[English] 日本語
 Yorodumi
Yorodumi- EMDB-28083: Helical reconstruction of the human cardiac actin-tropomyosin-myo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Helical reconstruction of the human cardiac actin-tropomyosin-myosin complex in the rigor form | |||||||||
 Map data Map data | Primary map for the human cardiac actin-tropomyosin-beta-myosin II complex in the rigor conformation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | actin / tropomyosin / myosin / cardiac / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of slow-twitch skeletal muscle fiber contraction / regulation of the force of skeletal muscle contraction / positive regulation of heart rate by epinephrine / muscle thin filament tropomyosin / actin-myosin filament sliding / muscle myosin complex / regulation of muscle contraction / regulation of the force of heart contraction / bleb / transition between fast and slow fiber ...regulation of slow-twitch skeletal muscle fiber contraction / regulation of the force of skeletal muscle contraction / positive regulation of heart rate by epinephrine / muscle thin filament tropomyosin / actin-myosin filament sliding / muscle myosin complex / regulation of muscle contraction / regulation of the force of heart contraction / bleb / transition between fast and slow fiber / ruffle organization / myosin filament / Striated Muscle Contraction / cardiac muscle hypertrophy in response to stress / muscle filament sliding / myosin complex / adult heart development / myosin II complex / structural constituent of muscle / sarcomere organization / ventricular cardiac muscle tissue morphogenesis / microfilament motor activity / myosin binding / heart contraction / negative regulation of vascular associated smooth muscle cell migration / regulation of heart contraction / mesenchyme migration / myofibril / negative regulation of vascular associated smooth muscle cell proliferation / Smooth Muscle Contraction / ATP metabolic process / striated muscle contraction / skeletal muscle contraction / cytoskeletal protein binding / positive regulation of stress fiber assembly / cardiac muscle contraction / stress fiber / cytoskeleton organization / positive regulation of cell adhesion / regulation of heart rate / muscle contraction / negative regulation of cell migration / actin filament organization / sarcomere / filopodium / actin filament / cellular response to reactive oxygen species / wound healing / structural constituent of cytoskeleton / ruffle membrane / Z disc / actin filament binding / regulation of cell shape / lamellipodium / actin cytoskeleton / actin binding / cell body / cytoskeleton / calmodulin binding / protein heterodimerization activity / positive regulation of gene expression / protein homodimerization activity / ATP binding / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
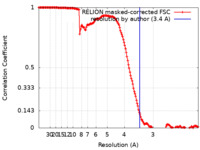
| Method | helical reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Doran MH / Lehman W / Rynkiewicz MJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Gen Physiol / Year: 2023 Journal: J Gen Physiol / Year: 2023Title: Myosin loop-4 is critical for optimal tropomyosin repositioning on actin during muscle activation and relaxation. Authors: Matthew H Doran / Michael J Rynkiewicz / Elumalai Pavadai / Skylar M L Bodt / David Rasicci / Jeffrey R Moore / Christopher M Yengo / Esther Bullitt / William Lehman /  Abstract: During force-generating steps of the muscle crossbridge cycle, the tip of the myosin motor, specifically loop-4, contacts the tropomyosin cable of actin filaments. In the current study, we determined ...During force-generating steps of the muscle crossbridge cycle, the tip of the myosin motor, specifically loop-4, contacts the tropomyosin cable of actin filaments. In the current study, we determined the corresponding effect of myosin loop-4 on the regulatory positioning of tropomyosin on actin. To accomplish this, we compared high-resolution cryo-EM structures of myosin S1-decorated thin filaments containing either wild-type or a loop-4 mutant construct, where the seven-residue portion of myosin loop-4 that contacts tropomyosin was replaced by glycine residues, thus removing polar side chains from residues 366-372. Cryo-EM analysis of fully decorated actin-tropomyosin filaments with wild-type and mutant S1, yielded 3.4-3.6 Å resolution reconstructions, with even higher definition at the actin-myosin interface. Loop-4 densities both in wild-type and mutant S1 were clearly identified, and side chains were resolved in the wild-type structure. Aside from loop-4, actin and myosin structural domains were indistinguishable from each other when filaments were decorated with either mutant or wild-type S1. In marked contrast, the position of tropomyosin on actin in the two reconstructions differed by 3 to 4 Å. In maps of filaments containing the mutant, tropomyosin was located closer to the myosin-head and thus moved in the direction of the C-state conformation adopted by myosin-free thin filaments. Complementary interaction energy measurements showed that tropomyosin in the mutant thin filaments sits on actin in a local energy minimum, whereas tropomyosin is positioned by wild-type S1 in an energetically unfavorable location. We propose that the high potential energy associated with tropomyosin positioning in wild-type filaments favors an effective transition to B- and C-states following release of myosin from the thin filaments during relaxation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28083.map.gz emd_28083.map.gz | 179.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28083-v30.xml emd-28083-v30.xml emd-28083.xml emd-28083.xml | 25.7 KB 25.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28083_fsc.xml emd_28083_fsc.xml | 15.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_28083.png emd_28083.png | 59.7 KB | ||
| Masks |  emd_28083_msk_1.map emd_28083_msk_1.map | 325 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28083.cif.gz emd-28083.cif.gz | 8.2 KB | ||
| Others |  emd_28083_additional_1.map.gz emd_28083_additional_1.map.gz emd_28083_half_map_1.map.gz emd_28083_half_map_1.map.gz emd_28083_half_map_2.map.gz emd_28083_half_map_2.map.gz | 294.5 MB 258.2 MB 258.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28083 http://ftp.pdbj.org/pub/emdb/structures/EMD-28083 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28083 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28083 | HTTPS FTP |
-Related structure data
| Related structure data |  8efiMC  8encC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28083.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28083.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Primary map for the human cardiac actin-tropomyosin-beta-myosin II complex in the rigor conformation | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.078 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28083_msk_1.map emd_28083_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Non-locally-filtered version of the primary map.
