-Search query
-Search result
Showing 1 - 50 of 64 items for (author: remaut & h)

EMDB-52871: 
Ex vivo spore-silk A-ENA fibers
Method: helical / : Sleutel M, Sogues A, Remaut H

EMDB-53314: 
3D cryoEM map of the BSAP-1 and B1RS complex
Method: single particle / : Pasveer EL, Remaut HK

EMDB-55504: 
Nonameric Ena1C ring of Bacillus cereus
Method: single particle / : Sleutel M, Remaut H

EMDB-48780: 
Cryo-EM of spore appendage from Anaerovoracaceae
Method: helical / : Bellis NF, Baquero DP, Egelman EH, Krupovic M, Wang F

PDB-9n0b: 
Cryo-EM of spore appendage from Anaerovoracaceae
Method: helical / : Bellis NF, Baquero DP, Egelman EH, Krupovic M, Wang F

EMDB-44403: 
Cryo-EM of Hyper2 tube, ~27 nm diameter
Method: helical / : Sonani RR, Miller JG, Conticello V, Egelman EH

EMDB-44404: 
Cryo-EM of Hyper2 tube, ~24 nm diameter
Method: helical / : Sonani RR, Miller JG, Conticello V, Egelman EH

PDB-9bab: 
Cryo-EM of Hyper2 tube, ~27 nm diameter
Method: helical / : Sonani RR, Miller JG, Conticello V, Egelman EH

PDB-9bac: 
Cryo-EM of Hyper2 tube, ~24 nm diameter
Method: helical / : Sonani RR, Miller JG, Conticello V, Egelman EH

EMDB-50893: 
Cryo-EM structure of LptDE-YedD complex from Escherichia Coli
Method: single particle / : Nguyen VS, Remaut H, Collet JF, Gennaris A, Thouvenel L

PDB-9fz5: 
Cryo-EM structure of LptDE-YedD complex from Escherichia Coli
Method: single particle / : Nguyen VS, Remaut H, Collet JF, Gennaris A, Thouvenel L

EMDB-52523: 
CryoEM structure of F-ENA fibers on the spores of Bacillus thuringiensis serovar kurstaki
Method: helical / : Sleutel M, Remaut H

EMDB-52650: 
Recombinant F-ENA-2 fibers
Method: helical / : Sleutel M, Remaut H

PDB-9hze: 
CryoEM structure of F-ENA fibers on the spores of Bacillus thuringiensis serovar kurstaki
Method: helical / : Sleutel M, Remaut H

EMDB-52564: 
Recombinant Ena2A fibers
Method: helical / : Sleutel M, Remaut H

EMDB-51935: 
Ex vivo cannulae fiber structure of Pyrodictium abyssi
Method: helical / : Sleutel M, Remaut H

PDB-9h8b: 
Ex vivo cannulae fiber structure of Pyrodictium abyssi
Method: helical / : Sleutel M, Remaut H

EMDB-45459: 
Structure of in vitro assembled B. anthracis S-layer protein Sap
Method: subtomogram averaging / : Leigh KE, Van der Verren SE, Remaut H, Kudryashev M

PDB-9g93: 
CryoET structure of the in vitro grown Bacillus anthracis Sap S-layer
Method: subtomogram averaging / : Sogues A, Leigh K, Van der Verren S, Kudryashev M, Pak A, Halingstad EV, Cecil AJ, Fioravanti A, Remaut H

EMDB-51414: 
Surface-layer (S-layer) PS2 protein from Corynebacterium glutamicum
Method: single particle / : Sogues A, Remaut H, Sleutel M
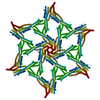
PDB-9gk2: 
Surface-layer (S-layer) PS2 protein from Corynebacterium glutamicum
Method: single particle / : Sogues A, Remaut H, Sleutel M

EMDB-17579: 
Ex vivo L-ENA type pili from Bacillus paranthracis endospores
Method: helical / : Sleutel M, Remaut H

EMDB-18560: 
SARS-CoV-2 S protein bound to human neutralising antibody UZGENT_G5
Method: single particle / : Remaut H, Reiter D, Vandenkerckhove L, Acar DD, Witkowski W, Gerlo S

