-Search query
-Search result
Showing 1 - 50 of 55 items for (author: marek & m)
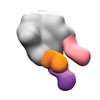
EMDB-40242: 
BG505 MD39 SOSIP in complex with Rh.NJ85 wk12 gp120GH, N611/FP, and base epitope polyclonal antibodies

EMDB-40243: 
BG505 MD39 SOSIP in complex with Rh.NJ86 wk12 V1V3, C3V5, N611/FP, and base epitope polyclonal antibodies

EMDB-40244: 
BG505 MD39 SOSIP in complex with Rh.NJ76 wk12 C3V5, N611/FP, and base epitope polyclonal antibodies
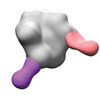
EMDB-40252: 
BG505 MD39 SOSIP in complex with Rh.NK04 wk12 gp120 glycan hole and base epitope polyclonal antibodies

EMDB-40254: 
BG505 MD39 SOSIP in complex with Rh.NJ75 wk12 gp120 glycan hole and base epitope polyclonal antibodies

EMDB-40255: 
BG505 MD39 SOSIP in complex with Rh.NJ87 wk12 C3V5 and base epitope polyclonal antibodies

EMDB-40256: 
BG505 MD39 SOSIP in complex with Rh.NJ84 wk12 V1V3, gp120 glycan hole and base epitope polyclonal antibodies

EMDB-40257: 
BG505 MD39 SOSIP in complex with Rh.NJ77 wk12 V1V3, C3V5, N611/FP and base epitope polyclonal

EMDB-17015: 
Cryo-EM map of the focused refinement of the subfamily III haloalkane dehalogenase from Haloferax mediterranei dimer forming hexameric assembly.

PDB-8ooh: 
Cryo-EM map of the focused refinement of the subfamily III haloalkane dehalogenase from Haloferax mediterranei dimer forming hexameric assembly.

EMDB-16998: 
Cryo-EM structure of subfamily III haloalkane dehalogenase DhmeA from Haloferax mediterranei

EMDB-26380: 
Cryo-EM structure of Shiga toxin 2 in complex with the native ribosomal P-stalk

EMDB-26381: 
Cryo-EM structure of Shiga toxin 2 in complex with the native ribosomal P-stalk

PDB-7u6v: 
Cryo-EM structure of Shiga toxin 2 in complex with the native ribosomal P-stalk
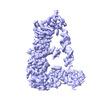
EMDB-14855: 
Mammalian Dicer in the "pre-dicing state" with pre-miR-15a substrate and TARBP2 subunit

PDB-7zpj: 
Mammalian Dicer in the "pre-dicing state" with pre-miR-15a substrate and TARBP2 subunit

EMDB-14383: 
Mammalian Dicer in the "pre-dicing state" with pre-miR-15a substrate
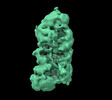
EMDB-14384: 
Mammalian Dicer in the dicing state with pre-miR-15a substrate

EMDB-14387: 
Mouse endoribonuclease Dicer (composite structure)

EMDB-14854: 
Mammalian Dicer in the "dicing state" with pre-miR-15a substrate

EMDB-14856: 
Mammalian Dicer in the "pre-dicing state" with pre-miR-15a substrate and TARBP2 subunit

PDB-7yym: 
Mammalian Dicer in the "pre-dicing state" with pre-miR-15a substrate

PDB-7yyn: 
Mammalian Dicer in the dicing state with pre-miR-15a substrate

PDB-7yz4: 
Mouse endoribonuclease Dicer (composite structure)

PDB-7zpi: 
Mammalian Dicer in the "dicing state" with pre-miR-15a substrate

PDB-7zpk: 
Mammalian Dicer in the "pre-dicing state" with pre-miR-15a substrate and TARBP2 subunit

EMDB-12679: 
Cryo-EM structure (model_1a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.4 A

EMDB-12680: 
Cryo-EM structure (model_2a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.5 A

EMDB-12681: 
Cryo-EM structure of the RC-dLH complex (model_1b) from Gemmatimonas phototrophica at 2.47 A

EMDB-12682: 
Cryo-EM structure (model_2b) of the RC-dLH complex from Gemmatimonas phototrophica at 2.44 A

PDB-7o0u: 
Cryo-EM structure (model_1a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.4 A

PDB-7o0v: 
Cryo-EM structure (model_2a) of the RC-dLH complex from Gemmatimonas phototrophica at 2.5 A

PDB-7o0w: 
Cryo-EM structure of the RC-dLH complex (model_1b) from Gemmatimonas phototrophica at 2.47 A

PDB-7o0x: 
Cryo-EM structure (model_2b) of the RC-dLH complex from Gemmatimonas phototrophica at 2.44 A

EMDB-13344: 
Cryo-EM structure of BMV-derived VLP expressed in E. coli and assembled in the presence of tRNA (tVLP)

EMDB-13345: 
Cryo-EM structure of BMV-derived VLP expressed in E. coli (eVLP)

PDB-7pe1: 
Cryo-EM structure of BMV-derived VLP expressed in E. coli and assembled in the presence of tRNA (tVLP)

PDB-7pe2: 
Cryo-EM structure of BMV-derived VLP expressed in E. coli (eVLP)

EMDB-13192: 
Structure of knob spiral in P. falciparum-infected erythrocytes

EMDB-4898: 
TssA protein from T6SS of Vibrio cholerae.

PDB-6riu: 
C-terminal domain of TssA protein from T6SS of Vibrio cholerae.

EMDB-3878: 
The distal end of a non-contractile T6SS sheath.

EMDB-3879: 
The baseplate of a non-contractile T6SS sheath.

PDB-5ojq: 
The modeled structure of of wild type extended type VI secretion system sheath/tube complex in vibrio cholerae based on cryo-EM reconstruction of the non-contractile sheath/tube complex

EMDB-3563: 
Subtomogram average of type VI secretion system (T6SS) extended sheath

EMDB-3564: 
Subtomogram average of type VI secretion system (T6SS) contracted sheath

EMDB-3566: 
VipA-N3, non-contractile sheath of the type VI secretion system

EMDB-3567: 
VipA-N2/VipB contracted sheath of type VI secretion system

PDB-5mxn: 
Atomic model of the VipA/VipB/Hcp, the type six secretion system non-contractile sheath-tube of Vibrio cholerae from cryo-EM

PDB-5myu: 
VipA-N2/VipB contracted sheath of type VI secretion system
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
