6LFH
 
 | |
1QUQ
 
 | | COMPLEX OF REPLICATION PROTEIN A SUBUNITS RPA14 AND RPA32 | | Descriptor: | PROTEIN (REPLICATION PROTEIN A 14 KD SUBUNIT), PROTEIN (REPLICATION PROTEIN A 32 KD SUBUNIT) | | Authors: | Bochkarev, A, Bochkareva, E, Frappier, L, Edwards, A.M. | | Deposit date: | 1999-07-02 | | Release date: | 1999-08-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of the complex of replication protein A subunits RPA32 and RPA14 reveals a mechanism for single-stranded DNA binding.
EMBO J., 18, 1999
|
|
5FO5
 
 | | Structure of the DNA-binding domain of Escherichia coli methionine biosynthesis regulator MetR | | Descriptor: | 1,2-ETHANEDIOL, HTH-TYPE TRANSCRIPTIONAL REGULATOR METR, MAGNESIUM ION | | Authors: | Punekar, A.S, Porter, J, Urbanowski, M.L, Stauffer, G.V, Carr, S.B, Phillips, S.E. | | Deposit date: | 2015-11-18 | | Release date: | 2016-06-08 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structural Basis for DNA Recognition by the Transcription Regulator Metr.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
6W30
 
 | | Protein Tyrosine Phosphatase 1B Bound to Amorphadiene | | Descriptor: | Amorphadiene, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Sarkar, A, Kim, E.Y, Hongdusit, A, Sankaran, B, Fox, J. | | Deposit date: | 2020-03-08 | | Release date: | 2021-05-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Microbially Guided Discovery and Biosynthesis of Biologically Active Natural Products.
Acs Synth Biol, 10, 2021
|
|
7ONF
 
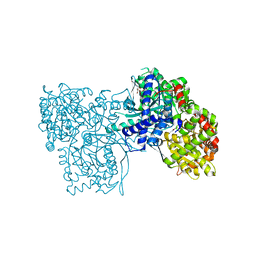 | | The binding of p-coumaroyl glucose to glycogen phosphorylase reveals the relationship between structural data and effects on cell metabolome | | Descriptor: | Glycogen phosphorylase, muscle form, INOSINIC ACID, ... | | Authors: | Tsagkarakou, A.S, Koulas, S.M, Kyriakis, E, Drakou, C.E, Leonidas, D.D. | | Deposit date: | 2021-05-25 | | Release date: | 2022-04-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure activity relationship of the binding of p-coumaroyl glucose to glycogen phosphorylase and its effect on hepatic cell metabolic pathways
Eur J Med Chem Rep, 3, 2021
|
|
1B3T
 
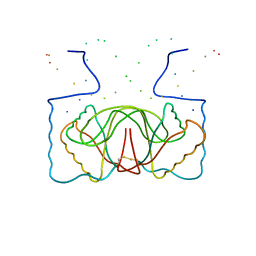 | | EBNA-1 NUCLEAR PROTEIN/DNA COMPLEX | | Descriptor: | DNA (5'-D(*GP*GP*GP*AP*AP*GP*CP*AP*TP*AP*TP*GP*CP*TP*TP*CP*CP*C)-3'), PROTEIN (NUCLEAR PROTEIN EBNA1) | | Authors: | Bochkarev, A, Bochkareva, E, Edwards, A, Frappier, L. | | Deposit date: | 1998-12-14 | | Release date: | 1998-12-15 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The 2.2 A structure of a permanganate-sensitive DNA site bound by the Epstein-Barr virus origin binding protein, EBNA1.
J.Mol.Biol., 284, 1998
|
|
1BCJ
 
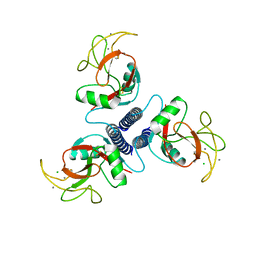 | |
1BCH
 
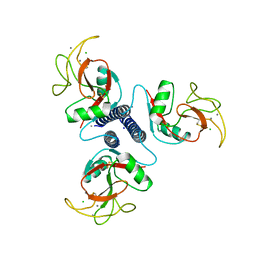 | |
1AFD
 
 | |
1AFB
 
 | |
1AFA
 
 | |
6COM
 
 | | 2.3A crystal structure of E. coli phosphoenolpyruvate carboxykinase mutant Asp269Asn | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, MAGNESIUM ION, ... | | Authors: | Sokaribo, A.S, Cotelesage, J.H, Novakovski, B, Goldie, H, Sanders, D. | | Deposit date: | 2018-03-12 | | Release date: | 2018-04-04 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Kinetic and structural analysis of Escherichia coli phosphoenolpyruvate carboxykinase mutants.
Biochim Biophys Acta Gen Subj, 1864, 2020
|
|
6D2P
 