EMDB-18571: 
SARS-CoV-2 S protein bound to neutralising antibody UZGENT_A3
Method: single particle / : Remaut H, Reiter D, Vandenkerckhove L, Acar DD, Witkowski W, Gerlo S

PDB-8qpr: 
SARS-CoV-2 S protein bound to human neutralising antibody UZGENT_G5
Method: single particle / : Remaut H, Reiter D, Vandenkerckhove L, Acar DD, Witkowski W, Gerlo S

PDB-8qq0: 
SARS-CoV-2 S protein bound to neutralising antibody UZGENT_A3
Method: single particle / : Remaut H, Reiter D, Vandenkerckhove L, Acar DD, Witkowski W, Gerlo S

EMDB-17627: 
Recombinant Ena3A L-Type endospore appendages
Method: helical / : Sleutel M, Remaut H

EMDB-16431: 
Pontibacter korlensis curli subunit CsgA
Method: helical / : Remaut H, Sleutel M, Pradhan B

PDB-8c50: 
Pontibacter korlensis curli subunit CsgA
Method: helical / : Remaut H, Sleutel M, Pradhan B

EMDB-26546: 
Cryo-EM of self-assembled cannula CanA
Method: helical / : Wang F, Miller JG, Egelman EH, Conticello VP

PDB-7uii: 
Cryo-EM of self-assembled cannula CanA
Method: helical / : Wang F, Miller JG, Egelman EH, Conticello VP

EMDB-14352: 
Structure of the GroEL chaperonin in complex with the CnoX chaperedoxin
Method: single particle / : Van der Verren SE, Remaut H

PDB-7ywy: 
Structure of the GroEL chaperonin in complex with the CnoX chaperedoxin
Method: single particle / : Van der Verren SE, Remaut H, Collet JF, Dupuy E

EMDB-12950: 
Cryo-EM structure of undecameric SlyB from Escherichia coli K12
Method: single particle / : Nguyen VS, Remaut H

PDB-7ojg: 
CRYO-EM STRUCTURE OF UNDECAMERIC SLYB FROM ESCHERICHIA COLI K12
Method: single particle / : Nguyen VS, Remaut H

PDB-7ojf: 
CRYO-EM STRUCTURE OF SLYB13-BAMA FROM ESCHERICHIA COLI.
Method: single particle / : Nguyen VS, Remaut H

EMDB-12945: 
Cryo-EM reconstruction of decameric SlyB from Escherichia coli K12
Method: single particle / : Nguyen VS, Remaut H

EMDB-12946: 
Cryo-EM reconstruction of dodecameric SlyB from Escherichia coli K12
Method: single particle / : Nguyen VS, Remaut H

EMDB-12947: 
Cryo-EM structure of tridecameric SlyB from Escherichia coli K12
Method: single particle / : Nguyen VS, Remaut H

EMDB-12948: 
Cryo-EM reconstruction of SlyB14-BamA from Escherichia coli
Method: single particle / : Nguyen VS, Remaut H

EMDB-11610: 
CryoEM structure of a beta3K279T GABA(A)R homomer in complex with megabody MbNbF3c7HopQ
Method: single particle / : Uchanski T, Masiulis S

EMDB-4542: 
CryoEM structure of a beta3K279T GABA(A)R homomer in complex with histamine and megabody Mb25
Method: single particle / : Uchanski T, Masiulis S

PDB-6qfa: 
CryoEM structure of a beta3K279T GABA(A)R homomer in complex with histamine and megabody Mb25
Method: single particle / : Uchanski T, Masiulis S, Fischer B, Kalichuk V, Wohlkoening A, Zoegg T, Remaut H, Vranken W, Aricescu AR, Pardon E, Steyaert J

EMDB-11591: 
Bacillus endospore appendages form a novel family of disulfide-linked pili
Method: helical / : Pradhan B, Liedtke J

EMDB-11592: 
3D cryoEM map of ex vivo Ena1 from Bacillus cereus
Method: helical / : Pradhan B, Sleutel M, Liedtke J, Lindback T, Llarena AK, Brynildsrud O, Aspholm M, Remaut H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search








 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
