2Y0I
 
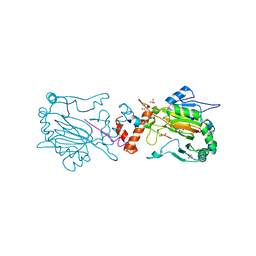 | | FACTOR INHIBITING HIF-1 ALPHA IN COMPLEX WITH TANKYRASE-2 (TNKS2) FRAGMENT PEPTIDE (21-MER) | | Descriptor: | 2-OXOGLUTARIC ACID, FE (II) ION, GLYCEROL, ... | | Authors: | Chowdhury, R, McDonough, M.A, Schofield, C.J. | | Deposit date: | 2010-12-02 | | Release date: | 2011-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Factor-inhibiting hypoxia-inducible factor (FIH) catalyses the post-translational hydroxylation of histidinyl residues within ankyrin repeat domains.
FEBS J., 278, 2011
|
|
2XYP
 
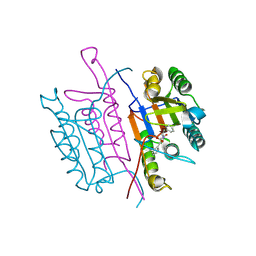 | | Caspase-3:CAS26049945 | | Descriptor: | CASPASE-3 SUBUNIT P12, CASPASE-3 SUBUNIT P17, PHENYLMETHYL N-[(2S)-4-CHLORO-3-OXO-1-PHENYL-BUTAN-2-YL]CARBAMATE | | Authors: | Ganesan, R, Jelakovic, S, Grutter, M.G, Mittl, P.R. | | Deposit date: | 2010-11-18 | | Release date: | 2011-08-17 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | In Silico Identification and Crystal Structure Validation of Caspase-3 Inhibitors without a P1 Aspartic Acid Moiety.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
3FIT
 
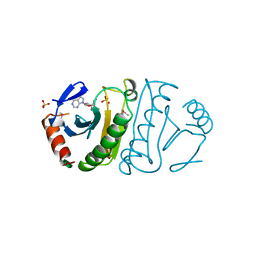 | | FHIT (FRAGILE HISTIDINE TRIAD PROTEIN) IN COMPLEX WITH ADENOSINE/SULFATE AMP ANALOG | | Descriptor: | ADENOSINE MONOPHOSPHATE, FRAGILE HISTIDINE PROTEIN, SULFATE ION, ... | | Authors: | Lima, C.D, D'Amico, K.L, Naday, I, Rosenbaum, G, Westbrook, E.M, Hendrickson, W.A. | | Deposit date: | 1997-05-17 | | Release date: | 1997-11-19 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | MAD analysis of FHIT, a putative human tumor suppressor from the HIT protein family.
Structure, 5, 1997
|
|
3FP8
 
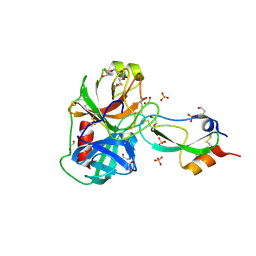 | | Anionic trypsin variant S195A in complex with bovine pancreatic trypsin inhibitor (BPTI) determined to the 1.46 A resolution limit | | Descriptor: | 1,2-ETHANEDIOL, Anionic trypsin-2, CALCIUM ION, ... | | Authors: | Zakharova, E, Horvath, M.P, Goldenberg, D.P, Curtice, K. | | Deposit date: | 2009-01-04 | | Release date: | 2009-02-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Structure of a serine protease poised to resynthesize a peptide bond.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3FPP
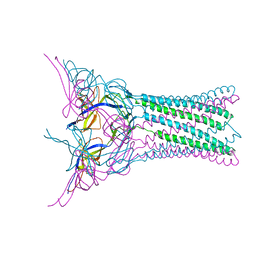 
 | | Crystal structure of E.coli MacA | | Descriptor: | Macrolide-specific efflux protein macA | | Authors: | Yum, S, Xu, Y, Piao, S, Ha, N.-C. | | Deposit date: | 2009-01-06 | | Release date: | 2009-04-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Crystal structure of the periplasmic component of a tripartite macrolide-specific efflux pump
J.Mol.Biol., 387, 2009
|
|
3TBG
 
