4D2G
 
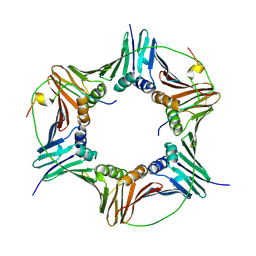 | | Crystal structure of human PCNA in complex with p15 peptide | | Descriptor: | P15, PROLIFERATING CELL NUCLEAR ANTIGEN | | Authors: | DeBiasio, A, Ibanez, A, Mortuza, G, Molina, R, Cordeiro, T.N, Castillo, F, Villate, M, Merino, N, Lelli, M, Diercks, T, Luque, I, Bernardo, P, Montoya, G, Blanco, F.J. | | Deposit date: | 2014-05-09 | | Release date: | 2015-03-18 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structure of P15(Paf)-PCNA Complex and Implications for Clamp Sliding During DNA Replication and Repair.
Nat.Commun., 6, 2015
|
|
1L1C
 
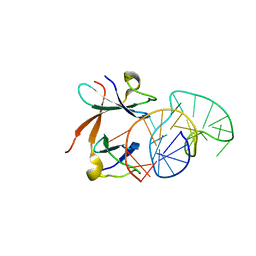 | | Structure of the LicT Bacterial Antiterminator Protein in Complex with its RNA Target | | Descriptor: | Transcription antiterminator licT, licT mRNA antiterminator hairpin | | Authors: | Yang, Y, Declerck, N, Manival, X, Aymerich, S, Kochoyan, M. | | Deposit date: | 2002-02-15 | | Release date: | 2002-03-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the LicT-RNA antitermination complex: CAT clamping RAT.
EMBO J., 21, 2002
|
|
1L5V
 
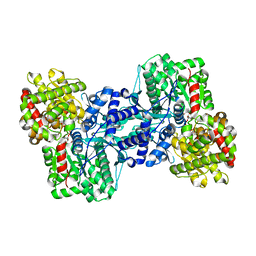 | | Crystal Structure of the Maltodextrin Phosphorylase complexed with Glucose-1-phosphate | | Descriptor: | 1-O-phosphono-alpha-D-glucopyranose, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MALTODEXTRIN PHOSPHORYLASE, ... | | Authors: | Geremia, S, Campagnolo, M, Schinzel, R, Johnson, L.N. | | Deposit date: | 2002-03-08 | | Release date: | 2002-04-10 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Enzymatic catalysis in crystals of Escherichia coli maltodextrin phosphorylase
J.Mol.Biol., 322, 2002
|
|
1L5W
 
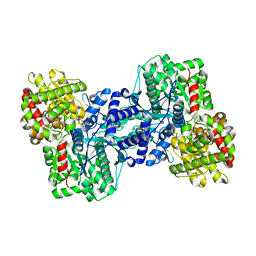 | | Crystal Structure of the Maltodextrin Phosphorylase Complexed with the Products of the Enzymatic Reaction between Glucose-1-phosphate and Maltotetraose | | Descriptor: | MALTODEXTRIN PHOSPHORYLASE, PHOSPHATE ION, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Geremia, S, Campagnolo, M, Schinzel, R, Johnson, L.N. | | Deposit date: | 2002-03-08 | | Release date: | 2002-04-10 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Enzymatic catalysis in crystals of Escherichia coli maltodextrin phosphorylase
J.Mol.Biol., 322, 2002
|
|
2V76
 
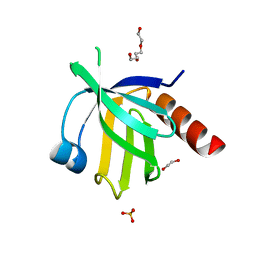 | | Crystal structure of the human dok1 PTB domain | | Descriptor: | 1,2-ETHANEDIOL, DOCKING PROTEIN 1, GLYCEROL, ... | | Authors: | Oxley, C.L, Anthis, N.J, Lowe, E.D, Campbell, I.D, Wegener, K.L. | | Deposit date: | 2007-07-26 | | Release date: | 2008-01-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | An Integrin Phosphorylation Switch: The Effect of {Beta}3 Integrin Tail Phosphorylation on Dok1 and Talin Binding.
J.Biol.Chem., 283, 2008
|
|
2VA6
 
