3DY7
 
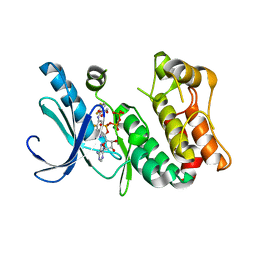 | | X-ray structure of the human mitogen-activated protein kinase kinase 1 (MEK1) in a complex with ligand and MgATP | | Descriptor: | (5S)-4,5-difluoro-6-[(2-fluoro-4-iodophenyl)imino]-N-(2-hydroxyethoxy)cyclohexa-1,3-diene-1-carboxamide, ADENOSINE-5'-TRIPHOSPHATE, Dual specificity mitogen-activated protein kinase kinase 1, ... | | Authors: | Ohren, J.F, Pavlovsky, A, Zhang, E. | | Deposit date: | 2008-07-25 | | Release date: | 2009-06-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Beyond the MEK-pocket: can current MEK kinase inhibitors be utilized to synthesize novel type III NCKIs? Does the MEK-pocket exist in kinases other than MEK?
Bioorg.Med.Chem.Lett., 19, 2009
|
|
8IS5
 
 | |
8IS4
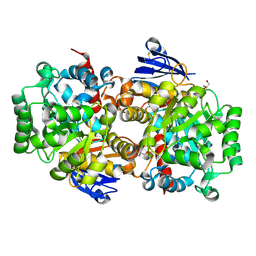 
 | | Structure of an Isocytosine specific deaminase Vcz in complexed with 5-FU | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 5-FLUOROURACIL, GLYCEROL, ... | | Authors: | Guo, W.T, Li, X.J, Wu, B.X. | | Deposit date: | 2023-03-20 | | Release date: | 2023-08-30 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural characterization of an isocytosine-specific deaminase VCZ reveals its application potential in the anti-cancer therapy.
Iscience, 26, 2023
|
|
2B52
 
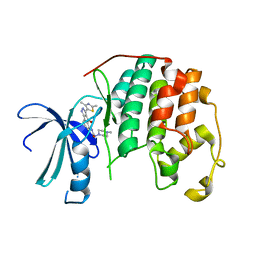 | | Human cyclin dependent kinase 2 (CDK2) complexed with DPH-042562 | | Descriptor: | 1-(3-(2,4-DIMETHYLTHIAZOL-5-YL)-4-OXO-2,4-DIHYDROINDENO[1,2-C]PYRAZOL-5-YL)-3-(4-METHYLPIPERAZIN-1-YL)UREA, Cell division protein kinase 2 | | Authors: | Muckelbauer, J. | | Deposit date: | 2005-09-27 | | Release date: | 2005-10-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Synthesis and Evaluation of Indenopyrazoles as Cyclin-Dependent Kinase Inhibitors. Part 4: Heterocycles at C3
Bioorg.Med.Chem.Lett., 14, 2004
|
|
5WYS
 
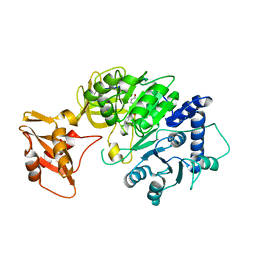 | | luciferase with inhibitor 3i | | Descriptor: | 5-[(3R)-3-(4-boranylphenyl)-3-oxidanyl-propyl]-2-oxidanyl-benzoic acid, Luciferin 4-monooxygenase | | Authors: | Gu, L, Su, J, Wang, F. | | Deposit date: | 2017-01-15 | | Release date: | 2017-12-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.999 Å) | | Cite: | Inhibiting Firefly Bioluminescence by Chalcones
Anal. Chem., 89, 2017
|
|
2B55
 
