance 1,Tyrosine-protein phosphatase non-receptor type 12 | | Authors: | Hexnerova, R, Veverka, V. | | Deposit date: | 2017-05-22 | | Release date: | 2017-09-20 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of CAS SH3 domain selectivity and regulation reveals new CAS interaction partners.
Sci Rep, 7, 2017
|
|
5O2M
 
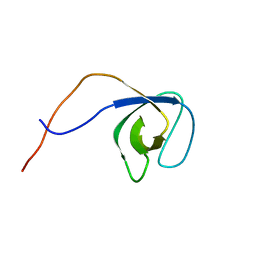 | | p130Cas SH3 domain | | Descriptor: | Breast cancer anti-estrogen resistance 1 | | Authors: | Hexnerova, R, Veverka, V. | | Deposit date: | 2017-05-22 | | Release date: | 2017-09-20 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of CAS SH3 domain selectivity and regulation reveals new CAS interaction partners.
Sci Rep, 7, 2017
|
|
5O2Q
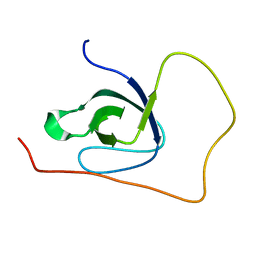 
 | | p130Cas SH3 domain Vinculin peptide chimera | | Descriptor: | Breast cancer anti-estrogen resistance 1,Vinculin | | Authors: | Hexnerova, R, Veverka, V. | | Deposit date: | 2017-05-22 | | Release date: | 2017-09-20 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of CAS SH3 domain selectivity and regulation reveals new CAS interaction partners.
Sci Rep, 7, 2017
|
|
2KZT
 
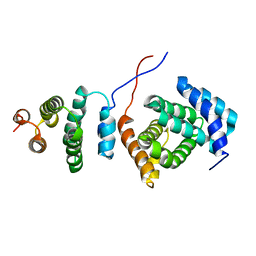 | | Structure of the Tandem MA-3 Region of Pdcd4 | | Descriptor: | Programmed cell death protein 4 | | Authors: | Waters, L.C, Strong, S.L, Oka, O, Muskett, F.W, Veverka, V, Banerjee, S, Schmedt, T, Henry, A.J, Klempnauer, K.H, Carr, M.D. | | Deposit date: | 2010-06-24 | | Release date: | 2011-03-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of the tandem MA-3 region of Pdcd4 protein and characterization of its interactions with eIF4A and eIF4G: molecular mechanisms of a tumor suppressor
J.Biol.Chem., 286, 2011
|
|
2M2N
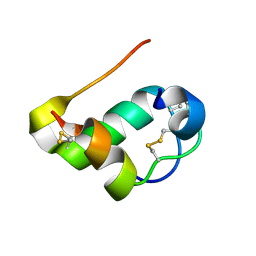 
 | |
2M2O
 
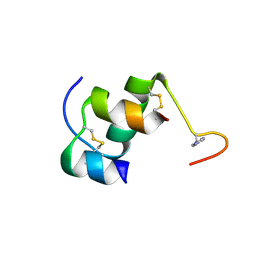 | |
2MSR
 
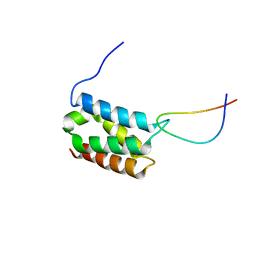 | | Solution structure of LEDGF/p75 IBD in complex with MLL1 peptide (140-160) | | Descriptor: | Histone-lysine N-methyltransferase 2A, PC4 and SFRS1-interacting protein | | Authors: | Cermakova, K, Tesina, P, Demeulemeester, J, El Ashkar, S, Mereau, H, Schwaller, J, Rezacova, P, Veverka, V, De Rijck, J. | | Deposit date: | 2014-08-05 | | Release date: | 2014-08-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Validation and Structural Characterization of the LEDGF/p75-MLL Interface as a New Target for the Treatment of MLL-Dependent Leukemia.
Cancer Res., 74, 2014
|
|
2MVD
 
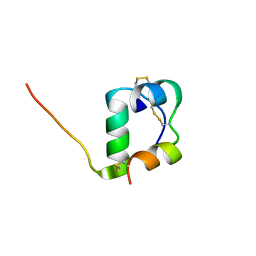 | | Solution structure of [GlnB22]-insulin mutant at pH 1.9 | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Hexnerova, R, Krizkova, K, Maletinska, L, Jiracek, J, Brzozowski, A.M, Zakova, L, Veverka, V. | | Deposit date: | 2014-10-02 | | Release date: | 2014-12-10 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural and Functional Study of the GlnB22-Insulin Mutant Responsible for Maturity-Onset Diabetes of the Young.
Plos One, 9, 2014
|
|
2N7E
 
