9CBL
 
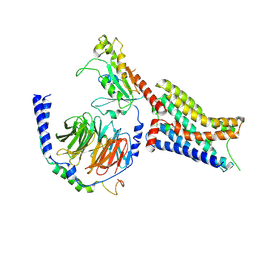 | | Cryo-EM structure of epinephrine-bound alpha-2A-adrenergic receptor in complex with heterotrimeric Gi-protein | | Descriptor: | Endolysin,Alpha-2A adrenergic receptor, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Lou, J.S, Su, M, Wang, J, Do, H.N, Miao, Y, Huang, X.Y. | | Deposit date: | 2024-06-19 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Distinct binding conformations of epinephrine with alpha- and beta-adrenergic receptors.
Exp.Mol.Med., 56, 2024
|
|
9CBM
 
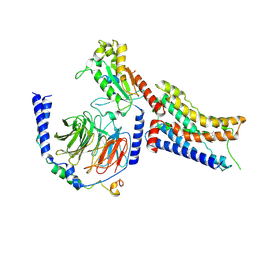 | | Cryo-EM structure of dexmedetomidine-bound alpha-2A-adrenergic receptor in complex with heterotrimeric Gi-protein | | Descriptor: | 4-[(1~{S})-1-(2,3-dimethylphenyl)ethyl]-1~{H}-imidazole, Endolysin,Alpha-2A adrenergic receptor, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Lou, J.S, Su, M, Wang, J, Do, H.N, Miao, Y, Huang, X.Y. | | Deposit date: | 2024-06-19 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Distinct binding conformations of epinephrine with alpha- and beta-adrenergic receptors.
Exp.Mol.Med., 56, 2024
|
|
3FAX
 
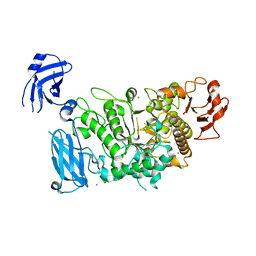 | |
3FDM
 
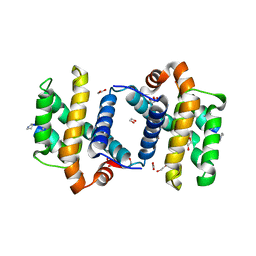 | | alpha/beta foldamer in complex with Bcl-xL | | Descriptor: | 1,2-ETHANEDIOL, Apoptosis regulator Bcl-X, alpha/beta-peptide foldamer | | Authors: | Fairlie, W.D, Lee, E.F, Smith, B.J, Czabotar, P.E, Colman, P.M, Sadowsky, J.D, Peterson-Kaufman, K.J, Gellman, S.H. | | Deposit date: | 2008-11-26 | | Release date: | 2009-03-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | High-Resolution Structural Characterization of a Helical alpha/beta-Peptide Foldamer Bound to the Anti-Apoptotic Protein Bcl-x(L)
Angew.Chem.Int.Ed.Engl., 48, 2009
|
|
2AX6
 
 | | Crystal Structure Of The Androgen Receptor Ligand Binding Domain T877A Mutant In Complex With Hydroxyflutamide | | Descriptor: | Androgen receptor, HYDROXYFLUTAMIDE | | Authors: | Bohl, C.E, Miller, D.D, Chen, J, Bell, C.E, Dalton, J.T. | | Deposit date: | 2005-09-03 | | Release date: | 2005-09-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural Basis for Accommodation of Nonsteroidal Ligands in the Androgen Receptor
J.Biol.Chem., 280, 2005
|
|
3FC5
 
 | | G586S mutant nNOSoxy | | Descriptor: | 5,6,7,8-TETRAHYDROBIOPTERIN, ARGININE, Nitric oxide synthase, ... | | Authors: | Bruckmann, C, Mowat, C.G. | | Deposit date: | 2008-11-21 | | Release date: | 2009-11-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Oxygen Activation in Neuronal NO Synthase: Stabilisation of a Novel Intermediate in the G586S Mutant
To be Published
|
|
3FHE
 
 | |
2BIJ
 
 | | Crystal structure of the human protein tyrosine phosphatase PTPN5 (STEP, striatum enriched enriched Phosphatase) | | Descriptor: | SULFATE ION, TYROSINE-PROTEIN PHOSPHATASE, NON-RECEPTOR TYPE 5 | | Authors: | Barr, A.J, Debreczeni, J.E, Eswaran, J, Smee, C, Burgess, N, Gileadi, O, Sundstrom, M, Arrowsmith, C, Edwards, A, Knapp, S, von Delft, F. | | Deposit date: | 2005-01-21 | | Release date: | 2005-03-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal structures and inhibitor identification for PTPN5, PTPRR and PTPN7: a family of human MAPK-specific protein tyrosine phosphatases.
Biochem. J., 395, 2006
|
|
9C10
 