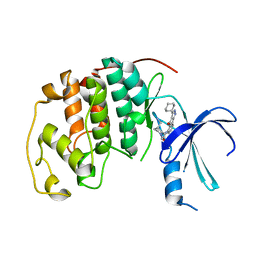 | | Human cyclin dependent kinase 2 (cdk2) complexed with indenopyraxole DIN-101312 | | Descriptor: | 2-(4-(AMINOMETHYL)PIPERIDIN-1-YL)-N-(3_CYCLOHEXYL-4-OXO-2,4-DIHYDROINDENO[1,2-C]PYRAZOL-5-YL)ACETAMIDE, Cell division protein kinase 2 | | Authors: | Muckelbauer, J. | | Deposit date: | 2005-09-27 | | Release date: | 2005-10-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Synthesis and Evaluation of Indenopyrazoles as Cyclin-Dependent Kinase Inhibitors. 3. Structure Activity Relationships at C3
J.Med.Chem., 45, 2002
|
|
2B53
 
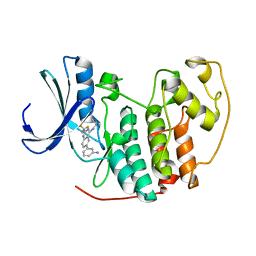 | |
2DRJ
 
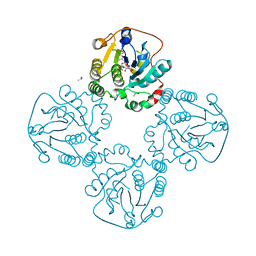 | |
2F2H
 
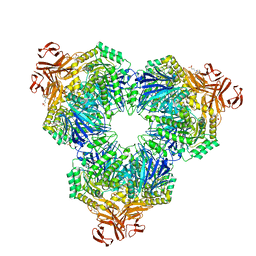 | | Structure of the YicI thiosugar Michaelis complex | | Descriptor: | 3[N-MORPHOLINO]PROPANE SULFONIC ACID, 4-NITROPHENYL 6-THIO-6-S-ALPHA-D-XYLOPYRANOSYL-BETA-D-GLUCOPYRANOSIDE, GLYCEROL, ... | | Authors: | Kim, Y.-W, Lovering, A.L, Strynadka, N.C.J, Withers, S.G. | | Deposit date: | 2005-11-16 | | Release date: | 2006-02-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Expanding the Thioglycoligase Strategy to the Synthesis of alpha-linked Thioglycosides Allows Structural Investigation of the Parent Enzyme/Substrate Complex
J.Am.Chem.Soc., 128, 2006
|
|
4C7T
 
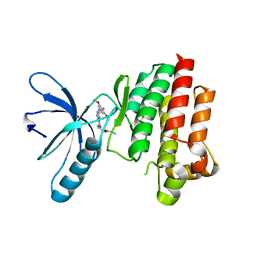 | | Focal Adhesion Kinase catalytic domain in complex with a diarylamino- 1,3,5-triazine inhibitor | | Descriptor: | FOCAL ADHESION KINASE 1, N-methyl-2-[[4-[(3,4,5-trimethoxyphenyl)amino]-1,3,5-triazin-2-yl]amino]benzamide, SULFATE ION | | Authors: | Le Coq, J, Lietha, D. | | Deposit date: | 2013-09-25 | | Release date: | 2014-04-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Inhibition of Both Focal Adhesion Kinase and Fibroblast Growth Factor Receptor 2 Pathways Induces Anti-Tumor and Anti-Angiogenic Activities.
Cancer Lett., 348, 2014
|
|
4BRX
 
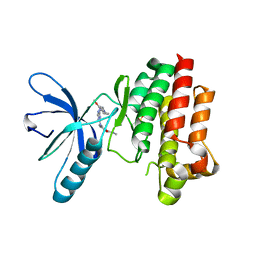 | | Focal Adhesion Kinase catalytic domain in complex with a diarylamino- 1,3,5-triazine inhibitor | | Descriptor: | 2-(4-(2-methoxy-4-morpholinophenylamino)-1,3,5-triazin-2-ylamino)-N-methylbenzamide, FOCAL ADHESION KINASE 1, SULFATE ION | | Authors: | Le Coq, J, Lietha, D. | | Deposit date: | 2013-06-05 | | Release date: | 2013-07-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Synthesis of Novel Diarylamino-1,3,5-Triazine Derivatives as Fak Inhibitors with Anti-Angiogenic Activity.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
8JNI
 
