9YQ4
 
 | |
9YGU
 
 | | Flagella filament structure in H. pylori composed of flagellin FlaA | | Descriptor: | 5,7-diamino-3,5,7,9-tetradeoxy-L-glycero-alpha-L-manno-non-2-ulopyranosonic acid, Flagellin | | Authors: | Kumar, R, Yu, H, Tachiyama, S, Liu, J. | | Deposit date: | 2025-09-29 | | Release date: | 2026-02-25 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | Assembly and glycosylation of Helicobacter pylori sheathed flagella.
Pnas Nexus, 5, 2026
|
|
9YQ2
 
 | |
9YQ5
 
 | |
9ZRG
 
 | | Structure of naked mole-rat ribosome (non-rotated) | | Descriptor: | 18S ribosomal RNA, 28S ribosomal RNA, 40S ribosomal protein S12, ... | | Authors: | Gutierrez-Vargas, C, De, S, Maji, S, Liu, Z, Nieb, M, Seluanov, A, Gorbunova, V, Frank, J. | | Deposit date: | 2025-12-19 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structures of naked mole-rat, tuco-tuco, and guinea pig ribosomes-is rRNA fragmentation linked to translational fidelity?
Nucleic Acids Res., 54, 2026
|
|
9Z6U
 
 | |
9YNH
 
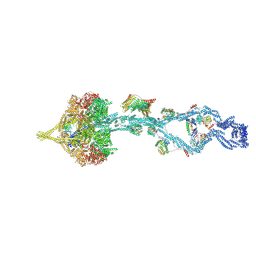 | | Full-length human cytoplasmic dynein-1 in phi-like state bound to dynactin-p150glued and LIS1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Cytoplasmic dynein 1 heavy chain 1, ... | | Authors: | Yang, J, Rao, Q, Chai, P, Zhang, K. | | Deposit date: | 2025-10-10 | | Release date: | 2026-01-28 | | Last modified: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (5.5 Å) | | Cite: | Roles of microtubules and LIS1 in dynein transport machinery assembly.
Nature, 2026
|
|
9YNC
 
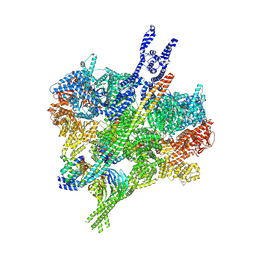 | | Motor domains of phi-like human dynein-1 bound to dynactin-p150glued and LIS1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Cytoplasmic dynein 1 heavy chain 1, ... | | Authors: | Yang, J, Rao, Q, Chai, P, Zhang, K. | | Deposit date: | 2025-10-10 | | Release date: | 2026-01-28 | | Last modified: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (3.93 Å) | | Cite: | Roles of microtubules and LIS1 in dynein transport machinery assembly.
Nature, 2026
|
|
9YRY
 
 | | Crystal structure of HIV-1 Protease WT (NL4-3) in Complex with NR02-73 | | Descriptor: | (3R,3aS,6aR)-hexahydrofuro[2,3-b]furan-3-yl [(2S,3R)-3-hydroxy-4-{[(1S)-1-hydroxy-2,3-dihydro-1H-indene-5-sulfonyl][(2S)-2-methylbutyl]amino}-1-phenylbutan-2-yl]carbamate, Protease, SULFATE ION | | Authors: | Lockbaum, G.J, Kaur, J, Shaqra, A.M, Schiffer, C.A. | | Deposit date: | 2025-10-17 | | Release date: | 2025-12-24 | | Last modified: | 2026-01-07 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Fluorinated HIV-1 protease inhibitors containing chiral hydroxyethylbenzene and indanol as P2' ligands with potent activity against drug-resistant variants.
Eur.J.Med.Chem., 304, 2025
|
|
9YNG
 
 | | Dynactin and dynein-1 tail region of dynein-dynactin complex on microtubule in the presence of LIS1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Actin, ... | | Authors: | Yang, J, Rao, Q, Chai, P, Zhang, K. | | Deposit date: | 2025-10-10 | | Release date: | 2026-01-28 | | Last modified: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (4.07 Å) | | Cite: | Roles of microtubules and LIS1 in dynein transport machinery assembly.
Nature, 2026
|
|
9Z2D
 
