6UUH
 
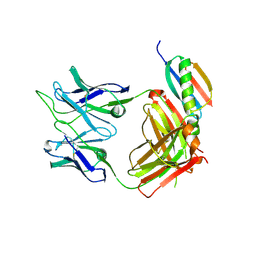 | |
7Y3O
 
 | | Crystal structure of SARS-CoV-2 receptor binding domain in complex with human antibody BIOLS56 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of BIOLS56, Light chain of BIOLS56, ... | | Authors: | Rao, X, Gao, F, Wu, Y, Gao, F. | | Deposit date: | 2022-06-11 | | Release date: | 2023-12-27 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Defining a de novo non-RBM antibody as RBD-8 and its synergistic rescue of immune-evaded antibodies to neutralize Omicron SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
7Y3N
 
 | | Crystal structure of SARS-CoV receptor binding domain in complex with human antibody BIOLS56 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of BIOLS56, ... | | Authors: | Rao, X, Chai, Y, Wu, Y, Gao, F. | | Deposit date: | 2022-06-11 | | Release date: | 2023-12-27 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.97 Å) | | Cite: | Defining a de novo non-RBM antibody as RBD-8 and its synergistic rescue of immune-evaded antibodies to neutralize Omicron SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8QJJ
 
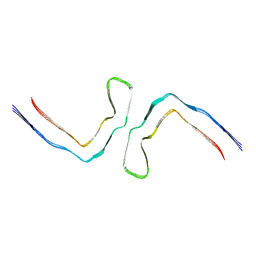 | | CTE type II (tau intermediate amyloid) | | Descriptor: | Isoform Tau-D of Microtubule-associated protein tau | | Authors: | Lovestam, S, Scheres, S.H.W. | | Deposit date: | 2023-09-13 | | Release date: | 2023-10-18 | | Last modified: | 2024-01-17 | | Method: | ELECTRON MICROSCOPY (3.35 Å) | | Cite: | Disease-specific tau filaments assemble via polymorphic intermediates.
Nature, 625, 2024
|
|
9IK9
 
 | | Cryo-EM Structure of SST analogs bond SSTR1-Gi complex | | Descriptor: | (4J2)(DCY)(DTY)(DTR)K(DVA)(DCY)(ALO)(NH2), Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Wong, T.S, Zeng, Z.C, Xiong, T.T, Gan, S.Y, Du, Y. | | Deposit date: | 2024-06-26 | | Release date: | 2025-04-30 | | Last modified: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (3.37 Å) | | Cite: | Structural insights into the binding modes of lanreotide and pasireotide with somatostatin receptor 1.
Acta Pharm Sin B, 15, 2025
|
|
9IK8
 
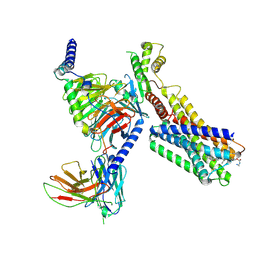 | | Cryo-EM Structure of SSTR1-Gi SST analogs complex | | Descriptor: | DTR-LYS-TY5-PHA-A1D5E-004, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Wong, T.S, Zeng, Z.C, Xiong, T.T, Gan, S.Y, Du, Y. | | Deposit date: | 2024-06-26 | | Release date: | 2025-04-30 | | Last modified: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (2.82 Å) | | Cite: | Structural insights into the binding modes of lanreotide and pasireotide with somatostatin receptor 1.
Acta Pharm Sin B, 15, 2025
|
|
4B61
 | | In meso structure of alginate transporter, AlgE, from Pseudomoas aeruginosa, PAO1. Crystal form 3. | | Descriptor: | (2R)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, (2S)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, ACETATE ION, ... | | Authors: | Tan, J, Pye, V.E, Aragao, D, Caffrey, M. | | Deposit date: | 2012-08-08 | | Release date: | 2013-07-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.402 Å) | | Cite: | A Conformational Landscape for Alginate Secretion Across the Outer Membrane of Pseudomonas Aeruginosa.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4AZL
 
 | | In meso structure of alginate transporter, AlgE, from Pseudomoas aeruginosa, PAO1, crystal form 2. | | Descriptor: | (2R)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, (2S)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, ALGINATE PRODUCTION PROTEIN ALGE, ... | | Authors: | Tan, J, Pye, V.E, Aragao, D, Caffrey, M. | | Deposit date: | 2012-06-26 | | Release date: | 2013-07-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A Conformational Landscape for Alginate Secretion Across the Outer Membrane of Pseudomonas Aeruginosa.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
7XST
 
