-Search query
-Search result
Showing 1 - 50 of 110 items for (author: orlova & ev)

EMDB-14342: 
Six DNA Helix Bundle nanopore - State 1

EMDB-14343: 
Six DNA duplex bundle nanopore - State 2

EMDB-14344: 
Six DNA Helix Bundle nanopore - State 3

EMDB-14345: 
Six DNA Helix Bundle nanopore - State 4

EMDB-14346: 
Six DNA Helix Bundle nanopore - State 5

EMDB-14509: 
gp6/gp15/gp16 connector complex of bacteriophage SPP1

EMDB-13664: 
GA1 bacteriophage portal protein

EMDB-13665: 
PhiCPV4 bacteriophage Portal Protein

PDB-7pv2: 
GA1 bacteriophage portal protein

PDB-7pv4: 
PhiCPV4 bacteriophage Portal Protein

EMDB-12707: 
O-Layer C14 at 2.58A - Local refinement with C14 symmetry of the O-layer of the outer membrane core complex from the fully-assembled R388 type IV secretion system.

EMDB-12708: 
I-Layer C16 at 3.08A - Local refinement with C16 symmetry of I-layer of the outer membrane core complex from the fully-assembled R388 type IV secretion system.

EMDB-12709: 
Stalk C1 at 3.71A - Local refinement without symmetry of the Stalk complex from the fully-assembled R388 type IV secretion system.

EMDB-12715: 
IMC-Arches C6 at 8.33A - Refinement with C6 symmetry of the Inner Membrane Complex (IMC) with the Arches from the fully-assembled R388 type IV secretion system.
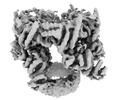
EMDB-12716: 
Trimer of dimers TrwK/VirB4unbound C1 at 4.14A - Refinement without symmetry of TrwK/VirB4unbound trimer of dimers complex (with Hcp1) from the R388 type IV secretion system.

EMDB-12717: 
Dimer TrwK/VirB4unbound C1 at 3.49A - Local refinement without symmetry of the TrwK/VirB4unbound dimer complex from the R388 type IV secretion system.

EMDB-12933: 
IMC protomer C1 at 3.75A - Local refinement without symmetry of the inner membrane complex (IMC) protomer from the fully-assembled R388 type IV secretion system.

EMDB-13765: 
OMCC C1 at 3.28A - Refinement without symmetry of the outer membrane core complex (O- and I-layer) from the fully-assembled R388 type IV secretion system.

EMDB-13766: 
Ab Initio model for IMC-Arches-Stalk from the fully-assembled R388 type IV secretion system determined by cryo-EM.

EMDB-13767: 
IMC-Arches-Stalk C1 at 6.18A - Refinement of the IMC, Arches and Stalk complex without symmetry from the fully-assembled R388 type IV secretion system.

EMDB-13768: 
Stalk C5 at 3.28A - Local refinement with C5 symmetry of the Stalk complex from the fully-assembled R388 type IV secretion system.

EMDB-11852: 
Bovine Papillomavirus E1 DNA helicase-replication fork complex

EMDB-4620: 
A dimer component of alpha-1 antitrypsin heat-induced polymers generated from wild-type M plasma protein and decorated with Fab 4B12

EMDB-4631: 
A dimer component of alpha-1 antitrypsin polymers isolated from ZZ explant liver tissue and decorated with Fab 4B12 (component B with ~90 degree rotation around the dimer axis)

EMDB-4632: 
A dimer component of alpha-1 antitrypsin polymers isolated from ZZ explant liver tissue and decorated with Fab 4B12 (component A with ~60 degree rotation around the dimer axis)

EMDB-10002: 
BACTERIOPHAGE SPP1 PROCAPSID-II PROTEIN

EMDB-4716: 
BACTERIOPHAGE SPP1 MATURE CAPSID PROTEIN

EMDB-4717: 
BACTERIOPHAGE SPP1 PROCAPSID-I PROTEIN

PDB-6ny1: 
CasX-gRNA-DNA(30bp) State II

PDB-6ny2: 
CasX-gRNA-DNA(45bp) state I

PDB-6ny3: 
CasX ternary complex with 30bp target DNA

EMDB-8987: 
ApoCasX

EMDB-8988: 
CasX binary complex

EMDB-8989: 
CasX-gRNA-DNA (45T-20NT) State I

EMDB-8990: 
CasX-gRNA-DNA (45T-20NT) State II

EMDB-8991: 
CasX-gRNA-DNA (45T-45NT) State II

EMDB-8994: 
CasX-gRNA-DNA (30bp) State II

EMDB-8980: 
CasX ternary complex with 30bp target DNA

EMDB-8996: 
CasX-gRNA-DNA(45bp) state I

EMDB-3585: 
Structure of the VirD4 bound to a Type IV secretion system

EMDB-3725: 
Negative-stain EM structure of the A. tumefaciens outer-membrane core complex

EMDB-3077: 
Novel inter-subunit contacts in Barley Stripe Mosaic Virus revealed by cryo-EM

EMDB-3078: 
Novel inter-subunit contacts in Barley Stripe Mosaic Virus revealed by cryo-EM

EMDB-3073: 
Novel inter-subunit contacts in Barley Stripe Mosaic Virus revealed by cryo-EM

EMDB-3074: 
Novel inter-subunit contacts in Barley Stripe Mosaic Virus revealed by cryo-EM

PDB-5a79: 
Novel inter-subunit contacts in Barley Stripe Mosaic Virus revealed by cryo-EM

PDB-5a7a: 
Novel inter-subunit contacts in Barley Stripe Mosaic Virus revealed by cryo-EM

EMDB-3087: 
Structural basis for DNA strand separation by a hexameric replicative helicase

EMDB-3088: 
Structural basis for DNA strand separation by a hexameric replicative helicase

EMDB-3089: 
Structural basis for DNA strand separation by a hexameric replicative helicase
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
