8IU0
 
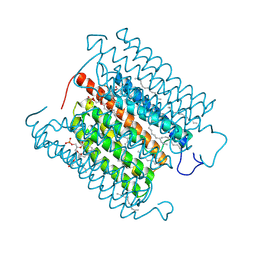 | | Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 H225F mutant in lipid nanodisc | | Descriptor: | (7R,17E,20E)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSA-17,20-DIEN-1-AMINIUM 4-OXIDE, HcKCR1, PALMITIC ACID, ... | | Authors: | Tajima, S, Kim, Y, Nakamura, S, Yamashita, K, Fukuda, M, Deisseroth, K, Kato, H.E. | | Deposit date: | 2023-03-23 | | Release date: | 2023-09-06 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (2.66 Å) | | Cite: | Structural basis for ion selectivity in potassium-selective channelrhodopsins.
Cell, 186, 2023
|
|
8Y6Y
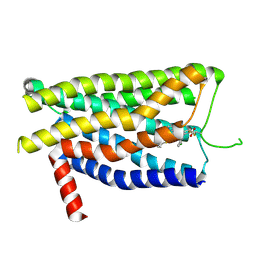 
 | | GLPG0974-bound human FFA2 | | Descriptor: | 4-[[(2~{R})-1-(1-benzothiophen-3-ylcarbonyl)-2-methyl-azetidin-2-yl]carbonyl-[(3-chlorophenyl)methyl]amino]butanoic acid, Free fatty acid receptor 2,Soluble cytochrome b562 | | Authors: | Kugawa, M, Kawakami, K, Kise, R, Kobayashi, K, Kojima, A, Inoue, W, Fukuda, M, Inoue, A, Kato, H.E. | | Deposit date: | 2024-02-03 | | Release date: | 2025-04-02 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (3.36 Å) | | Cite: | Structural insights into lipid chain-length selectivity and allosteric regulation of FFA2.
Nat Commun, 16, 2025
|
|
8Y6W
 
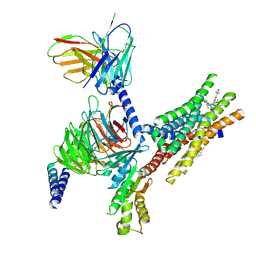 | | TUG-1375 and 4-CMTB-bound human FFA2 in complex with Gi | | Descriptor: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, (2~{S})-2-(4-chlorophenyl)-3-methyl-~{N}-(1,3-thiazol-2-yl)butanamide, Free fatty acid receptor 2, ... | | Authors: | Kugawa, M, Kawakami, K, Kise, R, Kobayashi, K, Kojima, A, Inoue, W, Fukuda, M, Inoue, A, Kato, H.E. | | Deposit date: | 2024-02-03 | | Release date: | 2025-04-02 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (3.19 Å) | | Cite: | Structural insights into lipid chain-length selectivity and allosteric regulation of FFA2.
Nat Commun, 16, 2025
|
|
7W9W
 
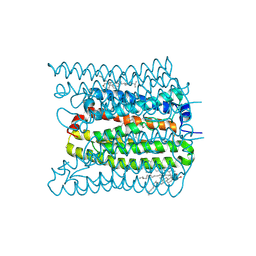 | | 2.02 angstrom cryo-EM structure of the pump-like channelrhodopsin ChRmine | | Descriptor: | CHOLESTEROL, ChRmine, PALMITIC ACID, ... | | Authors: | Kishi, K.E, Kim, Y, Fukuda, M, Yamashita, K, Deisseroth, K, Kato, H.E. | | Deposit date: | 2021-12-11 | | Release date: | 2022-02-02 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (2.02 Å) | | Cite: | Structural basis for channel conduction in the pump-like channelrhodopsin ChRmine.
Cell, 185, 2022
|
|
7UTZ
 
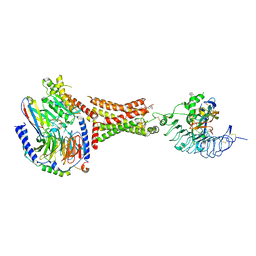 | | Human thyrotropin analog TR1402 bound to human Thyrotropin receptor in complex with miniGs399 (composite structure) | | Descriptor: | (2S)-3-hydroxypropane-1,2-diyl dihexadecanoate, 2-acetamido-2-deoxy-beta-D-glucopyranose, Glycoprotein hormones alpha chain analog TR1402, ... | | Authors: | Faust, B, Cheng, Y, Manglik, A. | | Deposit date: | 2022-04-28 | | Release date: | 2022-08-03 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (2.4 Å) | | Cite: | Autoantibody mimicry of hormone action at the thyrotropin receptor.
Nature, 609, 2022
|
|
8E0G
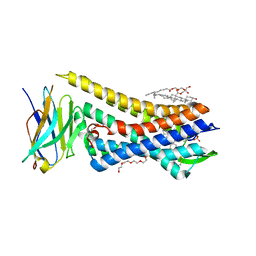 
 | |
9BQJ
 