 | |
6BV9
 
 | | Structure of proteinaceous RNase P 1 (PRORP1) from A. thaliana after overnight soak with juglone | | Descriptor: | 5-hydroxynaphthalene-1,4-dione, CHLORIDE ION, Proteinaceous RNase P 1, ... | | Authors: | Karasik, A, Wu, N, Fierke, C.A, Koutmos, M. | | Deposit date: | 2017-12-12 | | Release date: | 2019-06-12 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Inhibition of protein-only RNase P with Gambogic acid and Juglone
To Be Published
|
|
6BV5
 
 | | Structure of proteinaceous RNase P 1 (PRORP1) from A. thaliana after 45-minute soak with juglone | | Descriptor: | 5-hydroxynaphthalene-1,4-dione, CHLORIDE ION, MANGANESE (II) ION, ... | | Authors: | Karasik, A, Wu, N, Fierke, C.A, Koutmos, M. | | Deposit date: | 2017-12-12 | | Release date: | 2019-06-12 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Inhibition of protein-only RNase P with Gambogic acid and Juglone
To Be Published
|
|
7YC6
 
 | |
6V2M
 
 | |
6V2L
 
 | | E. coli Phosphoenolpyruvate carboxykinase S250A | | Descriptor: | ACETATE ION, ADENOSINE-5'-TRIPHOSPHATE, MANGANESE (II) ION, ... | | Authors: | Sokaribo, A.S, Goldie, H, Sanders, D. | | Deposit date: | 2019-11-25 | | Release date: | 2019-12-11 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Kinetic and structural analysis of Escherichia coli phosphoenolpyruvate carboxykinase mutants.
Biochim Biophys Acta Gen Subj, 1864, 2020
|
|
6BV8
 
 | | Structure of proteinaceous RNase P 1 (PRORP1) from A. thaliana complexed with Mn after 3-hour soak with juglone | | Descriptor: | 5-hydroxynaphthalene-1,4-dione, CHLORIDE ION, MANGANESE (II) ION, ... | | Authors: | Karasik, A, Wu, N, Fierke, C.A, Koutmos, M. | | Deposit date: | 2017-12-12 | | Release date: | 2019-06-12 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Inhibition of protein-only RNase P with Gambogic acid and Juglone
To Be Published
|
|
6BV6
 
 | | Structure of proteinaceous RNase P 1 (PRORP1) from A. thaliana after 3-hour soak with juglone | | Descriptor: | 5-hydroxynaphthalene-1,4-dione, Proteinaceous RNase P 1, chloroplastic/mitochondrial, ... | | Authors: | Karasik, A, Wu, N, Fierke, C.A, Koutmos, M. | | Deposit date: | 2017-12-12 | | Release date: | 2019-06-12 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Inhibition of protein-only RNase P with Gambogic acid and Juglone
To Be Published
|
|
6V2N
 
 | | Crystal structure of E. coli phosphoenolpyruvate carboxykinase mutant Lys254Ser | | Descriptor: | ACETATE ION, CALCIUM ION, Phosphoenolpyruvate carboxykinase (ATP) | | Authors: | Sokaribo, A.S, Cotelesage, J.H, Novakovski, B, Goldie, H, Sanders, D. | | Deposit date: | 2019-11-25 | | Release date: | 2019-12-25 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Kinetic and structural analysis of Escherichia coli phosphoenolpyruvate carboxykinase mutants.
Biochim Biophys Acta Gen Subj, 1864, 2020
|
|
6CRT
 
 | | Arg65Gln Mutagenic E.coli PCK | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MANGANESE (II) ION, Phosphoenolpyruvate carboxykinase (ATP) | | Authors: | Sokaribo, A.S, Cotelesage, J.H, Novakovski, B, Goldie, H, Sanders, D. | | Deposit date: | 2018-03-19 | | Release date: | 2018-04-04 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.995 Å) | | Cite: | Arg65Gln Mutagenic E.coli PCK
To Be Published
|
|
5HT8
 | | Crystal structure of clostrillin double mutant (S17H,S19H) in complex with nickel | | Descriptor: | Beta and gamma crystallin, NICKEL (II) ION | | Authors: | Jamkhindikar, A, Srivastava, S.S, Sankaranarayanan, R. | | Deposit date: | 2016-01-26 | | Release date: | 2017-02-01 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | A Transition Metal-Binding, Trimeric beta gamma-Crystallin from Methane-Producing Thermophilic Archaea, Methanosaeta thermophila
Biochemistry, 56, 2017
|
|
5U2H
 
 | |
5U1V
 | |
