1LZC
 
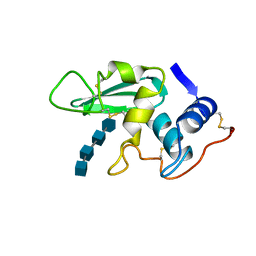 | | DISSECTION OF PROTEIN-CARBOHYDRATE INTERACTIONS IN MUTANT HEN EGG-WHITE LYSOZYME COMPLEXES AND THEIR HYDROLYTIC ACTIVITY | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, HEN EGG WHITE LYSOZYME | | Authors: | Maenaka, K, Matsushima, M, Song, H, Watanabe, K, Kumagai, I. | | Deposit date: | 1995-02-10 | | Release date: | 1995-05-08 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Dissection of protein-carbohydrate interactions in mutant hen egg-white lysozyme complexes and their hydrolytic activity.
J.Mol.Biol., 247, 1995
|
|
2GD8
 
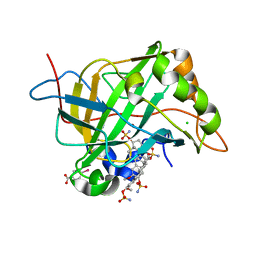 | | Crystal structure analysis of the human carbonic anhydrase II in complex with a 2-substituted estradiol bis-sulfamate | | Descriptor: | (9BETA,13ALPHA,14BETA,17ALPHA)-2-METHOXYESTRA-1,3,5(10)-TRIENE-3,17-DIYL DISULFAMATE, CHLORIDE ION, Carbonic anhydrase 2, ... | | Authors: | De Simone, G, Di Fiore, A. | | Deposit date: | 2006-03-15 | | Release date: | 2007-02-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | 2-substituted estradiol bis-sulfamates, multitargeted antitumor agents: synthesis, in vitro SAR, protein crystallography, and in vivo activity.
J.Med.Chem., 49, 2006
|
|
3L4U
 
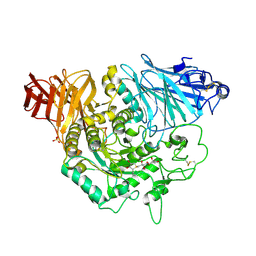 | | Crystal complex of N-terminal Human Maltase-Glucoamylase with de-O-sulfonated kotalanol | | Descriptor: | (2R,3S,4S)-1-[(2S,3S,4R,5R,6S)-2,3,4,5,6,7-hexahydroxyheptyl]-3,4-dihydroxy-2-(hydroxymethyl)tetrahydrothiophenium (non-preferred name), 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Sim, L, Rose, D.R. | | Deposit date: | 2009-12-21 | | Release date: | 2010-02-09 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | New glucosidase inhibitors from an ayurvedic herbal treatment for type 2 diabetes: structures and inhibition of human intestinal maltase-glucoamylase with compounds from Salacia reticulata.
Biochemistry, 49, 2010
|
|
2LL0
 
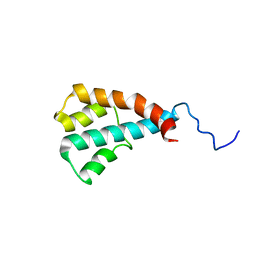 | | NMR structure of the putative ATPase regulatory protein YP_916642.1 from Paracoccus denitrificans | | Descriptor: | Uncharacterized protein | | Authors: | Serrano, P, Geralt, M, Wuthrich, K, Morales-Rios, E, Zarco-Zavala, M, Garcia-Trejo, J.J, Dutta, S.K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2011-10-24 | | Release date: | 2011-12-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the putative ATPase regulatory protein YP_916642.1 from Paracoccus denitrificans
To be Published
|
|
1JZF
 | | Pseudomonas aeruginosa Oxidized Azurin(Cu2+) Ru(tpy)(phen)(His83) | | Descriptor: | (2,2':6',2'-TERPYRIDINE)-(1,10-PHENANTHROLINE) RUTHENIUM (II), AZURIN, COPPER (II) ION, ... | | Authors: | Crane, B.R, Di Bilio, A.J, Winkler, J.R, Gray, H.B. | | Deposit date: | 2001-09-16 | | Release date: | 2001-10-17 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Electron tunneling in single crystals of Pseudomonas aeruginosa azurins.
J.Am.Chem.Soc., 123, 2001
|
|
1TTV
 