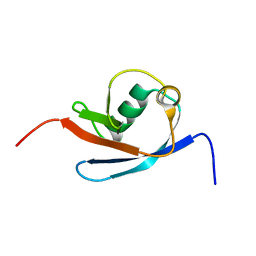 | |
2N7D
 
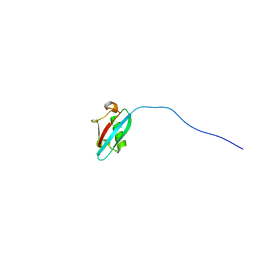 | |
2MVC
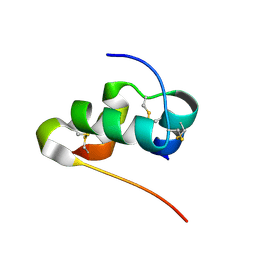 
 | | Solution structure of human insulin at pH 1.9 | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Hexnerova, R, Krizkova, K, Maletinska, L, Jiracek, J, Brzozowski, A.M, Zakova, L, Veverka, V. | | Deposit date: | 2014-10-02 | | Release date: | 2014-12-10 | | Last modified: | 2016-06-01 | | Method: | SOLUTION NMR | | Cite: | Structural and Functional Study of the GlnB22-Insulin Mutant Responsible for Maturity-Onset Diabetes of the Young.
Plos One, 9, 2014
|
|
2N3A
 
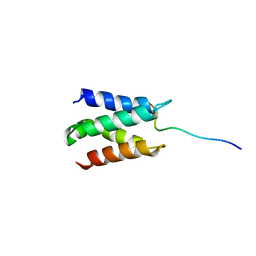 | | Solution structure of LEDGF/p75 IBD in complex with POGZ peptide (1389-1404) | | Descriptor: | PC4 and SFRS1-interacting protein, Pogo transposable element with ZNF domain | | Authors: | Tesina, P, Cermakova, K, Horejsi, M, Prochazkova, K, Fabry, M, Sharma, S, Christ, F, Demeulemeester, J, Debyser, Z, De Rijck, J, Veverka, V, Rezacova, P. | | Deposit date: | 2015-05-26 | | Release date: | 2015-08-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Multiple cellular proteins interact with LEDGF/p75 through a conserved unstructured consensus motif.
Nat Commun, 6, 2015
|
|
2MPG
 
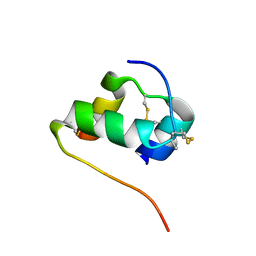 | | Solution structure of the [AibB8,LysB28,ProB29]-insulin analogue | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Kosinova, L, Jiracek, J, Zakova, L, Veverka, V. | | Deposit date: | 2014-05-17 | | Release date: | 2014-06-11 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Insight into the structural and biological relevance of the T/R transition of the N-terminus of the B-chain in human insulin.
Biochemistry, 53, 2014
|
|
2M2P
 
