-Search query
-Search result
Showing 1 - 50 of 79 items for (author: li & wf)

EMDB-75195: 
S305I Frontotemporal Lobar Degeneration (FTLD) type I tau filament
Method: helical / : Pan HS, Merz GE, Tse E, Southworth DR

EMDB-75196: 
S305I Frontotemporal Lobar Degeneration (FTLD) type II tau filament
Method: helical / : Pan HS, Merz GE, Tse E, Southworth DR

EMDB-49161: 
CryoEM structure of U7 Sm Ring in complex with symplekin N-terminal domain
Method: single particle / : Desotell A, Tong L

EMDB-49206: 
CRYO-EM STRUCTURE OF human U7 SNRNP WITH MUTANT LSM11 that disrupts contacts with CPSF73
Method: single particle / : Desotell A, Tong L

EMDB-49389: 
CRYO-EM STRUCTURE OF HUMAN U7 SNRNP WITH METHYLATED H2A* SUBSTRATE PRE-MRNA FOCUS MAP
Method: single particle / : Desotell A, Tong L

EMDB-49402: 
CRYO-EM STRUCTURE OF HUMAN U7 SNRNP WITH METHYLATED noncleavable H2A* SUBSTRATE PRE-MRNA (core region)
Method: single particle / : Desotell A, Tong L

EMDB-49403: 
CRYO-EM STRUCTURE OF HUMAN U7 SNRNP WITH METHYLATED noncleavable H2A* SUBSTRATE PRE-MRNA (COMPOSITE MAP)
Method: single particle / : Desotell A, Tong L

EMDB-72178: 
Cereblon Ternary Complex with Blimp1 and compound 5
Method: single particle / : Watson ER, Lander GC

EMDB-53417: 
Human UPF1 in complex with the histone stem loop RNA
Method: single particle / : Machado de Amorim A, Loll B, Hilal T, Chakrabarti S

EMDB-65926: 
Cryo-EM structure of the Type II secretion system protein from Acidithiobacillus caldus
Method: single particle / : Liu RH, Zhang K, Feng QS, Dai X, Zhang X, Li W, Bao ZP, Lin WF, Fu Y, Li Y

EMDB-62181: 
Cryo-EM structure of the Type II secretion system protein from Vibrio cholerae
Method: single particle / : Liu RH, Feng QS, Zhang K, Dai X, Dai J, Guo XR, Lin WF, Wang ZF, Fu Y, Li Y

EMDB-19627: 
Cryo-EM structure of CAK modified by covalent inhibitor SY-1365
Method: single particle / : Feng J, Koh AF, Kotecha A, Greber BJ

EMDB-19628: 
Cryo-EM structure of CAK in complex with SY-5609
Method: single particle / : Feng J, Cronin NB, Marineau JJ, Greber BJ

EMDB-17766: 
CryoEM structure of Nal1 protein, allele SPIKE, from Oryza sativa japonica group
Method: single particle / : Huang LY, Rety S, Xi XG

EMDB-17768: 
CryoEM structure of Nal1 protein, allele IR64, from Oryza sativa indica cultivar
Method: single particle / : Huang LY, Rety S, Xi XG

EMDB-29458: 
Alzheimer's disease paired-helical filament in complex with PET tracer GTP-1
Method: helical / : Merz GE, Tse E, Southworth DR

EMDB-17118: 
20S proteasome & CBR3 complex
Method: single particle / : Deshmukh FK, Sharon M

EMDB-34678: 
Cyanophage Pam3 neck
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-33799: 
Cyanophage Pam3 fiber
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-33802: 
Cyanophage Pam3 baseplate proteins
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-34679: 
Cyanophage Pam3 portal-adaptor
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-34680: 
Cyanophage Pam3 capsid asymmetric unit
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-34681: 
Cyanophage Pam3 Sheath-tube
Method: helical / : Yang F, Jiang YL, Zhou CZ

EMDB-31078: 
Cyanophage Pam1 capsid asymmetric unit
Method: single particle / : Zhang JT, Jiang YL

EMDB-31079: 
Cyanophage Pam1 portal-adaptor complex
Method: single particle / : Zhang JT, Jiang YL

EMDB-31080: 
Cyanophage Pam1 tail machine
Method: single particle / : Zhang JT, Jiang YL

EMDB-22627: 
Negative stain EM map of SARS-COV-2 spike protein (trimer) with Fab COV2-2082
Method: single particle / : Binshtein E, Crowe JE