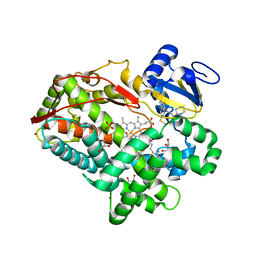 | | Human cytochrome P450 2D6 with two thioridazines bound in active site | | Descriptor: | 10-{2-[(2R)-1-methylpiperidin-2-yl]ethyl}-2-(methylsulfanyl)-10H-phenothiazine, Cytochrome P450 2D6, GLYCEROL, ... | | Authors: | Wang, A, Stout, C.D, Johnson, E.F. | | Deposit date: | 2011-08-05 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Contributions of Ionic Interactions and Protein Dynamics to Cytochrome P450 2D6 (CYP2D6) Substrate and Inhibitor Binding.
J.Biol.Chem., 290, 2015
|
|
3TCS
 
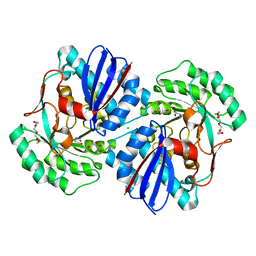 | | Crystal structure of a putative racemase from Roseobacter denitrificans | | Descriptor: | CHLORIDE ION, D-ALANINE, GLYCEROL, ... | | Authors: | Eswaramoorthy, S, Chamala, S, Evans, B, Foti, R, Gizzi, A, Hillerich, B, Kar, A, LaFleur, J, Seidel, R, Villigas, G, Zencheck, W, Almo, S.C, Swaminathan, S, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2011-08-09 | | Release date: | 2011-08-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Crystal structure of a putative racemase from Roseobacter denitrificans
To be Published
|
|
2VKM
 
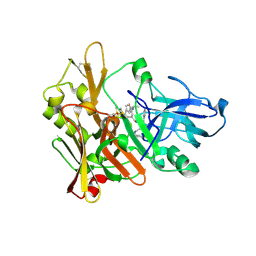 | | Crystal structure of GRL-8234 bound to BACE (Beta-secretase) | | Descriptor: | BETA-SECRETASE 1, N-{(1S,2R)-1-benzyl-2-hydroxy-3-[(3-methoxybenzyl)amino]propyl}-5-[methyl(methylsulfonyl)amino]-N'-[(1R)-1-phenylethyl]benzene-1,3-dicarboxamide | | Authors: | Hong, L, Tang, J, Ghosh, A.K. | | Deposit date: | 2007-12-04 | | Release date: | 2008-12-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Potent Memapsin 2 (Beta-Secretase) Inhibitors: Design, Synthesis, Protein-Ligand X-Ray Structure, and in Vivo Evaluation.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
4FED
 
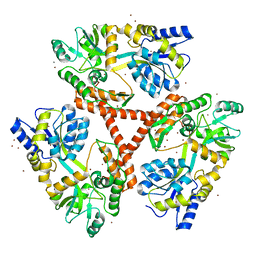 | | Crystal Structure of Htt36Q3H | | Descriptor: | CALCIUM ION, Maltose-binding periplasmic protein,Huntingtin, ZINC ION | | Authors: | Kim, M. | | Deposit date: | 2012-05-30 | | Release date: | 2013-03-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.807 Å) | | Cite: | Beta conformation of polyglutamine track revealed by a crystal structure of Huntingtin N-terminal region with insertion of three histidine residues.
Prion, 7, 2013
|
|
3TDL
 