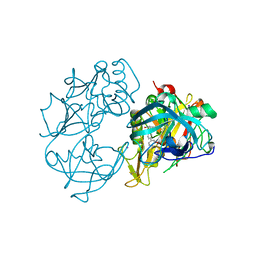 | | X-ray crystal structure of beta secretase complexed with compound 24 | | Descriptor: | (6S)-2-amino-6-(3'-methoxybiphenyl-3-yl)-3,6-dimethyl-5,6-dihydropyrimidin-4(3H)-one, BETA SECRETASE 1, IODIDE ION | | Authors: | Edwards, P.D, Albert, J.S, Sylvester, M, Aharony, D, Andisik, D, Callaghan, O, Campbell, J.B, Carr, R.A, Chessari, G, Congreve, M, Frederickson, M, Folmer, R.H.A, Geschwindner, S, Koether, G, Kolmodin, K, Krumrine, J, Mauger, R.C, Murray, C.W, Olsson, L.L, Patel, S, Spear, N, Tian, G. | | Deposit date: | 2007-08-30 | | Release date: | 2007-11-13 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Application of Fragment-Based Lead Generation to the Discovery of Novel, Cyclic Amidine Beta-Secretase Inhibitors with Nanomolar Potency, Cellular Activity, and High Ligand Efficiency.
J.Med.Chem., 50, 2007
|
|
1L6I
 
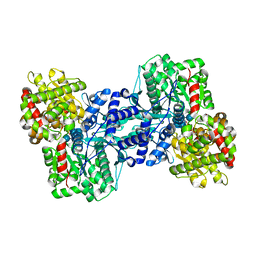 | | Crystal Structure of the Maltodextrin Phosphorylase complexed with the products of the enzymatic reaction between glucose-1-phosphate and maltopentaose | | Descriptor: | MALTODEXTRIN PHOSPHORYLASE, PHOSPHATE ION, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Geremia, S, Campagnolo, M, Schinzel, R, Johnson, L.N. | | Deposit date: | 2002-03-11 | | Release date: | 2002-04-10 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Enzymatic catalysis in crystals of Escherichia coli maltodextrin phosphorylase
J.Mol.Biol., 322, 2002
|
|
6LNP
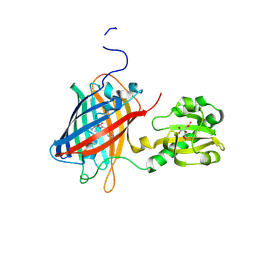 
 | | Crystal structure of citrate Biosensor | | Descriptor: | CITRIC ACID, Fusion protein of Green fluorescent protein and Sensor histidine kinase CitA | | Authors: | Wen, Y, Campbell, R. | | Deposit date: | 2019-12-31 | | Release date: | 2020-05-06 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.993 Å) | | Cite: | High-Performance Intensiometric Direct- and Inverse-Response Genetically Encoded Biosensors for Citrate.
Acs Cent.Sci., 6, 2020
|
|
8QUR
 
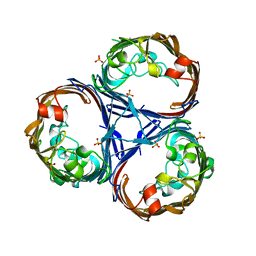 | | Crystal structure of Ompk36 GD at 3500 eV with no absorption corrections | | Descriptor: | OmpK36, SULFATE ION | | Authors: | Duman, R, Wagner, A, Beis, K, Wong, J. | | Deposit date: | 2023-10-16 | | Release date: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Ray-tracing analytical absorption correction for X-ray crystallography based on tomographic reconstructions.
J.Appl.Crystallogr., 57, 2024
|
|
8QVV
 
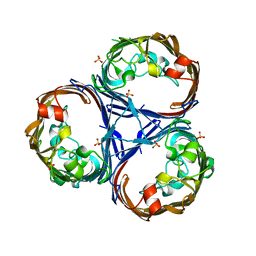 | | Crystal structure of Ompk36 GD at 3500 eV based on analytical absorption corrections | | Descriptor: | OmpK36, SULFATE ION | | Authors: | Duman, R, Wagner, A, Beis, K, Wong, J. | | Deposit date: | 2023-10-18 | | Release date: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Ray-tracing analytical absorption correction for X-ray crystallography based on tomographic reconstructions.
J.Appl.Crystallogr., 57, 2024
|
|
8QUQ
 
