6WAQ
 
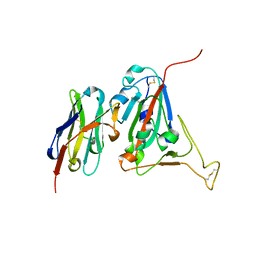 | |
6WAR
 
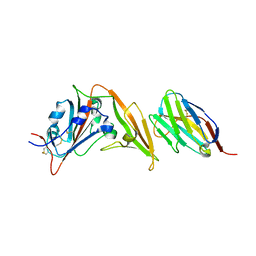 | |
6U7K
 
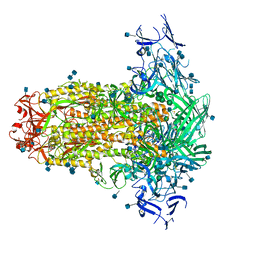 | | Prefusion structure of PEDV spike | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Wrapp, D, McLellan, J.S. | | Deposit date: | 2019-09-03 | | Release date: | 2019-09-18 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | The 3.1-Angstrom Cryo-electron Microscopy Structure of the Porcine Epidemic Diarrhea Virus Spike Protein in the Prefusion Conformation.
J.Virol., 93, 2019
|
|
6VSB
 
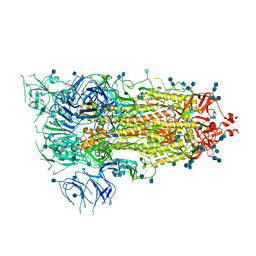 | | Prefusion 2019-nCoV spike glycoprotein with a single receptor-binding domain up | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein | | Authors: | Wrapp, D, Wang, N, Corbett, K.S, Goldsmith, J.A, Hsieh, C, Abiona, O, Graham, B.S, McLellan, J.S. | | Deposit date: | 2020-02-10 | | Release date: | 2020-02-26 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.46 Å) | | Cite: | Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation.
Science, 367, 2020
|
|
8EU8
 
 | | Cryo-EM structure of CH848 10.17DT DS-SOSIP-2P Env | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CH848 10.17DT SOSIP Envelope glycoprotein gp160 | | Authors: | Wrapp, D, Acharya, P, Haynes, B.F. | | Deposit date: | 2022-10-18 | | Release date: | 2023-01-04 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (3.73 Å) | | Cite: | Structure-Based Stabilization of SOSIP Env Enhances Recombinant Ectodomain Durability and Yield.
J.Virol., 97, 2023
|
|
7KBB
 | |
7KBA
 
 | |
7M1C
 
 | |
7LYV
 
 | |
7M30
 
 | |
7M22
 
 | |
7LYW
 
 | |
6XKL
 
 | | SARS-CoV-2 HexaPro S One RBD up | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein | | Authors: | Wrapp, D, Hsieh, C.-L, Goldsmith, J.A, McLellan, J.S. | | Deposit date: | 2020-06-26 | | Release date: | 2020-07-15 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.21 Å) | | Cite: | Structure-based design of prefusion-stabilized SARS-CoV-2 spikes.
Science, 369, 2020
|
|
9OFP
 
 | | HCoV-229E S2P bound by two DH1533 Fabs | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DH1533 heavy chain, ... | | Authors: | Wrapp, D. | | Deposit date: | 2025-04-30 | | Release date: | 2025-12-03 | | Last modified: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | A neutralizing human antibody induces movement of the HCoV-229E receptor binding domain.
Commun Biol, 9, 2025
|
|
9OFQ
 
 | | HCoV-229E S2P bound by one DH1533 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DH1533 FAB heavy chain, ... | | Authors: | Wrapp, D. | | Deposit date: | 2025-04-30 | | Release date: | 2025-12-03 | | Last modified: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | A neutralizing human antibody induces movement of the HCoV-229E receptor binding domain.
Commun Biol, 9, 2025
|
|
9OFO
 
 | | HCoV-229E S2P bound by three DH1533 Fabs | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DH1533 FAB heavy chain, ... | | Authors: | Wrapp, D. | | Deposit date: | 2025-04-30 | | Release date: | 2025-12-03 | | Last modified: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (2.82 Å) | | Cite: | A neutralizing human antibody induces movement of the HCoV-229E receptor binding domain.
Commun Biol, 9, 2025
|
|
6UOE
 