 | |
7XMX
 
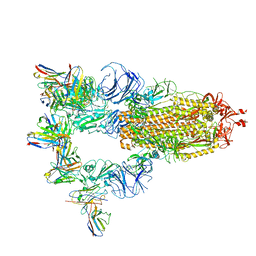
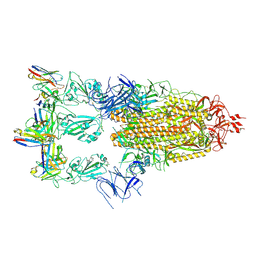 | | Cryo-EM structure of SARS-CoV-2 spike glycoprotein in complex with three F61 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, F61 heavy chain, F61 light chain, ... | | Authors: | Wang, X, Li, X. | | Deposit date: | 2022-04-27 | | Release date: | 2022-11-23 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.62 Å) | | Cite: | Structural basis of a two-antibody cocktail exhibiting highly potent and broadly neutralizing activities against SARS-CoV-2 variants including diverse Omicron sublineages.
Cell Discov, 8, 2022
|
|
6V6W
 
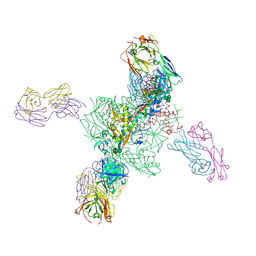 | |
4AFK
 
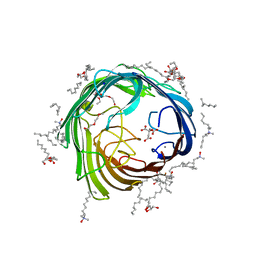 | | In meso structure of alginate transporter, AlgE, from Pseudomonas aeruginosa, PAO1 | | Descriptor: | (2R)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, (2S)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, 3,6,9,12,15,18,21,24-OCTAOXAHEXACOSAN-1-OL, ... | | Authors: | Tan, J, Pye, V.E, Aragao, D, Caffrey, M. | | Deposit date: | 2012-01-19 | | Release date: | 2013-02-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.897 Å) | | Cite: | A Conformational Landscape for Alginate Secretion Across the Outer Membrane of Pseudomonas Aeruginosa.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
9JCV
 
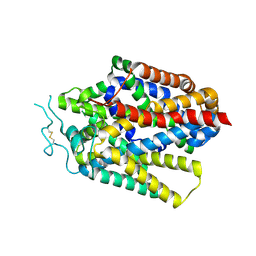 | | Cryo-EM structure of human TauT in the apo state, determined in an inward-facing open conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, Sodium- and chloride-dependent taurine transporter | | Authors: | Qu, Q, Chao, Y, Zhou, Z. | | Deposit date: | 2024-08-30 | | Release date: | 2025-03-19 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (3.41 Å) | | Cite: | Transport and inhibition mechanism for human TauT-mediated taurine uptake.
Cell Res., 35, 2025
|
|
6UTK
 
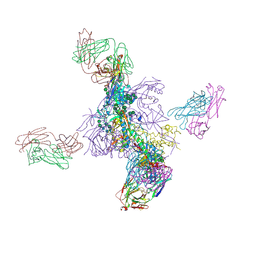 | |
9JD6
 
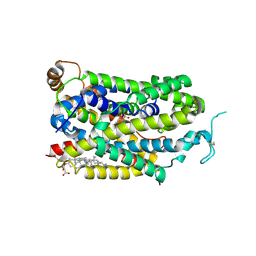 | | Cryo-EM structure of human TauT in presence of Piperidine-4-sulfonate, determined in an inward-facing occluded conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, CHOLESTEROL HEMISUCCINATE, ... | | Authors: | Qu, Q, Chao, Y, Zhou, Z. | | Deposit date: | 2024-08-30 | | Release date: | 2025-03-19 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Transport and inhibition mechanism for human TauT-mediated taurine uptake.
Cell Res., 35, 2025
|
|
9JD5
 
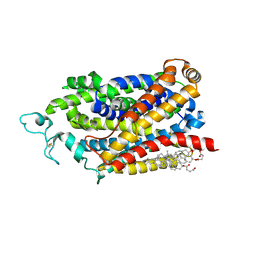 | | Cryo-EM structure of human TauT in presence of Taurocyamine, determined in an inward-facing occluded conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, CHOLESTEROL HEMISUCCINATE, ... | | Authors: | Qu, Q, Chao, Y, Zhou, Z. | | Deposit date: | 2024-08-30 | | Release date: | 2025-03-19 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (2.58 Å) | | Cite: | Transport and inhibition mechanism for human TauT-mediated taurine uptake.
Cell Res., 35, 2025
|
|
9JCZ
 | | Cryo-EM structure of human TauT in presence of taurine, determined in an inward-facing occluded conformation | | Descriptor: | 2-AMINOETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Qu, Q, Chao, Y, Zhou, Z. | | Deposit date: | 2024-08-30 | | Release date: | 2025-03-19 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (2.64 Å) | | Cite: | Transport and inhibition mechanism for human TauT-mediated taurine uptake.
Cell Res., 35, 2025
|
|
9JLN
 