 | | RO76 bound muOR-Gi1-scFv16 complex structure | | Descriptor: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Guanine nucleotide-binding protein G(i) subunit alpha-1, ... | | Authors: | Wang, H, Majumdar, S, Kobilka, B.K. | | Deposit date: | 2024-05-10 | | Release date: | 2024-09-11 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Signaling Modulation Mediated by Ligand Water Interactions with the Sodium Site at mu OR.
Acs Cent.Sci., 10, 2024
|
|
9B65
 
 | | Biased agonist bound CB1-Gi structure | | Descriptor: | Cannabinoid receptor 1, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Rangari, V.A, O'Brien, E.S, Kobilka, B.K, Krishna Kumar, K, Majumdar, S. | | Deposit date: | 2024-03-23 | | Release date: | 2025-03-05 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (3.03 Å) | | Cite: | A cryptic pocket in CB1 drives peripheral and functional selectivity.
Nature, 640, 2025
|
|
9B54
 
 | | Biased agonist bound CB1-Gi structure | | Descriptor: | Cannabinoid receptor 1, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Rangari, V.A, O'Brien, E.S, Kobilka, B.K, Krishna Kumar, K, Majumdar, S. | | Deposit date: | 2024-03-22 | | Release date: | 2025-03-05 | | Last modified: | 2025-04-16 | | Method: | ELECTRON MICROSCOPY (2.86 Å) | | Cite: | A cryptic pocket in CB1 drives peripheral and functional selectivity.
Nature, 640, 2025
|
|
7YIT
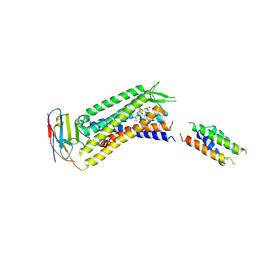 
 | |
5TCX
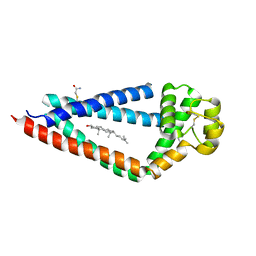 
 | | Crystal structure of human tetraspanin CD81 | | Descriptor: | CD81 antigen, CHOLESTEROL | | Authors: | Zimmerman, B, McMillan, B.J, Seegar, T.C.M, Kruse, A.C, Blacklow, S.C. | | Deposit date: | 2016-09-16 | | Release date: | 2016-11-09 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.955 Å) | | Cite: | Crystal Structure of a Full-Length Human Tetraspanin Reveals a Cholesterol-Binding Pocket.
Cell, 167, 2016
|
|
7N98
 
 | | Cryo-EM structure of MFSD2A | | Descriptor: | Sodium-dependent lysophosphatidylcholine symporter 1 | | Authors: | Zhang, J, Feng, L. | | Deposit date: | 2021-06-17 | | Release date: | 2021-08-04 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structure and mechanism of blood-brain-barrier lipid transporter MFSD2A.
Nature, 596, 2021
|
|
8GHV
 
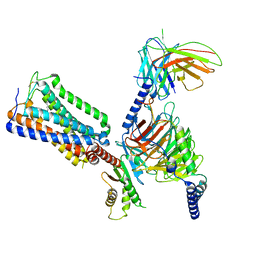 | | Cannabinoid Receptor 1-G Protein Complex | | Descriptor: | (5Z,8Z,11Z,13S,14Z)-N-[(2R)-1-hydroxypropan-2-yl]-13-methylicosa-5,8,11,14-tetraenamide, Cannabinoid receptor 1, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Krishna Kumar, K, Robertson, M.J, Skiniotis, G, Kobilka, B.K. | | Deposit date: | 2023-03-12 | | Release date: | 2023-05-24 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Structural basis for activation of CB1 by an endocannabinoid analog.
Nat Commun, 14, 2023
|
|
8FX5
 