 | | Structure of human serum albumin in complex with DAUDA | | Descriptor: | 11-({[5-(dimethylamino)naphthalen-1-yl]sulfonyl}amino)undecanoic acid, MYRISTIC ACID, Serum albumin | | Authors: | Wang, Y, Luo, Z, Shi, X, Wang, H, Nie, L. | | Deposit date: | 2011-08-11 | | Release date: | 2012-06-27 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A fluorescent fatty acid probe, DAUDA, selectively displaces two myristates bound in human serum albumin
Protein Sci., 20, 2011
|
|
3H1U
 
 | | Structure of ubiquitin in complex with Cd ions | | Descriptor: | CADMIUM ION, Ubiquitin | | Authors: | Qureshi, I.A, Ferron, F, Cheung, P, Lescar, J. | | Deposit date: | 2009-04-14 | | Release date: | 2009-05-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystallographic structure of ubiquitin in complex with cadmium ions
BMC RES NOTES, 2, 2009
|
|
2VGM
 
 | |
2VIT
 
 | | INFLUENZA VIRUS HEMAGGLUTININ, MUTANT WITH THR 155 REPLACED BY ILE, COMPLEXED WITH A NEUTRALIZING ANTIBODY | | Descriptor: | HEMAGGLUTININ, IMMUNOGLOBULIN (IGG1, LAMBDA), ... | | Authors: | Bizebard, T, Fleury, D, Gigant, B, Wharton, S.A, Skehel, J.J, Knossow, M. | | Deposit date: | 1997-12-22 | | Release date: | 1998-04-29 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Antigen distortion allows influenza virus to escape neutralization.
Nat.Struct.Biol., 5, 1998
|
|
2VGN
 
 | | Structure of S. cerevisiae Dom34, a translation termination-like factor involved in RNA quality control pathways and interacting with Hbs1 (SelenoMet-labeled protein) | | Descriptor: | DOM34, GLYCEROL, PHOSPHATE ION | | Authors: | Graille, M, Chaillet, M, Van Tilbeurgh, H. | | Deposit date: | 2007-11-14 | | Release date: | 2008-01-22 | | Last modified: | 2021-03-17 | | Method: | X-RAY DIFFRACTION (2.505 Å) | | Cite: | Structure of Yeast Dom34: A Protein Related to Translation Termination Factor Erf1 and Involved in No-Go Decay.
J.Biol.Chem., 283, 2008
|
|
2V7D
 
 | |
3THI
 
 | |
2V64
 
 | | Crystallographic structure of the conformational dimer of the Spindle Assembly Checkpoint protein Mad2. | | Descriptor: | MBP1, MITOTIC SPINDLE ASSEMBLY CHECKPOINT PROTEIN MAD2A | | Authors: | Mapelli, M, Massimiliano, L, Santaguida, S, Musacchio, A. | | Deposit date: | 2007-07-13 | | Release date: | 2007-11-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The MAD2 Conformational Dimer: Structure and Implications for the Spindle Assembly Checkpoint
Cell(Cambridge,Mass.), 131, 2007
|
|
3H4S
 
 | |
2V9F
 
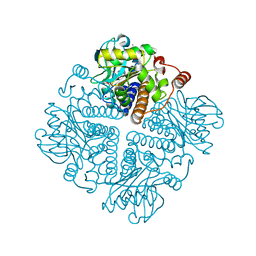 | |
2V9W
 
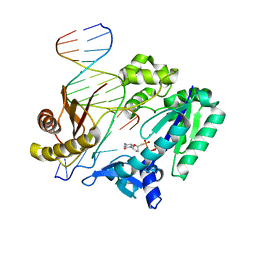 | | Complex structure of Sulfolobus solfataricus DPO4 and DNA duplex containing a hydrophobic thymine isostere 2,4-difluorotoluene nucleotide in the template strand | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, 5'-D(*GP*GP*GP*GP*GP*AP*AP*GP*GP*AP *CP*TP*G)-3', 5'-D(*TP*TP*CP*AP*GP*DFTP*AP*GP*TP*CP*CP *TP*TP*CP*CP*CP*CP*C)-3', ... | | Authors: | Irimia, A, Pallan, P.S, Egli, M. | | Deposit date: | 2007-08-27 | | Release date: | 2007-10-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure and Activity of Y-Class DNA Polymerase Dpo4 from Sulfolobus Solfataricus with Templates Containing the Hydrophobic Thymine Analog 2,4-Difluorotoluene.
J.Biol.Chem., 282, 2007
|
|
2VAN
 
