[English] 日本語
 Yorodumi
Yorodumi- EMDB-3564: Subtomogram average of type VI secretion system (T6SS) contracted... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3564 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of type VI secretion system (T6SS) contracted sheath | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology | Type VI secretion system TssC-like / TssC1, N-terminal / TssC1, C-terminal / EvpB/VC_A0108, tail sheath N-terminal domain / EvpB/VC_A0108, tail sheath gpW/gp25-like domain / Type VI secretion system sheath protein TssB1 / Type VI secretion system, VipA, VC_A0107 or Hcp2 / Type VI secretion protein / Type VI secretion system contractile sheath small subunit Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
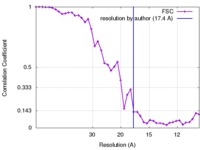
| Method | subtomogram averaging / cryo EM / Resolution: 17.4 Å | |||||||||
 Authors Authors | Wang J / Basler M | |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2017 Journal: Nat Microbiol / Year: 2017Title: Cryo-EM structure of the extended type VI secretion system sheath-tube complex. Authors: Jing Wang / Maximilian Brackmann / Daniel Castaño-Díez / Mikhail Kudryashev / Kenneth N Goldie / Timm Maier / Henning Stahlberg / Marek Basler /   Abstract: The bacterial type VI secretion system (T6SS) uses contraction of a long sheath to quickly thrust a tube with associated effectors across membranes of eukaryotic and bacterial cells . Only limited ...The bacterial type VI secretion system (T6SS) uses contraction of a long sheath to quickly thrust a tube with associated effectors across membranes of eukaryotic and bacterial cells . Only limited structural information is available about the inherently unstable precontraction state of the T6SS. Here, we obtain a 3.7 Å resolution structure of a non-contractile sheath-tube complex using cryo-electron microscopy and show that it resembles the extended T6SS inside Vibrio cholerae cells. We build a pseudo-atomic model of the complete sheath-tube assembly, which provides a mechanistic understanding of coupling sheath contraction with pushing and rotating the inner tube for efficient target membrane penetration. Our data further show that sheath contraction exposes a buried recognition domain to specifically trigger the disassembly and recycling of the T6SS sheath by the cognate ATP-dependent unfoldase ClpV. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3564.map.gz emd_3564.map.gz | 3.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3564-v30.xml emd-3564-v30.xml emd-3564.xml emd-3564.xml | 9.3 KB 9.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3564_fsc.xml emd_3564_fsc.xml | 4.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_3564.png emd_3564.png | 114.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3564 http://ftp.pdbj.org/pub/emdb/structures/EMD-3564 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3564 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3564 | HTTPS FTP |
-Related structure data
| Related structure data |  3563C  3566C  3567C  5mxnC  5myuC  5ojqC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3564.map.gz / Format: CCP4 / Size: 4.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3564.map.gz / Format: CCP4 / Size: 4.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Type VI secretion system contracted sheath
| Entire | Name: Type VI secretion system contracted sheath |
|---|---|
| Components |
|
-Supramolecule #1: Type VI secretion system contracted sheath
| Supramolecule | Name: Type VI secretion system contracted sheath / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: vipA
| Macromolecule | Name: vipA / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Sequence | String: MSKEGSVAPK ERINIKYIPA TGDAQAEVEL PLKTLVVGDF KGHAEQTPLE ERATITVDKN NFEAVMRESE LKITATVKN KLTDDENAEL PVELNFKSLA DFAPDAVASQ VPELKKLIEL REALVALKGP LGNIPAFRER L QSLLNSEE SREKLLAELN LLSGQEEPQA |
-Macromolecule #2: vipB
| Macromolecule | Name: vipB / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Sequence | String: MMSTTEKVLE RPQLAQGSLL DEIMAQTRIA PSEEGYDIAK KGVAAFIENL MGSQHSAEPV NKSLVDQMLV ELDKKISAQ MDEILHNSQF QAMESAWRGL KLFVDRTDFR ENNKVEILHV TKDELLEDFE FAPETAQSGL Y KHVYSAGY GQFGGEPVGA IIGNYAFTPS ...String: MMSTTEKVLE RPQLAQGSLL DEIMAQTRIA PSEEGYDIAK KGVAAFIENL MGSQHSAEPV NKSLVDQMLV ELDKKISAQ MDEILHNSQF QAMESAWRGL KLFVDRTDFR ENNKVEILHV TKDELLEDFE FAPETAQSGL Y KHVYSAGY GQFGGEPVGA IIGNYAFTPS TPDMKLLQYM GALGAMAHAP FISSVGPEFF GIDSFEELPN IK DLKSTFE SPKYTKWRSL RESEDARYLG LTAPRFLLRV PYDPIENPVK SFNYAENVSA SHEHYLWGNT AFA FATRLT DSFAKYRWCP NIIGPQSGGA VEDLPVHVFE SMGALQSKIP TEVLITDRKE FELAEEGFIA LTMR KGSDN AAFFSANSIQ KPKVFPNTKE GKEAETNYKL GTQLPYMMII NRLAHYVKVL QREQIGAWKE RQDLE RELN SWIKQYVADQ ENPPADVRSR RPLRAARIEV MDVEGNPGWY QVSLSVRPHF KYMGANFELS LVGRLD QA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)