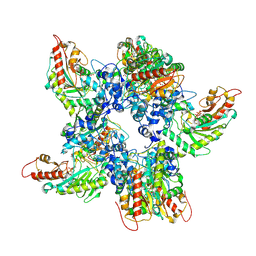 | | Mitochondrial Creatine Kinase in complex with ADP, creatine, and uncompetitive inhibitor uci | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Creatine kinase U-type, mitochondrial, ... | | Authors: | Demir, M, Zhao, J, Sergienko, E. | | Deposit date: | 2025-11-04 | | Release date: | 2026-01-28 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Uncompetitive Allosteric Inhibitor of Mitochondrial Creatine Kinase Prevents Binding and Release of Creatine by Stabilization of Loop Closure.
Biorxiv, 2026
|
|
9Z2F
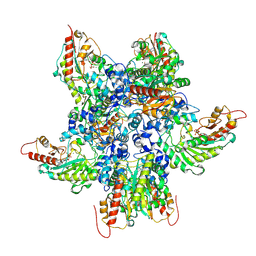 
 | |
9YZV
 
 | | Joint Xray/Neutron structure of human DJ-1 at room temperature | | Descriptor: | Protein deglycase DJ-1, deuterium(1+) | | Authors: | Lin, J, Kovalevsky, A, Walker, A.R, Wilson, M.A. | | Deposit date: | 2025-10-30 | | Release date: | 2025-11-19 | | Last modified: | 2026-04-01 | | Method: | NEUTRON DIFFRACTION (1.63 Å), X-RAY DIFFRACTION | | Cite: | Environmental Contributions to Proton Sharing in Protein Low-Barrier Hydrogen Bonds.
Biochemistry, 65, 2026
|
|
9ZRU
 
 | |
9YND
 
 | | Motor domain of human dynein-1 in pre-power stroke bound to dynactin-p150glued-CC1B and LIS1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Cytoplasmic dynein 1 heavy chain 1, ... | | Authors: | Yang, J, Rao, Q, Chai, P, Zhang, K. | | Deposit date: | 2025-10-10 | | Release date: | 2026-01-28 | | Last modified: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (4.26 Å) | | Cite: | Roles of microtubules and LIS1 in dynein transport machinery assembly.
Nature, 2026
|
|
9YKP
 
 | | HIV-1 Protease WT (NL4-3) with Inhibitor NR01-141 | | Descriptor: | (3R,3aS,6aR)-hexahydrofuro[2,3-b]furan-3-yl [(2S,3R)-3-hydroxy-4-{[(1S)-1-hydroxy-2,3-dihydro-1H-indene-5-sulfonyl](2-methylpropyl)amino}-1-phenylbutan-2-yl]carbamate, Protease, SULFATE ION | | Authors: | Lockbaum, G.J, Kaur, J, Shaqra, A.M, Schiffer, C.A. | | Deposit date: | 2025-10-07 | | Release date: | 2025-12-24 | | Last modified: | 2026-01-07 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Fluorinated HIV-1 protease inhibitors containing chiral hydroxyethylbenzene and indanol as P2' ligands with potent activity against drug-resistant variants.
Eur.J.Med.Chem., 304, 2025
|
|
9YZW
 
 | | Joint Xray/Neutron structure of Escherichia coli YajL at room temperature | | Descriptor: | Chaperone YajL, deuterium(1+) | | Authors: | Lin, J, Kovalevsky, A, Walker, A.R, Wilson, M.A. | | Deposit date: | 2025-10-30 | | Release date: | 2025-11-19 | | Last modified: | 2026-04-01 | | Method: | NEUTRON DIFFRACTION (1.65 Å), X-RAY DIFFRACTION | | Cite: | Environmental Contributions to Proton Sharing in Protein Low-Barrier Hydrogen Bonds.
Biochemistry, 65, 2026
|
|
9YR6
 