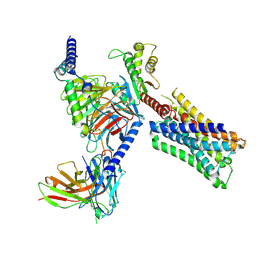 | | Human M4 muscarinic acetylcholine receptor complex with Gi1 and xanomeline | | Descriptor: | Antibody fragment scFv16, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Vuckovic, Z, Mobbs, J.I, Glukhova, A, Sexton, P.M, Danev, R, Thal, D.M. | | Deposit date: | 2023-01-23 | | Release date: | 2023-09-13 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (2.45 Å) | | Cite: | Xanomeline displays concomitant orthosteric and allosteric binding modes at the M 4 mAChR.
Nat Commun, 14, 2023
|
|
8H86
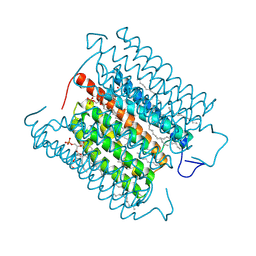 
 | | Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 in lipid nanodisc | | Descriptor: | (7R,17E,20E)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSA-17,20-DIEN-1-AMINIUM 4-OXIDE, HcKCR1, PALMITIC ACID, ... | | Authors: | Tajima, S, Kim, Y, Yamashita, K, Fukuda, M, Deisseroth, K, Kato, H.E. | | Deposit date: | 2022-10-21 | | Release date: | 2023-09-06 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (2.56 Å) | | Cite: | Structural basis for ion selectivity in potassium-selective channelrhodopsins.
Cell, 186, 2023
|
|
8H87
 
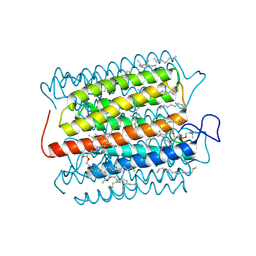 | | Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR2 in lipid nanodisc | | Descriptor: | (7R,17E,20E)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSA-17,20-DIEN-1-AMINIUM 4-OXIDE, HcKCR2, PALMITIC ACID, ... | | Authors: | Tajima, S, Kim, Y, Yamashita, K, Fukuda, M, Deisseroth, K, Kato, H.E. | | Deposit date: | 2022-10-21 | | Release date: | 2023-09-06 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (2.53 Å) | | Cite: | Structural basis for ion selectivity in potassium-selective channelrhodopsins.
Cell, 186, 2023
|
|
7TB3
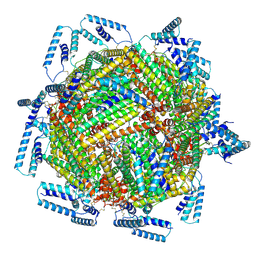 
 | | cryo-EM structure of MBP-KIX-apoferritin | | Descriptor: | Isoform 2 of CREB-binding protein,Ferritin heavy chain, N-terminally processed | | Authors: | Zhang, K, Horikoshi, N, Li, S, Powers, A, Hameedi, M, Pintilie, G, Chae, H, Khan, Y, Suomivuori, C, Dror, R, Sakamoto, K, Chiu, W, Wakatsuki, S. | | Deposit date: | 2021-12-21 | | Release date: | 2022-03-16 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (2.57 Å) | | Cite: | Cryo-EM, Protein Engineering, and Simulation Enable the Development of Peptide Therapeutics against Acute Myeloid Leukemia.
Acs Cent.Sci., 8, 2022
|
|
7TBH
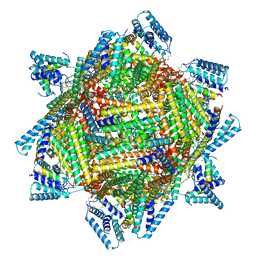 
 | | cryo-EM structure of MBP-KIX-apoferritin complex with peptide 7 | | Descriptor: | Isoform 2 of CREB-binding protein,Ferritin heavy chain, N-terminally processed, LEU-SER-ARG-ARG-PRO-SEP-TYR-ARG-LYS-ILE-LEU-ASN-ASP-LEU-SER-SER-ASP-ALA-PRO | | Authors: | Zhang, K, Horikoshi, N, Li, S, Powers, A, Hameedi, M, Pintilie, G, Chae, H, Khan, Y, Suomivuori, C, Dror, R, Sakamoto, K, Chiu, W, Wakatsuki, S. | | Deposit date: | 2021-12-22 | | Release date: | 2022-03-16 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Cryo-EM, Protein Engineering, and Simulation Enable the Development of Peptide Therapeutics against Acute Myeloid Leukemia.
Acs Cent.Sci., 8, 2022
|
|
4XT3
 
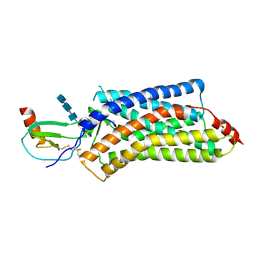 | | Structure of a viral GPCR bound to human chemokine CX3CL1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Fractalkine, G-protein coupled receptor homolog US28, ... | | Authors: | Burg, J.S, Jude, K.M, Waghray, D, Garcia, K.C. | | Deposit date: | 2015-01-22 | | Release date: | 2015-03-04 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3.801 Å) | | Cite: | Structural biology. Structural basis for chemokine recognition and activation of a viral G protein-coupled receptor.
Science, 347, 2015
|
|
4XT1
 