 | | AMG 193, a clinical stage MTA-cooperative PRMT5 inhibitor, drives anti-tumor activity preclinically and in patients with MTAP-deleted cancers | | Descriptor: | (4-amino-1,3-dihydrofuro[3,4-c][1,7]naphthyridin-8-yl){(3S)-3-[4-(trifluoromethyl)phenyl]morpholin-4-yl}methanone, 5'-DEOXY-5'-METHYLTHIOADENOSINE, DIMETHYL SULFOXIDE, ... | | Authors: | Ghimire-Rijal, S, Mukund, S. | | Deposit date: | 2024-05-28 | | Release date: | 2024-10-02 | | Last modified: | 2025-01-22 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | AMG 193, a Clinical Stage MTA-Cooperative PRMT5 Inhibitor, Drives Antitumor Activity Preclinically and in Patients with MTAP-Deleted Cancers.
Cancer Discov, 15, 2025
|
|
3F1K
 
 | |
9EJM
 
 | |
2BCJ
 
 | |
1VZT
 
 | | ROLES OF INDIVIDUAL RESIDUES OF ALPHA-1,3 GALACTOSYLTRANSFERASES IN SUBSTRATE BINDING AND CATALYSIS | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MANGANESE (II) ION, N-ACETYLLACTOSAMINIDE ALPHA-1,3-GALACTOSYLTRANSFERASE, ... | | Authors: | Zhang, Y, Swaminathan, G.J, Deshpande, A, Natesh, R, Xie, X, Acharya, K.R, Brew, K. | | Deposit date: | 2004-05-26 | | Release date: | 2005-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Roles of Individual Enzyme-Substrate Interactions by Alpha-1,3-Galactosyltransferase in Catalysis and Specificity.
Biochemistry, 42, 2003
|
|
9DQJ
 
 | | CryoEM structure of Gq-coupled MRGPRD with a new agonist EP-3945 | | Descriptor: | 2-({1-[1-(4-methoxyphenyl)cyclopropane-1-carbonyl]piperidin-4-yl}amino)quinazolin-4(3H)-one, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Cao, C, Wang, C, Liu, Y, Fay, J.F, Roth, B.L. | | Deposit date: | 2024-09-24 | | Release date: | 2024-12-11 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | High-affinity agonists reveal recognition motifs for the MRGPRD GPCR.
Cell Rep, 43, 2024
|
|
9DTR
 | |
9D61
 
 | | Kappa opioid receptor:Galphai protein in complex with inverse agonist JDTic , no scFv16 | | Descriptor: | (3R)-7-hydroxy-N-{(2S)-1-[(3R,4R)-4-(3-hydroxyphenyl)-3,4-dimethylpiperidin-1-yl]-3-methylbutan-2-yl}-1,2,3,4-tetrahydroisoquinoline-3-carboxamide, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Gati, C, Motiwala, Z, Tyson, A.S, Styrpejko, D, Han, G.W, Khan, S, Ramos-Gonzalez, N, Shenvi, R, Majumdar, S. | | Deposit date: | 2024-08-14 | | Release date: | 2025-01-15 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (3.58 Å) | | Cite: | Molecular mechanisms of inverse agonism via kappa-opioid receptor-G protein complexes.
Nat.Chem.Biol., 21, 2025
|
|
3F5Q
 
 | | CRYSTAL STRUCTURE OF putative short chain dehydrogenase from Escherichia coli CFT073 | | Descriptor: | dehydrogenase | | Authors: | Malashkevich, V.N, Toro, R, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-11-04 | | Release date: | 2009-01-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structure of an uncharacterized protein
to be published
|
|
3F92
 
 | | Crystal structure of ubiquitin-conjugating enzyme E2-25kDa (Huntington Interacting Protein 2) M172A mutant crystallized at pH 8.5 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, BETA-MERCAPTOETHANOL, CALCIUM ION, ... | | Authors: | Wilson, R.C, Hughes, R.C, Flatt, J.W, Meehan, E.J, Ng, J.D, Twigg, P.D. | | Deposit date: | 2008-11-13 | | Release date: | 2008-11-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Structure of full-length ubiquitin-conjugating enzyme E2-25K (huntingtin-interacting protein 2).
Acta Crystallogr.,Sect.F, 65, 2009
|
|
9EBS
 