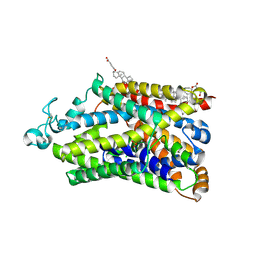 | | Cryo-EM structure of human TauT in presence of taurine, observed only one sodium ion, determined in an inward-occluded conformation | | Descriptor: | 2-AMINOETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Qu, Q, Chao, Y, Zhou, Z. | | Deposit date: | 2024-09-19 | | Release date: | 2025-03-19 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Transport and inhibition mechanism for human TauT-mediated taurine uptake.
Cell Res., 35, 2025
|
|
8UFD
 
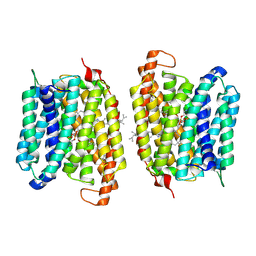 | | Multidrug efflux pump MtEfpA bound with inhibitor BRD8000.3 | | Descriptor: | (1S,3S)-N-[6-bromo-5-(pyrimidin-2-yl)pyridin-2-yl]-2,2-dimethyl-3-(2-methylpropyl)cyclopropane-1-carboxamide, Integral membrane efflux protein EFPA, PHOSPHATIDYLETHANOLAMINE | | Authors: | Wang, S, Liao, M. | | Deposit date: | 2023-10-04 | | Release date: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Structures of the Mycobacterium tuberculosis efflux pump EfpA reveal the mechanisms of transport and inhibition.
Nat Commun, 15, 2024
|
|
8UFE
 
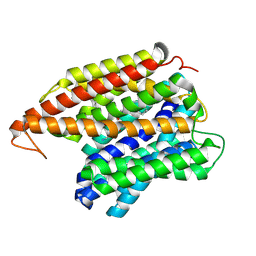 | | Multidrug efflux pump EfpA from mycobacterium smegmatis | | Descriptor: | Integral membrane efflux protein EfpA, PHOSPHATIDYLETHANOLAMINE | | Authors: | Wang, S, Liao, M. | | Deposit date: | 2023-10-04 | | Release date: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (3.68 Å) | | Cite: | Structures of the Mycobacterium tuberculosis efflux pump EfpA reveal the mechanisms of transport and inhibition.
Nat Commun, 15, 2024
|
|
8IM6
 
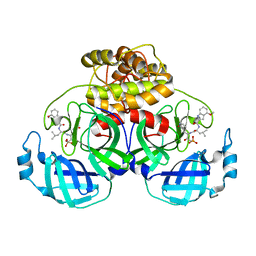 | | Crystal structure of HCoV 229E main protease in complex with PF07304814 | | Descriptor: | 3C-like proteinase, [(3~{S})-3-[[(2~{S})-2-[(4-methoxy-1~{H}-indol-2-yl)carbonylamino]-4-methyl-pentanoyl]amino]-2-oxidanylidene-4-[(3~{R})-2-oxidanylidene-3,4-dihydropyrrol-3-yl]butyl] dihydrogen phosphate | | Authors: | Zhou, Y.R, Zeng, P, Zhang, J, Li, J. | | Deposit date: | 2023-03-06 | | Release date: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structural basis of main proteases of HCoV-229E bound to inhibitor PF-07304814 and PF-07321332.
Biochem.Biophys.Res.Commun., 657, 2023
|
|
7OTJ
 
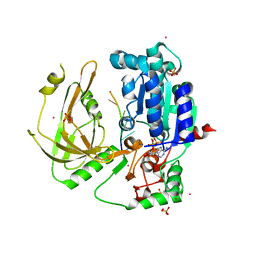 | | Crystal structure of Pif1 helicase from Candida albicans | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ATP-dependent DNA helicase PIF1, DNA (5'-D(P*TP*TP*TP*TP*TP*T)-3'), ... | | Authors: | Rety, S, Xi, X.G. | | Deposit date: | 2021-06-10 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Structural study of the function of Candida Albicans Pif1.
Biochem.Biophys.Res.Commun., 567, 2021
|
|
7TEI
 
 | |
7TF8
 
 | |
7THT
 
 | | CryoEM structure of SARS-CoV-2 S protein in complex with Receptor Binding Domain antibody DH1042 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DH1042 heavy chain, ... | | Authors: | Manne, K, May, A, Acharya, P. | | Deposit date: | 2022-01-12 | | Release date: | 2022-02-16 | | Last modified: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (3.42 Å) | | Cite: | Structural diversity of the SARS-CoV-2 Omicron spike.
Mol.Cell, 82, 2022
|
|