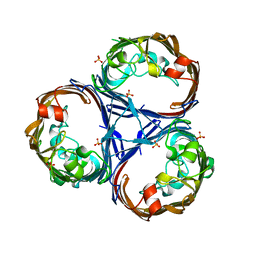 | | Crystal structure of Ompk36 GD at 3500 eV based on spherical harmonics absorption corrections | | Descriptor: | OmpK36, SULFATE ION | | Authors: | Duman, R, Wagner, A, Beis, K, Wong, J. | | Deposit date: | 2023-10-16 | | Release date: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Ray-tracing analytical absorption correction for X-ray crystallography based on tomographic reconstructions.
J.Appl.Crystallogr., 57, 2024
|
|
8CPM
 
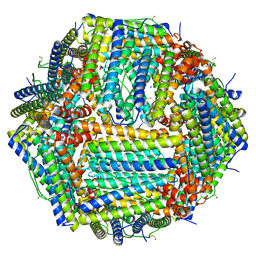 | | Human apoferritin after 405 nm laser exposure | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.81 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPX
 
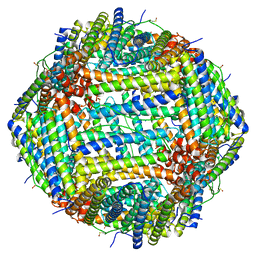 | | Human apoferritin after 488 nm laser exposure in presence of rsEGFP2 | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.76 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPV
 
 | | Human apoferritin | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.76 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPS
 
 | | Human apoferritin | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.82 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPW
 
 | | Human apoferritin after 405 nm + 488 nm laser exposure in presence of rsEGFP2 | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.79 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPU
 
 | | Human apoferritin after 561 nm laser exposure | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.76 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPT
 
 | | Human apoferritin after 488 nm laser exposure | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.79 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8QVS
 
 | | Crystal structure of Ompk36 GD at 3500 eV based on a combination of spherical harmonics and analytical absorption corrections | | Descriptor: | OmpK36, SULFATE ION | | Authors: | Duman, R, Wagner, A, Beis, K, Wong, J. | | Deposit date: | 2023-10-18 | | Release date: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Ray-tracing analytical absorption correction for X-ray crystallography based on tomographic reconstructions.
J.Appl.Crystallogr., 57, 2024
|
|
6XEZ
 
 | | Structure of SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ALUMINUM FLUORIDE, CHAPSO, ... | | Authors: | Chen, J, Malone, B, Llewellyn, E.C, Campbell, E.A, Darst, S.A. | | Deposit date: | 2020-06-14 | | Release date: | 2020-07-29 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural Basis for Helicase-Polymerase Coupling in the SARS-CoV-2 Replication-Transcription Complex.
Cell, 182, 2020
|
|
8YRM
 
 | | Iota toxin Ib pore serine-clamp mutant | | Descriptor: | CALCIUM ION, Iota toxin component Ib | | Authors: | Ninomiya, Y, Yoshida, T, Yamada, T, Kishikawa, J, Tsuge, H. | | Deposit date: | 2024-03-21 | | Release date: | 2025-03-26 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (2.36 Å) | | Cite: | Serine clamp of Clostridium perfringens binary toxin BECb (CPILEb)-pore confers cytotoxicity and enterotoxicity.
To Be Published
|
|
6YA9
 
 | | Crystal structure of rsGCaMP in the ON state (non-illuminated) | | Descriptor: | CALCIUM ION, rsCGaMP | | Authors: | Janowski, R, Fuenzalida-Werner, J.P, Mishra, K, Stiel, A.C, Niessing, D. | | Deposit date: | 2020-03-11 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Genetically encoded photo-switchable molecular sensors for optoacoustic and super-resolution imaging.
Nat.Biotechnol., 2021
|
|
4WFZ
 
 | |
4WFY
 
 | |
8BLR
 
 | | G13D mutant of KRAS4b (2-169) bound to GDP with the switch-I in fully open conformation | | Descriptor: | GTPase KRas, N-terminally processed, GUANOSINE-5'-DIPHOSPHATE | | Authors: | Moche, M, Jungholm, O, Strandback, E, Ampah-Korsah, H, Nyman, T, Orwar, O. | | Deposit date: | 2022-11-10 | | Release date: | 2024-05-22 | | Last modified: | 2024-09-11 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Novel druggable space in human KRAS G13D discovered using structural bioinformatics and a P-loop targeting monoclonal antibody.
Sci Rep, 14, 2024
|
|