 | | NMR Structure of a Complex Between MDM2 and a Small Molecule Inhibitor | | Descriptor: | 1-{[4,5-BIS(4-CHLOROPHENYL)-2-(2-ISOPROPOXY-4-METHOXYPHENYL)-4,5-DIHYDRO-1H-IMIDAZOL-1-YL]CARBONYL}PIPERAZINE, Ubiquitin-protein ligase E3 Mdm2 | | Authors: | Fry, D.C, Emerson, S.D, Palme, S, Vu, B.T, Liu, C.M, Podlaski, F. | | Deposit date: | 2004-06-23 | | Release date: | 2005-01-04 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR structure of a complex between MDM2 and a small molecule inhibitor.
J.Biomol.Nmr, 30, 2004
|
|
2FXK
 
 | | Crystal structure of the macro-domain of human core histone variant macroH2A1.1 (form A) | | Descriptor: | H2A histone family, member Y isoform 1 | | Authors: | Kustatscher, G, Hothorn, M, Pugieux, C, Scheffzek, K, Ladurner, A.G. | | Deposit date: | 2006-02-06 | | Release date: | 2006-02-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Splicing regulates NAD metabolite binding to histone macroH2A.
Nat.Struct.Mol.Biol., 12, 2005
|
|
2C0T
 
 | | Src family kinase Hck with bound inhibitor A-641359 | | Descriptor: | CALCIUM ION, N-(4-{4-AMINO-1-[1-(TETRAHYDRO-2H-PYRAN-4-YL)PIPERIDIN-4-YL]-1H-PYRAZOLO[3,4-D]PYRIMIDIN-3-YL}-2-METHOXYPHENYL)-1-METHYL-1H-INDOLE-2-CARBOXAMIDE, TYROSINE-PROTEIN KINASE HCK | | Authors: | Borhani, D.W, Burchat, A, Calderwood, D.J, Hirst, G.C, Li, B, Loew, A. | | Deposit date: | 2005-09-07 | | Release date: | 2006-09-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Discovery of A-770041, a Src-Family Selective Orally Active Lck Inhibitor that Prevents Organ Allograft Rejection.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
3Q7E
 
 | |
5PON
 
 | | PanDDA analysis group deposition -- Crystal Structure of BRD1 in complex with N10980a | | Descriptor: | 1,2-ETHANEDIOL, 1-[4-(pyridin-2-yl)piperazin-1-yl]ethan-1-one, Bromodomain-containing protein 1, ... | | Authors: | Pearce, N.M, Krojer, T, Talon, R, Bradley, A.R, Fairhead, M, Sethi, R, Wright, N, MacLean, E, Collins, P, Brandao-Neto, J, Douangamath, A, Renjie, Z, Dias, A, Ng, J, Brennan, P.E, Cox, O, Bountra, C, Arrowsmith, C.H, Edwards, A, von Delft, F. | | Deposit date: | 2017-02-07 | | Release date: | 2017-03-15 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.519 Å) | | Cite: | A multi-crystal method for extracting obscured crystallographic states from conventionally uninterpretable electron density.
Nat Commun, 8, 2017
|
|
4B95
 
 | | pVHL-EloB-EloB-EloC complex_(2S,4R)-1-(2-chlorophenyl)carbonyl-N-[(4-chlorophenyl)methyl]-4-oxidanyl-pyrrolidine-2-carboxamide bound | | Descriptor: | (2S,4R)-1-(2-chlorophenyl)carbonyl-N-[(4-chlorophenyl)methyl]-4-oxidanyl-pyrrolidine-2-carboxamide, ACETATE ION, TRANSCRIPTION ELONGATION FACTOR B POLYPEPTIDE 1, ... | | Authors: | Buckley, D.L, Gustafson, J.L, VanMolle, I, Roth, A.G, SeopTae, H, Gareiss, P.C, Jorgensen, W.L, Ciulli, A, Crews, C.M. | | Deposit date: | 2012-08-31 | | Release date: | 2012-10-24 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Small-Molecule Inhibitors of the Interaction between the E3 Ligase Vhl and Hif1Alpha
Angew.Chem.Int.Ed.Engl., 51, 2012
|
|
2C5O
 