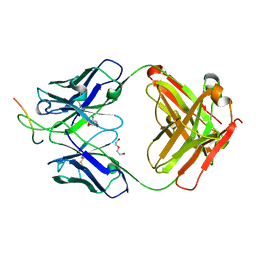 | |
8DF1
 
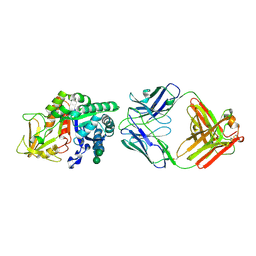 | | Chi3l1 bound by antibody C59 | | Descriptor: | C59 Fab Heavy chain, C59 Fab Light chain, Chitinase-3-like protein 1, ... | | Authors: | Wrapp, D, McLellan, J.S. | | Deposit date: | 2022-06-21 | | Release date: | 2023-03-01 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | A novel humanized Chi3l1 blocking antibody attenuates acetaminophen-induced liver injury in mice.
Antib Ther, 6, 2023
|
|
5TP3
 
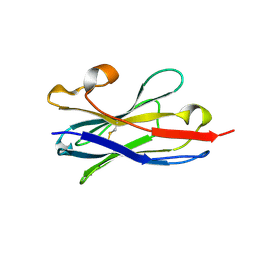 | |
9NW7
 
 | | CA117v2v8 Fab bound to HLA-E-VL9 | | Descriptor: | Beta-2-microglobulin, CA117v2v8 heavy chain, CA117v2v8 light chain, ... | | Authors: | Wrapp, D, Hwang, J.K, Marston, D.J, Haynes, B.F, Azoitei, M.L. | | Deposit date: | 2025-03-21 | | Release date: | 2026-03-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | HLA-E-VL9 antibodies mediate NKG2A+ NK cell and CD8+ T cell killing of HIV-infected CD4+ T cells
To Be Published
|
|
9NW9
 
 | | CA117v2v8 Fab bound to HLA-E-RL9HIV | | Descriptor: | Beta-2-microglobulin, CA117v2v8 heavy chain, CA117v2v8 light chain, ... | | Authors: | Wrapp, D, Hwang, J.K, Marston, D.J, Haynes, B.F, Azoitei, M.L. | | Deposit date: | 2025-03-21 | | Release date: | 2026-03-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | HLA-E-VL9 antibodies mediate NKG2A+ NK cell and CD8+ T cell killing of HIV-infected CD4+ T cells
To Be Published
|
|
9NW6
 
 | | CA147v24 Fab bound to HLA-E-VL9 | | Descriptor: | Beta-2-microglobulin, CA147v24 heavy chain, CA147v24 light chain, ... | | Authors: | Wrapp, D, Hwang, J.K, Marston, D.J, Haynes, B.F, Azoitei, M.L. | | Deposit date: | 2025-03-21 | | Release date: | 2026-03-25 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | HLA-E-VL9 antibodies mediate NKG2A+ NK cell and CD8+ T cell killing of HIV-infected CD4+ T cells
To Be Published
|
|
9NW8
 
 | | CA117v2v8 Fab bound to HLA-E-Mtb44 | | Descriptor: | Beta-2-microglobulin, CA117v2v8 heavy chain, CA117v2v8 light chain, ... | | Authors: | Wrapp, D, Hwang, J.K, Marston, D.J, Haynes, B.F, Azoitei, M.L. | | Deposit date: | 2025-03-21 | | Release date: | 2026-03-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | HLA-E-VL9 antibodies mediate NKG2A+ NK cell and CD8+ T cell killing of HIV-infected CD4+ T cells
To Be Published
|
|
9NW5
 
 | | 3H4v31 anti-HLA-E-VL9 Fab | | Descriptor: | 3H4v31 heavy chain, 3H4v31 light chain | | Authors: | Wrapp, D, Hwang, J.K, Marston, D.J, Haynes, B.F, Azoitei, M.L. | | Deposit date: | 2025-03-21 | | Release date: | 2026-03-25 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | HLA-E-VL9 antibodies mediate NKG2A+ NK cell and CD8+ T cell killing of HIV-infected CD4+ T cells
To Be Published
|
|
6APD
 
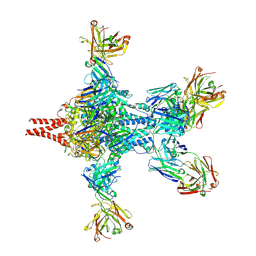 | | Crystal structure of RSV F bound by AM22 and the infant antibody ADI-19425 | | Descriptor: | AM22 Fab Heavy Chain,IGH@ protein, AM22 Fab Light Chain,Uncharacterized protein, Fusion glycoprotein F0,Envelope glycoprotein, ... | | Authors: | Wrapp, D, McLellan, J.S. | | Deposit date: | 2017-08-17 | | Release date: | 2018-03-21 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (4.1 Å) | | Cite: | Infants Infected with Respiratory Syncytial Virus Generate Potent Neutralizing Antibodies that Lack Somatic Hypermutation.
Immunity, 48, 2018
|
|