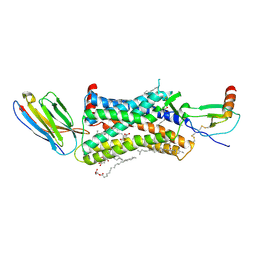 | | Structure of a nanobody-bound viral GPCR bound to human chemokine CX3CL1 | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, CHOLESTEROL, Fractalkine, ... | | Authors: | Burg, J.S, Jude, K.M, Waghray, D, Garcia, K.C. | | Deposit date: | 2015-01-22 | | Release date: | 2015-03-04 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.886 Å) | | Cite: | Structural biology. Structural basis for chemokine recognition and activation of a viral G protein-coupled receptor.
Science, 347, 2015
|
|
5UHS
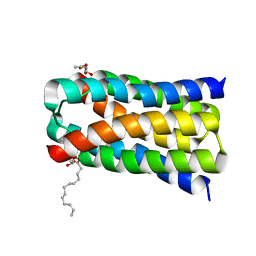 
 | | Structure of a SemiSWEET D57A mutant | | Descriptor: | MONOVACCENIN, Sugar transporter SemiSWEET, beta-D-glucopyranose | | Authors: | Fastman, N.M, Feng, L. | | Deposit date: | 2017-01-11 | | Release date: | 2017-08-02 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Mechanism of Substrate Translocation in an Alternating Access Transporter.
Cell, 169, 2017
|
|
5UHQ
 
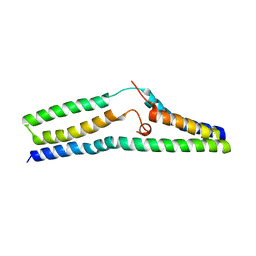 | | Structure of a SemiSWEET Q20A mutant | | Descriptor: | Sugar transporter SemiSWEET | | Authors: | Fastman, N.M, Feng, L. | | Deposit date: | 2017-01-11 | | Release date: | 2017-08-02 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.78 Å) | | Cite: | Mechanism of Substrate Translocation in an Alternating Access Transporter.
Cell, 169, 2017
|
|
9MNY
 
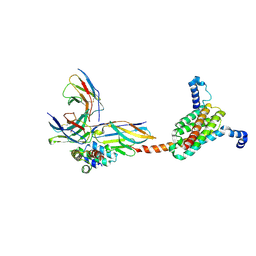 | | Cryo-EM structure of human MPC with pyruvate | | Descriptor: | Fab_8D3_2 heavy chain, Fab_8D3_2 light chain, MBP-PrA/G, ... | | Authors: | Zhang, J, He, Z, Feng, L. | | Deposit date: | 2024-12-24 | | Release date: | 2025-03-05 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Structure of mitochondrial pyruvate carrier and its inhibition mechanism.
Nature, 641, 2025
|
|
9MO0
 
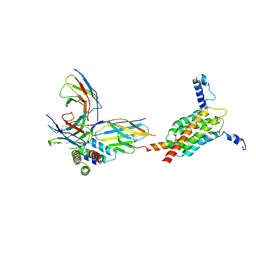 | | Cryo-EM structure of human MPC in complex with AKOS005153046 | | Descriptor: | (~{E})-2-cyano-3-[5-(2-nitrophenyl)furan-2-yl]prop-2-enoic acid, Fab_8D3_2 heavy chain, Fab_8D3_2 light chain, ... | | Authors: | Zhang, J, He, Z, Feng, L. | | Deposit date: | 2024-12-24 | | Release date: | 2025-03-05 | | Last modified: | 2025-05-14 | | Method: | ELECTRON MICROSCOPY (2.83 Å) | | Cite: | Structure of mitochondrial pyruvate carrier and its inhibition mechanism.
Nature, 641, 2025
|
|
9MNW
 
 | | Cryo-EM structure of human MPC in complex with GW604714 | | Descriptor: | (5E)-5-({(5P)-5-[6-(4-acetylpiperazin-1-yl)-3-nitropyridin-2-yl]-2-fluorophenyl}methylidene)-1,3-thiazolidine-2,4-dione, Fab_8D3_2 heavy chain, Fab_8D3_2 light chain, ... | | Authors: | Zhang, J, Feng, L. | | Deposit date: | 2024-12-24 | | Release date: | 2025-03-05 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.35 Å) | | Cite: | Structure of mitochondrial pyruvate carrier and its inhibition mechanism.
Nature, 641, 2025
|
|