 | | Differential Binding Of Inhibitors To Active And Inactive Cdk2 Provides Insights For Drug Design | | Descriptor: | 4-(2,4-DIMETHYL-1,3-THIAZOL-5-YL)PYRIMIDIN-2-AMINE, CELL DIVISION PROTEIN KINASE 2, CYCLIN A2 | | Authors: | Kontopidis, G, McInnes, C, Pandalaneni, S.R, McNae, I, Gibson, D, Mezna, M, Thomas, M, Wood, G, Wang, S, Walkinshaw, M.D, Fischer, P.M. | | Deposit date: | 2005-10-30 | | Release date: | 2006-03-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Differential Binding of Inhibitors to Active and Inactive Cdk2 Provides Insights for Drug Design.
Chem.Biol., 13, 2006
|
|
3A7D
 
 | | Crystal Structures of rat Catechol-O-Methyltransferase complexed with new bi-substrate type inhibitor | | Descriptor: | 5'-deoxy-5'-[4-({[(2,3-dihydroxy-5-nitrophenyl)carbonyl]amino}methyl)-1H-1,2,3-triazol-1-yl]adenosine, Catechol O-methyltransferase, MAGNESIUM ION, ... | | Authors: | Tsuji, E. | | Deposit date: | 2009-09-26 | | Release date: | 2010-09-15 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Hit to Lead: Comprehensive Strategy of de novo Scaffold Generation by FBDD. Part 2: Ligand Fishing using Mass Spectrometry by Detection of Ligand-Protein Non-Covalent Complex after Matrix Click Chemistry
To be Published
|
|
5TJL
 
 | | Crystal structure of GTA + A trisaccharide (mercury derivative) | | Descriptor: | Histo-blood group ABO system transferase, MERCURY (II) ION, alpha-L-fucopyranose-(1-2)-[2-acetamido-2-deoxy-alpha-D-galactopyranose-(1-3)]octyl beta-D-galactopyranoside | | Authors: | Legg, M.S.G, Gagnon, S.M.L, Evans, S.V. | | Deposit date: | 2016-10-04 | | Release date: | 2017-06-14 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | High-resolution crystal structures and STD NMR mapping of human ABO(H) blood group glycosyltransferases in complex with trisaccharide reaction products suggest a molecular basis for product release.
Glycobiology, 27, 2017
|
|
5TJO
 
 | | Crystal structure of GTB + B trisaccharide (mercury derivative) | | Descriptor: | Histo-blood group ABO system transferase, MERCURY (II) ION, alpha-L-fucopyranose-(1-2)-[alpha-D-galactopyranose-(1-3)]octyl beta-D-galactopyranoside | | Authors: | Legg, M.S.G, Gagnon, S.M.L, Evans, S.V. | | Deposit date: | 2016-10-04 | | Release date: | 2017-06-14 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | High-resolution crystal structures and STD NMR mapping of human ABO(H) blood group glycosyltransferases in complex with trisaccharide reaction products suggest a molecular basis for product release.
Glycobiology, 27, 2017
|
|
2HLJ
 
 | |
6LBV
 
 | | Crystal structure of barley exohydrolaseI W434F mutant in complex with methyl 6-thio-beta-gentiobioside | | Descriptor: | ACETATE ION, Beta-D-glucan exohydrolase isoenzyme ExoI, GLYCEROL, ... | | Authors: | Luang, S, Streltsov, V.A, Hrmova, M. | | Deposit date: | 2019-11-15 | | Release date: | 2020-09-30 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | The evolutionary advantage of an aromatic clamp in plant family 3 glycoside exo-hydrolases.
Nat Commun, 13, 2022
|
|
2GFG
 
