-Search query
-Search result
Showing all 46 items for (author: antony & e)

EMDB-18539: 
Structure of the human 80S ribosome at 1.9 A resolution - the molecular role of chemical modifications and ions in RNA
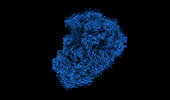
EMDB-18812: 
The structure of the human 80S ribosome at 1.9 angstrom resolution reveals the molecular role of chemical modifications and ions in RNA - Focused refinement of the of the 60S subunit
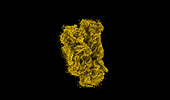
EMDB-18813: 
The structure of the human 80S ribosome at 1.9 angstrom resolution reveals the molecular role of chemical modifications and ions in RNA - Focused refinement of the of the 40S subunit body
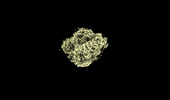
EMDB-18814: 
The structure of the human 80S ribosome at 1.9 angstrom resolution reveals the molecular role of chemical modifications and ions in RNA - Focused refinement of the of the 40S subunit head
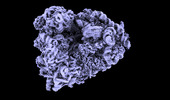
EMDB-18815: 
Structure of the human 80S ribosome at 1.9 A resolution - the molecular role of chemical modifications and ions in RNA - Global human 80S ribosome refinement before focused refinements.

PDB-8qoi: 
Structure of the human 80S ribosome at 1.9 A resolution - the molecular role of chemical modifications and ions in RNA

EMDB-16916: 
Bipartite interaction of TOPBP1 with the GINS complex

PDB-8ok2: 
Bipartite interaction of TOPBP1 with the GINS complex

EMDB-29695: 
CryoEM structure of yeast recombination mediator Rad52

PDB-8g3g: 
CryoEM structure of yeast recombination mediator Rad52

EMDB-16041: 
CryoEM structure of quinol-dependent Nitric Oxide Reductase (qNOR) from Alcaligenes xylosoxidans at 2.2 A resolution

PDB-8bgw: 
CryoEM structure of quinol-dependent Nitric Oxide Reductase (qNOR) from Alcaligenes xylosoxidans at 2.2 A resolution

EMDB-15414: 
Vaccinia C16 N-terminal domains

EMDB-15415: 
Vaccinia C16 protein bound to Ku70/Ku80

EMDB-15416: 
Vaccinia C16 protein bound to Ku70/Ku80

PDB-8ag3: 
Vaccinia C16 N-terminal domains

PDB-8ag4: 
Vaccinia C16 protein bound to Ku70/Ku80

PDB-8ag5: 
Vaccinia C16 protein bound to Ku70/Ku80

EMDB-13895: 
CryoEM structure of the Smc5/6-holocomplex (composite structure)

PDB-7qcd: 
CryoEM structure of the Smc5/6-holocomplex (composite structure)

EMDB-13893: 
Cryo-EM structure of the Smc5/6 holo-complex; map for head-end of complex.

EMDB-13894: 
CryoEM structure of the Smc5/6 holocomplex; map for hinge and arm region.

EMDB-14527: 
Structure of DNA-bound human RAD17-RFC clamp loader and 9-1-1 checkpoint clamp

PDB-7z6h: 
Structure of DNA-bound human RAD17-RFC clamp loader and 9-1-1 checkpoint clamp

EMDB-22884: 
Structure of the NiV F glycoprotein in complex with the 12B2 neutralizing antibody

EMDB-22885: 
Structure of the HeV F glycoprotein in complex with the 1F5 neutralizing antibody

PDB-7ki4: 
Structure of the NiV F glycoprotein in complex with the 12B2 neutralizing antibody

PDB-7ki6: 
Structure of the HeV F glycoprotein in complex with the 1F5 neutralizing antibody

EMDB-11002: 
CryoEM structure of bovine cytochrome bc1 in complex with a tetrahydroquinolone inhibitor

EMDB-4642: 
12 Angstrom structure of detergent solubilised LAT1-CD98hc

EMDB-0822: 
Cryo-EM structure of dimeric quinol dependent Nitric Oxide Reductase (qNOR) from the pathogen Neisseria meninigitidis

EMDB-10387: 
Glu-494-Ala inactive monomer of a quinol dependent Nitric Oxide Reductase (qNOR) from Alcaligenes xylosoxidans

PDB-6l3h: 
Cryo-EM structure of dimeric quinol dependent Nitric Oxide Reductase (qNOR) from the pathogen Neisseria meninigitidis

PDB-6t6v: 
Glu-494-Ala inactive monomer of a quinol dependent Nitric Oxide Reductase (qNOR) from Alcaligenes xylosoxidans

EMDB-4618: 
Cryo-EM structure of dimeric quinol dependent nitric oxide reductase (qNOR) from Alcaligenes xylosoxidans

EMDB-4619: 
Cryo-EM structure of dimeric quinol dependent nitric oxide reductase (qNOR) Val495Ala mutant from Alcaligenes xylosoxidans

PDB-6qq5: 
Cryo-EM structure of dimeric quinol dependent nitric oxide reductase (qNOR) from Alcaligenes xylosoxidans

PDB-6qq6: 
Cryo-EM structure of dimeric quinol dependent nitric oxide reductase (qNOR) Val495Ala mutant from Alcaligenes xylosoxidans

EMDB-4410: 
Yeast RPA bound to ssDNA

PDB-6i52: 
Yeast RPA bound to ssDNA

EMDB-4286: 
CryoEM structure of bovine cytochrome bc1 in complex with the anti-malarial compound GSK932121

EMDB-4288: 
CryoEM structure of bovine cytochrome bc1 with no ligand bound

EMDB-4292: 
CryoEM structure of bovine cytochrome bc1 in complex with the anti-malarial inhibitor SCR0911

PDB-6fo0: 
CryoEM structure of bovine cytochrome bc1 in complex with the anti-malarial compound GSK932121

PDB-6fo2: 
CryoEM structure of bovine cytochrome bc1 with no ligand bound

PDB-6fo6: 
CryoEM structure of bovine cytochrome bc1 in complex with the anti-malarial inhibitor SCR0911
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
