-Search query
-Search result
Showing all 36 items for (author: bullitt & e)

EMDB-28150: 
Cryo-EM reconstruction of the CFA/I bacterial adhesion pili
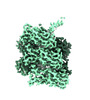
EMDB-28151: 
Cryo-EM reconstruction of the CS17 bacterial adhesion pili

EMDB-28152: 
Cryo-EM reconstruction of the CS20 bacterial adhesion pili

PDB-8ehr: 
Cryo-EM reconstruction of the CFA/I bacterial adhesion pili

PDB-8ehs: 
Cryo-EM reconstruction of the CS17 bacterial adhesion pili

PDB-8eht: 
Cryo-EM reconstruction of the CS20 bacterial adhesion pili

EMDB-28080: 
Human cardiac myosin II and associated essential light chain in the rigor conformation

EMDB-28081: 
Human beta-cardiac myosin II bound to ADP-MG2+ and the associated essential light chain

EMDB-28082: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in complex with ADP-Mg2+

PDB-8efd: 
Human cardiac myosin II and associated essential light chain in the rigor conformation

PDB-8efe: 
Human beta-cardiac myosin II bound to ADP-MG2+ and the associated essential light chain

PDB-8efh: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in complex with ADP-Mg2+

EMDB-28083: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in the rigor form

EMDB-28270: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin loop 4 7G mutant complex

PDB-8efi: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in the rigor form

PDB-8enc: 
Helical reconstruction of the human cardiac actin-tropomyosin-myosin loop 4 7G mutant complex

EMDB-22067: 
Bovine Cardiac Myosin in Complex with Chicken Skeletal Actin and Human Cardiac Tropomyosin in the Rigor State

PDB-6x5z: 
Bovine Cardiac Myosin in Complex with Chicken Skeletal Actin and Human Cardiac Tropomyosin in the Rigor State

EMDB-0497: 
Cryo-EM reconstruction of CFA/I pili

PDB-6nrv: 
Cryo-EM reconstruction of CFA/I pili

EMDB-9357: 
Outer Membrane Cytochrome S Filament from Geobacter Sulfurreducens

PDB-6nef: 
Outer Membrane Cytochrome S Filament from Geobacter Sulfurreducens

EMDB-7871: 
Class 3, Infectious Microvesicles from poliovirus-infected HeLa cells

EMDB-7872: 
Class 2, Infectious Microvesicles from poliovirus-infected HeLa cells

EMDB-7873: 
Class 1, Infectious Microvesicles from poliovirus-infected HeLa cells

EMDB-7877: 
Noodle-like structures in infectious microvesicles from poliovirus-infected HeLa cells

EMDB-7878: 
Noodle-like structure in infectious microvesicles (Class 3) from poliovirus-infected HeLa cells

EMDB-7879: 
Infectious exosomes from poliovirus-infected HeLa cells

EMDB-7880: 
Infectious microvesicles (Class 1 and 3) from poliovirus-infected HeLa cells, displaying clusters of virions and additional inner vesicular structures

EMDB-7881: 
Subcellular fractionation of viral replication membranes from poliovirus-infected HeLa cells at 5 hours post infection

EMDB-8654: 
Zika virus-infected Vero E6 cell at 48 hpi: dual-axis tilt series tomogram from 3 serial sections

EMDB-8655: 
Zika virus-infected Vero E6 cell at 20 hpi: dual-axis tilt series tomogram from 3 serial sections

EMDB-8656: 
Zika virus-infected Vero E6 cell at 48 hpi: dual-axis tilt series tomogram from 2 serial sections

EMDB-8657: 
Zika virus-infected Vero E6 cell at 48 hpi, showing spherule pore opening to cytoplasm

EMDB-5560: 
CryoEM of Poliovirus Polymerase Tube, including Ghost Reflections

EMDB-2270: 
Average reconstruction of cryoEM poliovirus polymerase tubes with (-32,6) helical symmetry
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
