-Search query
-Search result
Showing 1 - 50 of 58 items for (author: georgiou & g)

EMDB-47490: 
Cryo-EM structure of the human P2X7 receptor in the apo closed state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

EMDB-47491: 
Cryo-EM structure of the human P2X7 receptor in the ATP-bound open state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

EMDB-47492: 
Cryo-EM structure of the human P2X7 receptor in the UB-ALT-P30-bound inhibited state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

EMDB-47493: 
Cryo-EM structure of the human P2X7 receptor in the UB-MBX-46-bound inhibited state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

EMDB-47494: 
Cryo-EM structure of the mouse P2X7 receptor in the apo closed state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

PDB-9e3m: 
Cryo-EM structure of the human P2X7 receptor in the apo closed state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

PDB-9e3n: 
Cryo-EM structure of the human P2X7 receptor in the ATP-bound open state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

PDB-9e3o: 
Cryo-EM structure of the human P2X7 receptor in the UB-ALT-P30-bound inhibited state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

PDB-9e3p: 
Cryo-EM structure of the human P2X7 receptor in the UB-MBX-46-bound inhibited state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

PDB-9e3q: 
Cryo-EM structure of the mouse P2X7 receptor in the apo closed state
Method: single particle / : Oken AC, Turcu AL, Vazquez S, Mansoor SE

EMDB-42839: 
Structure of UT14 Fab in complex with the head domain of H3 (A/Singapore/INFIMH-16-0019/2016)
Method: single particle / : Park J, Georgiou G

PDB-8uzc: 
Structure of UT14 Fab in complex with the head domain of H3 (A/Singapore/INFIMH-16-0019/2016)
Method: single particle / : Park J, Georgiou G

EMDB-43250: 
SARS-CoV-2 spike omicron (BA.1) ectodomain trimer in complex with SC27 Fab, global refinement
Method: single particle / : Byrne PO, McLellan JS

EMDB-43261: 
SARS-CoV-2 spike omicron (BA.1) ectodomain dimer-of-trimers in complex with SC27 Fab, global refinement
Method: single particle / : Byrne PO, McLellan JS

EMDB-43260: 
SARS-CoV-2 spike omicron (BA.1) ectodomain trimer in complex with SC27 Fab, local refinement
Method: single particle / : Byrne PO, McLellan JS

EMDB-43315: 
SARS-CoV-2 spike omicron (BA.1) RBD ectodomain dimer-of-trimers in complex with SC27 Fabs
Method: single particle / : Byrne PO, McLellan JS

PDB-8vif: 
SARS-CoV-2 spike omicron (BA.1) ectodomain trimer in complex with SC27 Fab, local refinement
Method: single particle / : Byrne PO, McLellan JS

PDB-8vke: 
SARS-CoV-2 spike omicron (BA.1) RBD ectodomain dimer-of-trimers in complex with SC27 Fabs
Method: single particle / : Byrne PO, McLellan JS

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS

EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS

EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tm1: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tma: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS

EMDB-12798: 
Hexameric coxsackievirus B3 2C protein in complex with S-fluoxetine
Method: single particle / : Hurdiss DL, Forster F

EMDB-23717: 
SARS-CoV-2 S-NTD + Fab CM25
Method: single particle / : Johnson NV, Mclellan JS

PDB-7m8j: 
SARS-CoV-2 S-NTD + Fab CM25
Method: single particle / : Johnson NV, Mclellan JS

EMDB-22897: 
Disulfide Stabilized Norovirus GI.1 VLP Shell Region
Method: single particle / : Gorman J, Kwong PD

PDB-7kjp: 
Disulfide Stabilized Norovirus GI.1 VLP Shell Region
Method: single particle / : Gorman J, Kwong PD
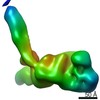
EMDB-10870: 
Cohesin complex with loader gripping DNA
Method: single particle / : Higashi TL, Eickhoff P, Sousa JS, Costa A, Uhlmann F

EMDB-10930: 
Cohesin complex with loader gripping DNA
Method: single particle / : Higashi TL, Eickhoff P

PDB-6yuf: 
Cohesin complex with loader gripping DNA
Method: single particle / : Higashi TL, Eickhoff P, Sousa JS, Costa A, Uhlmann F
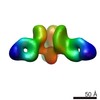
EMDB-7908: 
VIC170 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7909: 
VIC169 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
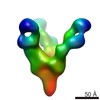
EMDB-7910: 
VIC167 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
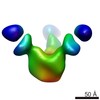
EMDB-7911: 
VIC166 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
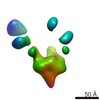
EMDB-7912: 
VIC165 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7914: 
VIC1164 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
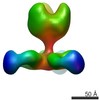
EMDB-7915: 
VIC163 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7926: 
VIC82 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
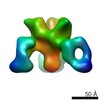
EMDB-7927: 
VIC43 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7928: 
VIC32 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7929: 
VIC26 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7930: 
VIC16 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7931: 
VIC15 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7932: 
VIC12 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
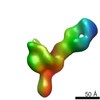
EMDB-7933: 
VIC11 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7922: 
VIC133 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7923: 
VIC101 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7924: 
VIC94 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7925: 
VIC91 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