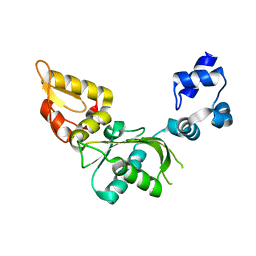 | | Nucleotidyl Transfer Mechanism of Mismatched dNTP Incorporation by DNA Polymerase b by Structural and Kinetic Analyses | | Descriptor: | DNA POLYMERSE BETA | | Authors: | Chan, H, Chou, C, Tang, K, Niebuhr, M, Tung, C, Tsai, M. | | Deposit date: | 2007-09-03 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Mismatched Dntp Incorporation by DNA Polymerase Beta Does not Proceed Via Globally Different Conformational Pathways.
Nucleic Acids Res., 36, 2008
|
|
2UY6
 
 | | Crystal structure of the P pilus rod subunit PapA | | Descriptor: | PAP FIMBRIAL MAJOR PILIN PROTEIN, PERIPLASMID CHAPERONE PAPD PROTEIN | | Authors: | Verger, D, Bullitt, E, Hultgren, S.J, Waksman, G. | | Deposit date: | 2007-04-02 | | Release date: | 2007-05-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of the P Pilus Rod Subunit Papa.
Plos Pathog., 3, 2007
|
|
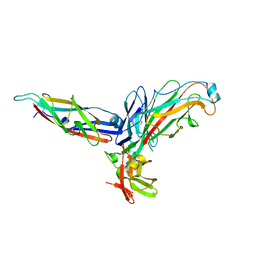
2VDT
 
 | | Crystallographic structure of Levansucrase from Bacillus subtilis mutant S164A | | Descriptor: | CALCIUM ION, LEVANSUCRASE | | Authors: | Ortiz-Soto, M.E, Rivera, M, Rudino-Pinera, E, Olvera, C, Lopez-Munguia, A. | | Deposit date: | 2007-10-11 | | Release date: | 2008-07-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Selected Mutations in Bacillus Subtilis Levansucrase Semi-Conserved Regions Affecting its Biochemical Properties
Protein Eng.Des.Sel., 21, 2008
|
|
2VJV
 
 | | Crystal structure of the IS608 transposase in complex with left end 26-mer DNA hairpin and a 6-mer DNA representing the left end cleavage site | | Descriptor: | 5'-D(*DA*DA*DA*DG*DC*DC*DC*DC*DT*DA*DG*DC*DTP*DT *DT*DT*DA*DG*DC*DT*DA*DT*DG*DG*DG*DGP)-3', 5'-D(*DT*DA*DT*DT*DA*DCP)-3', MAGNESIUM ION, ... | | Authors: | Barabas, O, Ronning, D.R, Guynet, C, Hickman, A.B, Ton-Hoang, B, Chandler, M, Dyda, F. | | Deposit date: | 2007-12-13 | | Release date: | 2008-02-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mechanism of is200/is605 Family DNA Transposases: Activation and Transposon-Directed Target Site Selection.
Cell(Cambridge,Mass.), 132, 2008
|
|
2VAW
 
 | | FtsZ Pseudomonas aeruginosa GDP | | Descriptor: | CELL DIVISION PROTEIN FTSZ, GUANOSINE-5'-DIPHOSPHATE | | Authors: | Oliva, M.A, Lowe, J. | | Deposit date: | 2007-09-04 | | Release date: | 2007-09-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural Insights Into the Conformational Variability of Ftsz
J.Mol.Biol., 373, 2007
|
|