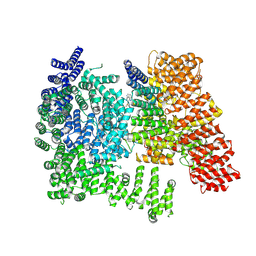 | | Structure of HTTQ23-HAP40 complex bound to a small molecule ligand | | Descriptor: | 2-[(1R)-2,2-difluorocyclopropyl]-N-[(1-methyl-1H-indazol-6-yl)methyl]aniline, 40-kDa huntingtin-associated protein, Huntingtin | | Authors: | Balakrishnan, S, Deme, J, Lea, S.M, Harding, R.J. | | Deposit date: | 2025-10-16 | | Release date: | 2025-12-03 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Identification and Characterization of a Small Molecule Ligand for the Huntingtin-HAP40 Complex
Biorxiv, 2025
|
|
9YV3
 
 | | Crystal Structure of a ternary complex of Saccharomyces cerevisiae Sec14 with a small molecule inhibitor and phosphatidylcholine | | Descriptor: | (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, (4-chloranyl-3-nitro-phenyl)-[4-(2-fluorophenyl)piperazin-1-yl]methanone, SEC14 cytosolic factor | | Authors: | Singh, P.K, Green, S.M, Krieger, I, Sacchettini, J, Igumenova, T.I, Bankaitis, V.A. | | Deposit date: | 2025-10-23 | | Release date: | 2025-11-05 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal Structure of a ternary complex of Saccharomyces cerevisiae Sec14 with a small molecule inhibitor and phosphatidylcholine
To Be Published
|
|
9ZH8
 
 | |
9YGW
 
 | | Perdeuterated E. coli YajL, 100K | | Descriptor: | CHLORIDE ION, Chaperone YajL, MAGNESIUM ION | | Authors: | Lin, J, Kovalevsky, A, Walker, A.R, Wilson, M.A. | | Deposit date: | 2025-09-29 | | Release date: | 2025-11-26 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (0.88 Å) | | Cite: | Environmental Contributions to Proton Sharing in Protein Low-Barrier Hydrogen Bonds.
Biochemistry, 65, 2026
|
|
9YKF
 
 | | Crystal Structure of E. coli D23N YajL | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Lin, J, Kovalevsky, A, Walker, A.R, Wilson, M.A. | | Deposit date: | 2025-10-07 | | Release date: | 2025-10-22 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (0.98 Å) | | Cite: | Environmental Contributions to Proton Sharing in Protein Low-Barrier Hydrogen Bonds.
Biochemistry, 65, 2026
|
|
9Z0M
 
 | |
9ZMD
 
 | | Equine Serum Albumin with Copper(II) | | Descriptor: | COPPER (II) ION, SODIUM ION, SULFATE ION, ... | | Authors: | Handing, K.B, Bijak, V, Gucwa, M, Slawek, J, Minor, W, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2025-12-10 | | Release date: | 2025-12-24 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Equine Serum Albumin with Copper(II)
To Be Published
|
|
9Z6W
 
 | | The structure of TMD with 4 TARPs from all native AMPA receptor subtypes | | Descriptor: | (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, 6-[2-chloro-6-(trifluoromethoxy)phenyl]-1H-benzimidazol-2-ol, Glutamate receptor 2, ... | | Authors: | Park, J, Gouaux, E. | | Deposit date: | 2025-11-14 | | Release date: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (2.99 Å) | | Cite: | Efficient and rapid isolation of native AMPA receptor complexes for cryo-EM.
Protein Sci., 35, 2026
|
|