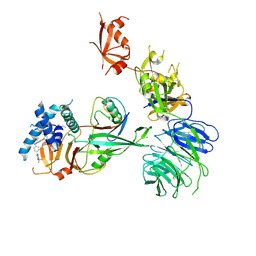 | | Cryo-EM structure of USP1-UAF1-Ubiquitin in complex with TNG348 | | Descriptor: | 2-(4-cyclopropyl-6-methoxypyrimidin-5-yl)-9-({4-[1-methyl-4-(trifluoromethyl)-1H-imidazol-2-yl]phenyl}methyl)-7-(2,2,2-trifluoroethyl)-7,9-dihydro-8H-purin-8-imine, 3-(methanesulfonyl)propan-1-amine, Ubiquitin, ... | | Authors: | Whittington, D.A. | | Deposit date: | 2024-11-13 | | Release date: | 2025-01-29 | | Last modified: | 2025-05-14 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Characterization of TNG348: A Selective, Allosteric USP1 Inhibitor That Synergizes with PARP Inhibitors in Tumors with Homologous Recombination Deficiency.
Mol.Cancer Ther., 24, 2025
|
|
9CLW
 
 | | Cryo-EM structure of Gq-coupled FFA2 in complex with TUG-1375 and 4-CMTB | | Descriptor: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, (2~{S})-2-(4-chlorophenyl)-3-methyl-~{N}-(1,3-thiazol-2-yl)butanamide, CHOLESTEROL, ... | | Authors: | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | Deposit date: | 2024-07-12 | | Release date: | 2025-06-25 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.19 Å) | | Cite: | Allosteric modulation and biased signalling at free fatty acid receptor 2.
Nature, 2025
|
|
9CM7
 
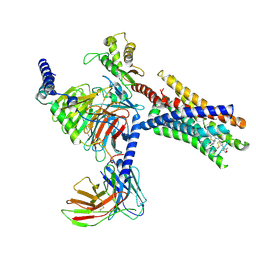 | | Cryo-EM structure of Gi-coupled FFA2 in complex with TUG-1375 and AZ-1729 | | Descriptor: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, Free fatty acid receptor 2, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | Deposit date: | 2024-07-12 | | Release date: | 2025-06-25 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.29 Å) | | Cite: | Allosteric modulation and biased signalling at free fatty acid receptor 2.
Nature, 2025
|
|
2BKM
 
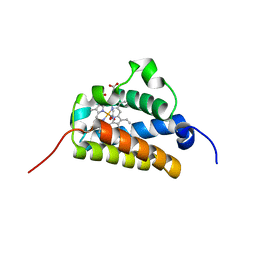 | | Crystal structure of the truncated hemoglobin from Geobacillus stearothermophilus | | Descriptor: | ACETATE ION, OXYGEN MOLECULE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Ilari, A, Kjelgaard, P, von Wachenfeldt, C, Boffi, A, Chiancone, E. | | Deposit date: | 2006-02-08 | | Release date: | 2006-11-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure and Ligand Binding Properties of the Truncated Hemoglobin from Geobacillus Stearothermophilus
Arch.Biochem.Biophys., 457, 2007
|
|
9CM3
 
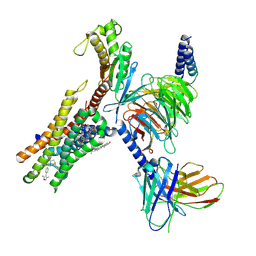 | | Cryo-EM structure of Gq-coupled FFA2 in complex with TUG-1375 and compound 187 | | Descriptor: | (2R,4R)-2-(2-chlorophenyl)-3-[4-(3,5-dimethyl-1,2-oxazol-4-yl)phenyl]carbonyl-1,3-thiazolidine-4-carboxylic acid, 4-[(2R,6S)-2,6-dimethylmorpholin-4-yl]-7-(2-fluorobenzene-1-sulfonyl)-2-methyl-5H-pyrrolo[3,2-d]pyrimidin-6-amine, CHOLESTEROL, ... | | Authors: | Zhang, X, Tikhonova, I, Milligan, G, Zhang, C. | | Deposit date: | 2024-07-12 | | Release date: | 2025-06-25 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Allosteric modulation and biased signalling at free fatty acid receptor 2.
Nature, 2025
|
|
3F7S
 
 | |
3FFK
 
 | | Crystal structure of human Gelsolin domains G1-G3 bound to Actin | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, actin, ... | | Authors: | Chumnarnsilpa, S, Robinson, R.C, Burtnick, L.D. | | Deposit date: | 2008-12-03 | | Release date: | 2009-10-06 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Ca2+ binding by domain 2 plays a critical role in the activation and stabilization of gelsolin.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