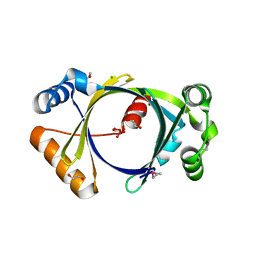 | |
5TLG
 
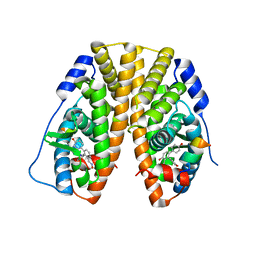 | | Crystal Structure of the ER-alpha Ligand-binding Domain (Y537S) in Complex with (E)-4,4''-dihydroxy-3'-((hydroxyiminio)methyl)-[1,1':2',1''-terphenyl]-4'-olate | | Descriptor: | 2~3~-[(E)-(hydroxyimino)methyl][1~1~,2~1~:2~2~,3~1~-terphenyl]-1~4~,2~4~,3~4~-triol, Estrogen receptor, NUCLEAR RECEPTOR COACTIVATOR 2 | | Authors: | Nwachukwu, J.C, Srinivasan, S, Bruno, N.E, Nowak, J, Kojetin, D.J, Elemento, O, Katzenellenbogen, J.A, Nettles, K.W. | | Deposit date: | 2016-10-11 | | Release date: | 2017-01-18 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.228 Å) | | Cite: | Systems Structural Biology Analysis of Ligand Effects on ER alpha Predicts Cellular Response to Environmental Estrogens and Anti-hormone Therapies.
Cell Chem Biol, 24, 2017
|
|
7FA0
 
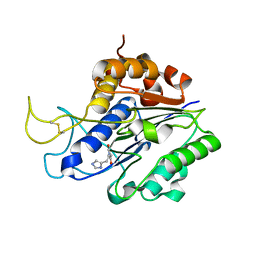 | | The crystal structure of VyPAL2-C214A, a dead mutant of VyPAL2 peptide asparaginyl ligase in form II | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Peptide Asparaginyl Ligases, ... | | Authors: | Hu, S, Sahili, A, Lescar, J. | | Deposit date: | 2021-07-05 | | Release date: | 2022-07-13 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for proenzyme maturation, substrate recognition, and ligation by a hyperactive peptide asparaginyl ligase.
Plant Cell, 34, 2022
|
|
5A7M
 
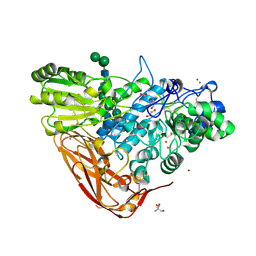 | | The structure of Hypocrea jecorina beta-xylosidase Xyl3A (Bxl1) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mikkelsen, N.E, Gudmundsson, M, Karkehabadi, S, Hansson, H, Sandgren, M, Larenas, E, Mitchinson, C, Keleman, B, Kaper, T. | | Deposit date: | 2015-07-08 | | Release date: | 2016-08-10 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Th Crystal Structure of a Fungal Glycoside Hydrolase Family 3 Beta-Xylosidase, Xyl3A from Hypocrea Jecorina
To be Published
|
|
3QTI
 
 | | c-Met Kinase in Complex with NVP-BVU972 | | Descriptor: | 6-{[6-(1-methyl-1H-pyrazol-4-yl)imidazo[1,2-b]pyridazin-3-yl]methyl}quinoline, CHLORIDE ION, GLYCEROL, ... | | Authors: | Appleton, B.A. | | Deposit date: | 2011-02-22 | | Release date: | 2011-07-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A Drug Resistance Screen Using a Selective MET Inhibitor Reveals a Spectrum of Mutations That Partially Overlap with Activating Mutations Found in Cancer Patients.
Cancer Res., 71, 2011
|
|
3QT0
 
 | | Revealing a steroid receptor ligand as a unique PPARgamma agonist | | Descriptor: | 11-(4-DIMETHYLAMINO-PHENYL)-17-HYDROXY-13-METHYL-17-PROP-1-YNYL-1,2,6,7,8,11,12,13,14,15,16,17-DODEC AHYDRO-CYCLOPENTA[A]PHENANTHREN-3-ONE, Nuclear receptor coactivator 1 peptide, Peroxisome proliferator-activated receptor gamma | | Authors: | Rong, H. | | Deposit date: | 2011-02-22 | | Release date: | 2012-02-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.496 Å) | | Cite: | Revealing a steroid receptor ligand as a unique PPAR gamma agonist.
Cell Res., 22, 2012
|
|
4OJC
 
 | |
3AI0
 | | Crystal structure of beta-glucosidase from termite Neotermes koshunensis in complex with para-nitrophenyl-beta-D-glucopyranoside | | Descriptor: | 4-nitrophenyl beta-D-glucopyranoside, GLYCEROL, beta-glucosidase | | Authors: | Jeng, W.-Y, Liu, C.-I, Wang, A.H.-J. | | Deposit date: | 2010-05-06 | | Release date: | 2010-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural and functional analysis of three beta-glucosidases from bacterium Clostridium cellulovorans, fungus Trichoderma reesei and termite Neotermes koshunensis
J.Struct.Biol., 173, 2011
|
|