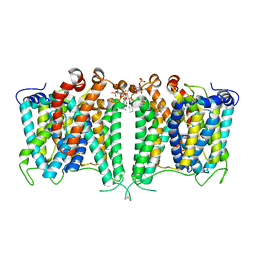 | | Structure of AE2 in complex with PIP2 | | Descriptor: | Anion exchange protein 2, CHLORIDE ION, [(2R)-1-octadecanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phospho ryl]oxy-propan-2-yl] (8Z)-icosa-5,8,11,14-tetraenoate | | Authors: | Yin, Y.X, Ding, D. | | Deposit date: | 2023-06-06 | | Release date: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structural and functional insights into the lipid regulation of human anion exchanger 2.
Nat Commun, 15, 2024
|
|
8JNJ
 
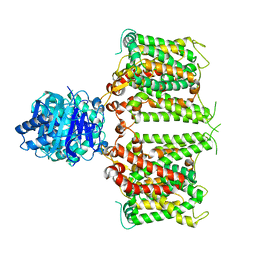 | | Structure of R932A/K1147A/H1148A mutant AE2 | | Descriptor: | Anion exchange protein 2 | | Authors: | Yin, Y.X, Ding, D. | | Deposit date: | 2023-06-06 | | Release date: | 2024-02-07 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structural and functional insights into the lipid regulation of human anion exchanger 2.
Nat Commun, 15, 2024
|
|
8KA3
 
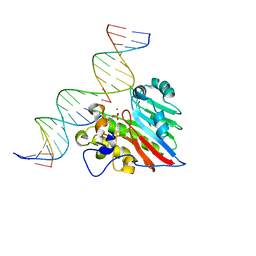 | |
8KA5
 
 | |
8KA4
 
 | |
7VWV
 
 | |
8SLX
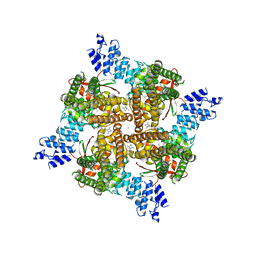 
 | | Rat TRPV2 bound with 1 CBD ligand in nanodiscs | | Descriptor: | SODIUM ION, Transient receptor potential cation channel subfamily V member 2, cannabidiol | | Authors: | Tan, X, Swartz, K.J. | | Deposit date: | 2023-04-24 | | Release date: | 2023-05-31 | | Last modified: | 2023-07-05 | | Method: | ELECTRON MICROSCOPY (3.23 Å) | | Cite: | Cannabidiol sensitizes TRPV2 channels to activation by 2-APB.
Elife, 12, 2023
|
|
8SLY
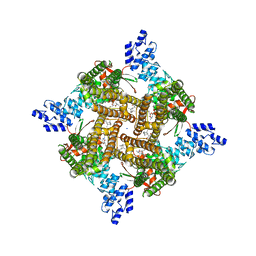 
 | | Rat TRPV2 bound with 2 CBD ligands in nanodiscs | | Descriptor: | (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, SODIUM ION, Transient receptor potential cation channel subfamily V member 2, ... | | Authors: | Tan, X, Swartz, K.J. | | Deposit date: | 2023-04-24 | | Release date: | 2023-05-31 | | Last modified: | 2023-07-05 | | Method: | ELECTRON MICROSCOPY (3.32 Å) | | Cite: | Cannabidiol sensitizes TRPV2 channels to activation by 2-APB.
Elife, 12, 2023
|
|
1C9B
 
 | |
7XS8
 
 | |
7XSA
 
 | |
7XSB
 
 | |
7XSC
 
 | |
3M37
 
 | |