| File | emd_28083_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Non-locally-filtered version of the primary map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1.
| File | emd_28083_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2.
| File | emd_28083_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human cardiac actin-tropomyosin-beta-myosin II complex in the rig...
| Entire | Name: Human cardiac actin-tropomyosin-beta-myosin II complex in the rigor form. |
|---|---|
| Components |
|
-Supramolecule #1: Human cardiac actin-tropomyosin-beta-myosin II complex in the rig...
| Supramolecule | Name: Human cardiac actin-tropomyosin-beta-myosin II complex in the rigor form. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 / Details: Cardiac actomyosin-tropomyosin complex. |
|---|
-Supramolecule #2: Cardiac F-actin complex
| Supramolecule | Name: Cardiac F-actin complex / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2 / Details: F-actin forms the backbone of the complex |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Human cardiac myosin II
| Supramolecule | Name: Human cardiac myosin II / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 Details: The motor domain of the myosin saturates the actin filament. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: Human cardiac tropomyosin
| Supramolecule | Name: Human cardiac tropomyosin / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 / Details: Tropomyosin wraps around the F-actin core. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Myosin-7
| Macromolecule | Name: Myosin-7 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 223.445984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGDSEMAVFG AAAPYLRKSE KERLEAQTRP FDLKKDVFVP DDKQEFVKAK IVSREGGKVT AETEYGKTVT VKEDQVMQQN PPKFDKIED MAMLTFLHEP AVLYNLKDRY GSWMIYTYSG LFCVTVNPYK WLPVYTPEVV AAYRGKKRSE APPHIFSISD N AYQYMLTD ...String: MGDSEMAVFG AAAPYLRKSE KERLEAQTRP FDLKKDVFVP DDKQEFVKAK IVSREGGKVT AETEYGKTVT VKEDQVMQQN PPKFDKIED MAMLTFLHEP AVLYNLKDRY GSWMIYTYSG LFCVTVNPYK WLPVYTPEVV AAYRGKKRSE APPHIFSISD N AYQYMLTD RENQSILITG ESGAGKTVNT KRVIQYFAVI AAIGDRSKKD QSPGKGTLED QIIQANPALE AFGNAKTVRN DN SSRFGKF IRIHFGATGK LASADIETYL LEKSRVIFQL KAERDYHIFY QILSNKKPEL LDMLLITNNP YDYAFISQGE TTV ASIDDA EELMATDNAF DVLGFTSEEK NSMYKLTGAI MHFGNMKFKL KQREEQAEPD GTEEADKSAY LMGLNSADLL KGLC HPRVK VGNEYVTKGQ NVQQVIYATG ALAKAVYERM FNWMVTRINA TLETKQPRQY FIGVLDIAGF EIFDFNSFEQ LCINF TNEK LQQFFNHHMF VLEQEEYKKE GIEWTFIDFG MDLQACIDLI EKPMGIMSIL EEECMFPKAT DMTFKAKLFD NHLGKS ANF QKPRNIKGKP EAHFSLIHYA GIVDYNIIGW LQKNKDPLNE TVVGLYQKSS LKLLSTLFAN YAGADAPIEK GKGKAKK GS SFQTVSALHR ENLNKLMTNL RSTHPHFVRC IIPNETKSPG VMDNPLVMHQ LRCNGVLEGI RICRKGFPNR ILYGDFRQ R YRILNPAAIP EGQFIDSRKG AEKLLSSLDI DHNQYKFGHT KVFFKAGLLG LLEEMRDERL SRIITRIQAQ SRGVLARME YKKLLERRDS LLVIQWNIRA FMGVKNWPWM KLYFKIKPLL KSAEREKEMA SMKEEFTRLK EALEKSEARR KELEEKMVSL LQEKNDLQL QVQAEQDNLA DAEERCDQLI KNKIQLEAKV KEMNERLEDE EEMNAELTAK KRKLEDECSE LKRDIDDLEL T LAKVEKEK HATENKVKNL TEEMAGLDEI IAKLTKEKKA LQEAHQQALD DLQAEEDKVN TLTKAKVKLE QQVDDLEGSL EQ EKKVRMD LERAKRKLEG DLKLTQESIM