[English] 日本語
 Yorodumi
Yorodumi- EMDB-4618: Cryo-EM structure of dimeric quinol dependent nitric oxide reduct... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4618 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of dimeric quinol dependent nitric oxide reductase (qNOR) from Alcaligenes xylosoxidans | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Proton Transfer / Membrane Protein / Homodimer / OXIDOREDUCTASE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnitric oxide reductase (cytochrome c) / nitric oxide reductase activity / cytochrome-c oxidase activity / aerobic respiration / heme binding / membrane Similarity search - Function | |||||||||||||||
| Biological species |  Achromobacter xylosoxidans (bacteria) / Achromobacter xylosoxidans (bacteria) /  Alcaligenes xylosoxydans xylosoxydans (bacteria) Alcaligenes xylosoxydans xylosoxydans (bacteria) | |||||||||||||||
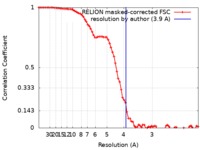
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||||||||
 Authors Authors | Gopalasingam CC / Johnson RM / Chiduza GN | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2019 Journal: Sci Adv / Year: 2019Title: Dimeric structures of quinol-dependent nitric oxide reductases (qNORs) revealed by cryo-electron microscopy. Authors: Chai C Gopalasingam / Rachel M Johnson / George N Chiduza / Takehiko Tosha / Masaki Yamamoto / Yoshitsugu Shiro / Svetlana V Antonyuk / Stephen P Muench / S Samar Hasnain /   Abstract: Quinol-dependent nitric oxide reductases (qNORs) are membrane-integrated, iron-containing enzymes of the denitrification pathway, which catalyze the reduction of nitric oxide (NO) to the major ozone ...Quinol-dependent nitric oxide reductases (qNORs) are membrane-integrated, iron-containing enzymes of the denitrification pathway, which catalyze the reduction of nitric oxide (NO) to the major ozone destroying gas nitrous oxide (NO). Cryo-electron microscopy structures of active qNOR from and an activity-enhancing mutant have been determined to be at local resolutions of 3.7 and 3.2 Å, respectively. They unexpectedly reveal a dimeric conformation (also confirmed for qNOR from ) and define the active-site configuration, with a clear water channel from the cytoplasm. Structure-based mutagenesis has identified key residues involved in proton transport and substrate delivery to the active site of qNORs. The proton supply direction differs from cytochrome c-dependent NOR (cNOR), where water molecules from the cytoplasm serve as a proton source similar to those from cytochrome c oxidase. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4618.map.gz emd_4618.map.gz | 25.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4618-v30.xml emd-4618-v30.xml emd-4618.xml emd-4618.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4618_fsc.xml emd_4618_fsc.xml | 7.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_4618.png emd_4618.png | 33.2 KB | ||
| Filedesc metadata |  emd-4618.cif.gz emd-4618.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4618 http://ftp.pdbj.org/pub/emdb/structures/EMD-4618 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4618 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4618 | HTTPS FTP |
-Related structure data
| Related structure data |  6qq5MC  4619C  6qq6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4618.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4618.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Quinol Dependent Nitric Oxide Reductase
| Entire | Name: Quinol Dependent Nitric Oxide Reductase |
|---|---|
| Components |
|
-Supramolecule #1: Quinol Dependent Nitric Oxide Reductase
| Supramolecule | Name: Quinol Dependent Nitric Oxide Reductase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Achromobacter xylosoxidans (bacteria) Achromobacter xylosoxidans (bacteria) |
| Molecular weight | Theoretical: 170 KDa |
-Macromolecule #1: Nitric oxide reductase subunit B
| Macromolecule | Name: Nitric oxide reductase subunit B / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: nitric oxide reductase (cytochrome c) |
|---|---|
| Source (natural) | Organism:  Alcaligenes xylosoxydans xylosoxydans (bacteria) Alcaligenes xylosoxydans xylosoxydans (bacteria) |
| Molecular weight | Theoretical: 83.121984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGPYRRLWFT LIAVLAVTFA LLGFYGGEVY RQAPPIPEEV ASADGTRLFG RDDILDGQTA WQSIGGMQLG SIWGHGAYQA PDWTADWLH RELMAWLDLA ARDAHGRDYG QLDAPAQAAL REQLKAEYRA NRADAAGGKL TLSPRRAQAV AQTEAYYDQL F SDAPALHR ...String: MGPYRRLWFT LIAVLAVTFA LLGFYGGEVY RQAPPIPEEV ASADGTRLFG RDDILDGQTA WQSIGGMQLG SIWGHGAYQA PDWTADWLH RELMAWLDLA ARDAHGRDYG QLDAPAQAAL REQLKAEYRA NRADAAGGKL TLSPRRAQAV AQTEAYYDQL F SDAPALHR SRENYAMKEN TLPDANRRRQ MTHFFFWTAW AAGTEREGTS VTYTNNWPHE PLIGNHPSSE NVMWSIISVV VL LAGIGLL IWAWAFLRGK EEDEPPAPAR DPLTTFALTP SQRALGKYLF LVVALFGFQV LLGGFTAHYT VEGQKFYGID LSQ WFPYSL VRTWHIQSAL FWIATGFLAA GLFLAPLING GRDPKYQKAG VDILFWALVL VVVGSFAGNY LAIAQIMPPD LNFW LGHQG YEYVDLGRLW QIGKFAGICF WLVLMLRGIV PALRTPGGDK NLLALLTASV GAIGLFYGAG FFYGERTHLT VMEYW RWWI VHLWVEGFFE VFATTALAFI FSTLGLVSRR MATTASLASA SLFMLGGIPG TFHHLYFAGT TTPVMAVGAS FSALEV VPL IVLGHEAWEN WRLKTRAPWM ENLKWPLMCF VAVAFWNMLG AGVFGFMINP PVSLYYIQGL NTTPVHAHAA LFGVYGF LA LGFTLLVLRY IRPQYALSPG LMKLAFWGLN LGLALMIFTS LLPIGLIQFH ASVSEGMWYA RSEAFMQQDI LKTLRWGR T FGDVVFLLGA LAMVVQVILG LLSGK UniProtKB: Nitric oxide reductase subunit B |
-Macromolecule #2: PROTOPORPHYRIN IX CONTAINING FE
| Macromolecule | Name: PROTOPORPHYRIN IX CONTAINING FE / type: ligand / ID: 2 / Number of copies: 4 / Formula: HEM |
|---|---|
| Molecular weight | Theoretical: 616.487 Da |
| Chemical component information |  ChemComp-HEM: |
-Macromolecule #3: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Blot for 6 seconds with blot force of 6 prior to plunge freezing. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 3213 / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -3.5 µm / Nominal defocus min: -1.5 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)