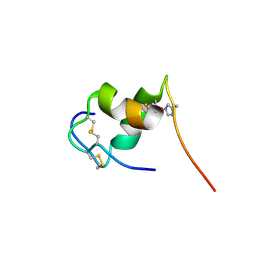 | |
2M2M
 
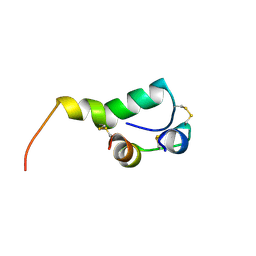 | |
4MUM
 
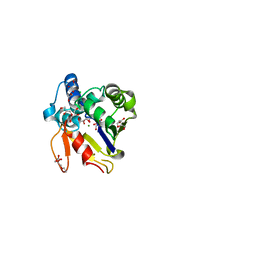 | |
4NFL
 
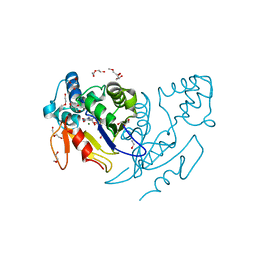 | | Crystal structure of human mitochondrial 5'(3')-deoxyribonucleotidase in complex with the inhibitor NPB-T | | Descriptor: | 1-{2-deoxy-3,5-O-[(4-nitrophenyl)(phosphono)methylidene]-beta-D-threo-pentofuranosyl}-5-methylpyrimidine-2,4(1H,3H)-dione, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 5'(3')-deoxyribonucleotidase, ... | | Authors: | Pachl, P, Rezacova, P, Brynda, J. | | Deposit date: | 2013-10-31 | | Release date: | 2014-09-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.375 Å) | | Cite: | Conformationally constrained nucleoside phosphonic acids - potent inhibitors of human mitochondrial and cytosolic 5'(3')-nucleotidases.
Org.Biomol.Chem., 12, 2014
|
|
5BQQ
 
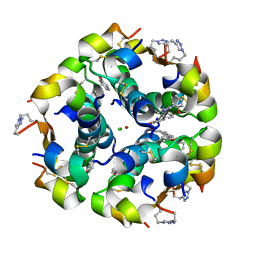 | | Human insulin with intra-chain chemical crosslink between modified B27 and B30 | | Descriptor: | CHLORIDE ION, Insulin, PHENOL, ... | | Authors: | Brzozowski, A.M, Turkenburg, J.P, Jiracek, J, Zakova, L. | | Deposit date: | 2015-05-29 | | Release date: | 2016-02-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Rational steering of insulin binding specificity by intra-chain chemical crosslinking.
Sci Rep, 6, 2016
|
|
7Z5A
 
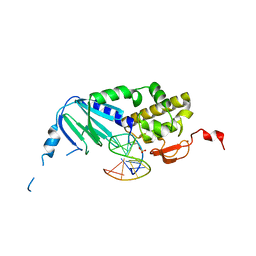 | |
6TRJ
 
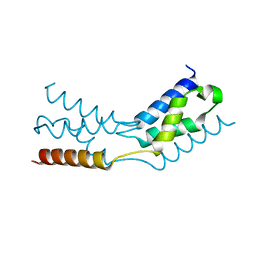 | | LEDGF/p75 IBD dimer | | Descriptor: | PC4 and SFRS1-interacting protein | | Authors: | Kugler, M, Brynda, J. | | Deposit date: | 2019-12-19 | | Release date: | 2020-09-09 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Fine-tuning of the LEDGF/p75 interaction network by dimerization
Structure
|
|
5OPK
 
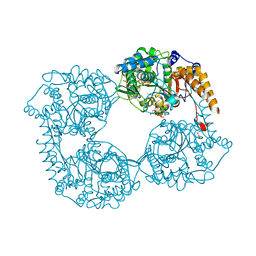 | | Crystal structure of D52N/R367Q cN-II mutant bound to dATP and free phosphate | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, Cytosolic purine 5'-nucleotidase, GLYCEROL, ... | | Authors: | Hnizda, A, Pachl, P, Rezacova, P. | | Deposit date: | 2017-08-10 | | Release date: | 2018-06-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Relapsed acute lymphoblastic leukemia-specific mutations in NT5C2 cluster into hotspots driving intersubunit stimulation.
Leukemia, 32, 2018
|
|
5OPP
 
 | | Crystal structure of S408R cN-II mutant | | Descriptor: | BROMIDE ION, Cytosolic purine 5'-nucleotidase, GLYCEROL, ... | | Authors: | Hnizda, A, Pachl, P, Rezacova, P. | | Deposit date: | 2017-08-10 | | Release date: | 2018-06-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Relapsed acute lymphoblastic leukemia-specific mutations in NT5C2 cluster into hotspots driving intersubunit stimulation.
Leukemia, 32, 2018
|
|
5OPN
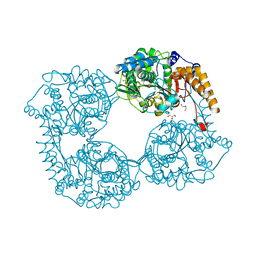 
 | | Crystal structure of R39Q cN-II mutant | | Descriptor: | Cytosolic purine 5'-nucleotidase, GLYCEROL | | Authors: | Hnizda, A, Pachl, P, Rezacova, P. | | Deposit date: | 2017-08-10 | | Release date: | 2018-06-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Relapsed acute lymphoblastic leukemia-specific mutations in NT5C2 cluster into hotspots driving intersubunit stimulation.
Leukemia, 32, 2018
|
|