DLENDKQQLD ERLKKKDFEL NALNARIEDE QALGSQLQKK LKELQARIEE LEE ELEAER TARAKVEKLR SDLSRELEEI SERLEEAGGA TSVQIEMNKK REAEFQKMRR DLEEATLQHE ATAAALRKKH ADSV AELGE QIDNLQRVKQ KLEKEKSEFK LELDDVTSNM EQIIKAKANL EKMCRTLEDQ MNEHRSKAEE TQRSVNDLTS QRAKL QTEN GELSRQLDEK EALISQLTRG KLTYTQQLED LKRQLEEEVK AKNALAHALQ SARHDCDLLR EQYEEETEAK AELQRV LSK ANSEVAQWRT KYETDAIQRT EELEEAKKKL AQRLQEAEEA VEAVNAKCSS LEKTKHRLQN EIEDLMVDVE RSNAAAA AL DKKQRNFDKI LAEWKQKYEE SQSELESSQK EARSLSTELF KLKNAYEESL EHLETFKREN KNLQEEISDL TEQLGSSG K TIHELEKVRK QLEAEKMELQ SALEEAEASL EHEEGKILRA QLEFNQIKAE IERKLAEKDE EMEQAKRNHL RVVDSLQTS LDAETRSRNE ALRVKKKMEG DLNEMEIQLS HANRMAAEAQ KQVKSLQSLL KDTQIQLDDA VRANDDLKEN IAIVERRNNL LQAELEELR AVVEQTERSR KLAEQELIET SERVQLLHSQ NTSLINQKKK MDADLSQLQT EVEEAVQECR NAEEKAKKAI T DAAMMAEE LKKEQDTSAH LERMKKNMEQ TIKDLQHRLD EAEQIALKGG KKQLQKLEAR VRELENELEA EQKRNAESVK GM RKSERRI KELTYQTEED RKNLLRLQDL VDKLQLKVKA YKRQAEEAEE QANTNLSKFR KVQHELDEAE ERADIAESQV NKL RAKSRD IGTKGLNEE UniProtKB: Myosin-7 |
-Macromolecule #2: Actin, alpha cardiac muscle 1
| Macromolecule | Name: Actin, alpha cardiac muscle 1 / type: protein_or_peptide / ID: 2 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.064891 KDa |
| Sequence | String: MCDDEETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIEHGIITN WDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLDSGD G VTHNVPIY ...String: MCDDEETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIEHGIITN WDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLDSGD G VTHNVPIY EGYALPHAIM RLDLAGRDLT DYLMKILTER GYSFVTTAER EIVRDIKEKL CYVALDFENE MATAASSSSL EK SYELPDG QVITIGNERF RCPETLFQPS FIGMESAGIH ETTYNSIMKC DIDIRKDLYA NNVLSGGTTM YPGIADRMQK EIT ALAPST MKIKIIAPPE RKYSVWIGGS ILASLSTFQQ MWISKQEYDE AGPSIVHRKC F UniProtKB: Actin alpha cardiac muscle 1 |
-Macromolecule #3: Tropomyosin alpha-1 chain
| Macromolecule | Name: Tropomyosin alpha-1 chain / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.763621 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDAIKKKMQM LKLDKENALD RAEQAEADKK AAEDRSKQLE DELVSLQKKL KGTEDELDKY SEALKDAQEK LELAEKKATD AEADVASLN RRIQLVEEEL DRAQERLATA LQKLEEAEKA ADESERGMKV IESRAQKDEE KMEIQEIQLK EAKHIAEDAD R KYEEVARK ...String: MDAIKKKMQM LKLDKENALD RAEQAEADKK AAEDRSKQLE DELVSLQKKL KGTEDELDKY SEALKDAQEK LELAEKKATD AEADVASLN RRIQLVEEEL DRAQERLATA LQKLEEAEKA ADESERGMKV IESRAQKDEE KMEIQEIQLK EAKHIAEDAD R KYEEVARK LVIIESDLER AEERAELSEG KCAELEEELK TVTNNLKSLE AQAEKYSQKE DRYEEEIKVL SDKLKEAETR AE FAERSVT KLEKSIDDLE DELYAQKLKY KAISEELDHA LNDMTSI UniProtKB: Tropomyosin alpha-1 chain |
-Macromolecule #4: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 5 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.13 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Support film - Material: GOLD / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 539 pixel / Digitization - Dimensions - Height: 539 pixel / Number grids imaged: 4 / Number real images: 3961 / Average exposure time: 3.12 sec. / Average electron dose: 53.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 80000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)