EMDB-22628: 
Negative stain EM map of SARS-COV-2 spike protein (trimer) with Fab COV2-2479
Method: single particle / : Binshtein E, Crowe JE

EMDB-0960: 
Cryo-EM structure of RbcL8-RbcS4 from Anabaena sp. PCC 7120
Method: single particle / : Xia LY, Jiang YL

EMDB-22148: 
Negative stain EM map of SARS-COV-2 spike protein open RBD (trimer) with Fab COV2-2096
Method: single particle / : Binshtein E, Crowe JE

EMDB-22149: 
Negative stain EM map of SARS-COV-2 spike protein (trimer) with Fab COV2-2832
Method: single particle / : Binshtein E, Crowe JE

EMDB-0959: 
Cryo-EM structure of RuBisCO-Raf1 from Anabaena sp. PCC 7120
Method: single particle / : Xia LY, Jiang YL

EMDB-0961: 
The RuBisCO assembly intermediate RbcL-RbcS-Raf1 (L8S4F8)
Method: single particle / : Xia LY, Jiang YL, Kong WW, Sun H, Li WF, Chen Y, Zhou CZ

EMDB-0962: 
The cryo-EM structure of RuBisCO from Anabaena sp. PCC 7120
Method: single particle / : Xia LY, Jiang YL, Kong WW, Sun H, Li WF, Chen Y, Zhou CZ

EMDB-20159: 
Rotavirus A-VP3-8mer ssRNA complex (RVA-VP3-RNA)
Method: single particle / : Kumar D, Yu X

EMDB-21045: 
Cryo-EM map of human symplekin CTD
Method: single particle / : Sun Y, Zhang Y, Walz T, Tong L

EMDB-21046: 
Cryo-EM map of the core of human histone pre-mRNA cleavage complex (CPSF73-CPSF100-Symplekin)
Method: single particle / : Sun Y, Zhang Y, Walz T, Tong L

EMDB-21047: 
Cryo-EM map of the overall structure of human histone pre-mRNA 3'-end processing machinery
Method: single particle / : Sun Y, Zhang Y, Walz T, Tong L

EMDB-21050: 
Cryo-EM structure of an active human histone pre-mRNA 3'-end processing machinery at 3.2 Angstrom resolution
Method: single particle / : Sun Y, Zhang Y, Walz T, Tong L
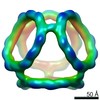
EMDB-21026: 
Truncated Tetrahedral RNA Nanostructures
Method: single particle / : Wu W, Zakrevsky P, Heinz W, Khant H, Kasprzak WK, Bindewald E, Dorjsuren N, Fields E, De Val N, Jaeger L, Shapiro BA

EMDB-9774: 
Capsid structure of a freshwater cyanophage Siphoviridae Mic1
Method: single particle / : Jin H, Jiang YL

EMDB-20367: 
Cryo-EM structure of the zebrafish TRPM2 channel in the apo conformation, processed with C4 symmetry
Method: single particle / : Yin Y, Wu M

EMDB-20368: 
Cryo-EM structure of the zebrafish TRPM2 channel in the apo conformation, processed with C2 symmetry (pseudo C4 symmetry)
Method: single particle / : Yin Y, Wu M

EMDB-20369: 
Cryo-EM structure of the zebrafish TRPM2 channel in the presence of ADPR and Ca2+
Method: single particle / : Yin Y, Wu M

EMDB-20192: 
Structure of the TRPV3 K169A sensitized mutant in apo form at 4.1 A resolution.
Method: single particle / : Zubcevic L, Borschel WF

EMDB-20194: 
Structure of the TRPV3 K169A sensitized mutant in the presence of 2-APB at 3.6 A resolution
Method: single particle / : Zubcevic L, Borschel WF

EMDB-7822: 
Cryo-EM structure of the zebrafish TRPM2 channel in the presence of Ca2+
Method: single particle / : Yin Y, Wu M

EMDB-9167: 
Electron cryo-tomography and subtomogram averaging of microtubule triplet from procentriole
Method: subtomogram averaging / : Li S, Fernandez JJ, Marshall W, Agard DA

EMDB-9168: 
Electron cryo-tomography and subtomogram averaging of microtubule triplet from procentriole
Method: subtomogram averaging / : Li S, Fernandez JJ, Marshall W, Agard DA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
