3FW1
 
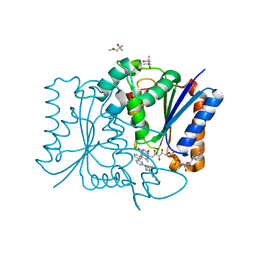 | | Quinone Reductase 2 | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, 4-(4-METHYL-PIPERAZIN-1-YLMETHYL)-N-[4-METHYL-3-(4-PYRIDIN-3-YL-PYRIMIDIN-2-YLAMINO)-PHENYL]-BENZAMIDE, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Winger, J.A, Hantschel, O, Superti-Furga, G, Kuriyan, J. | | Deposit date: | 2009-01-16 | | Release date: | 2009-03-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The structure of the leukemia drug imatinib bound to human quinone reductase 2 (NQO2).
Bmc Struct.Biol., 9, 2009
|
|
4I2V
 
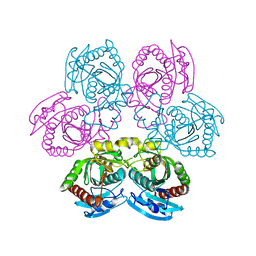 | | X-ray structure of the unliganded uridine phosphorylase from Yersinia pseudotuberculosis at 2.12A resolution | | Descriptor: | Uridine phosphorylase | | Authors: | Lashkov, A.A, Balaev, V.V, Prokofev, I.I, Betzel, C, Gabdoulkhakov, A.G, Mikhailov, A.M. | | Deposit date: | 2012-11-23 | | Release date: | 2013-12-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.121 Å) | | Cite: | X-ray structure of the unliganded uridine phosphorylase from Yersinia pseudotuberculosis at 2.12A resolution
To be Published
|
|
4I1M
 
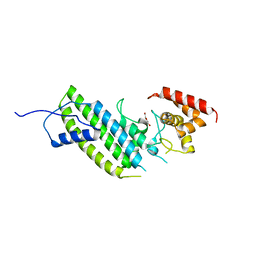 | | Crystal structure of the Legionella pneumophila GAP domain of LepB | | Descriptor: | DI(HYDROXYETHYL)ETHER, LepB | | Authors: | Streller, A, Gazdag, E.M, Vetter, I.R, Goody, R.S, Itzen, A. | | Deposit date: | 2012-11-21 | | Release date: | 2013-01-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.804 Å) | | Cite: | Mechanism of Rab1b deactivation by the Legionella pneumophila GAP LepB.
Embo Rep., 14, 2013
|
|
4J0R
 
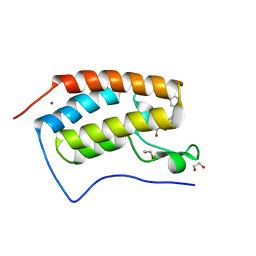 | | Crystal Structure of the first bromodomain of human BRD4 in complex with a 3,5-dimethylisoxazol ligand | | Descriptor: | 1,2-ETHANEDIOL, 3-(3,5-dimethyl-1,2-oxazol-4-yl)-5-[(R)-hydroxy(phenyl)methyl]phenol, Bromodomain-containing protein 4, ... | | Authors: | Filippakopoulos, P, Picaud, S, Qi, J, Felletar, I, Hewings, D.S, von Delft, F, Conway, S.J, Bountra, C, Arrowsmith, C.H, Edwards, A, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2013-01-31 | | Release date: | 2013-02-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Optimization of 3,5-dimethylisoxazole derivatives as potent bromodomain ligands.
J.Med.Chem., 56, 2013
|
|
4IZU
 
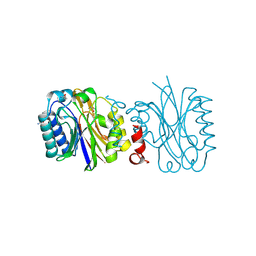 | |
4J3I
 
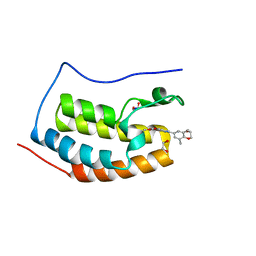 | | X-ray crystal structure of bromodomain complex to 1.24 A resolution | | Descriptor: | 1,2-ETHANEDIOL, 2-[4-(2-hydroxyethoxy)-3,5-dimethylphenyl]-5,7-dimethoxyquinazolin-4(3H)-one, Bromodomain-containing protein 4, ... | | Authors: | Stein, A.J, White, A, Suto, R.K. | | Deposit date: | 2013-02-05 | | Release date: | 2014-01-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | RVX-208, an Inducer of ApoA-I in Humans, Is a BET Bromodomain Antagonist.
Plos One, 8, 2013
|
|
3E2I
 
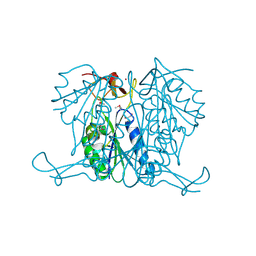 | | Crystal structure of Thymidine Kinase from S. aureus | | Descriptor: | GLYCEROL, Thymidine kinase, ZINC ION | | Authors: | Lam, R, Johns, K, Battaile, K.P, Romanov, V, Lam, K, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2008-08-05 | | Release date: | 2009-08-11 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Crystal structure of Thymidine Kinase from S. aureus
To be Published
|
|
4ITC
 
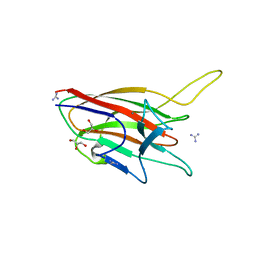 | | Crystal Structure Analysis of the K1 Cleaved Adhesin domain of Lys-gingipain (Kgp) from Porphyromonas gingivalis W83 | | Descriptor: | CALCIUM ION, GLYCEROL, GUANIDINE, ... | | Authors: | Ganuelas, L.A, Li, N, Hunter, N, Collyer, C.A. | | Deposit date: | 2013-01-18 | | Release date: | 2013-09-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The lysine gingipain adhesin domains from Porphyromonas gingivalis interact with erythrocytes and albumin: Structures correlate to function.
Eur J Microbiol Immunol (Bp), 3, 2013
|
|
4IVT
 
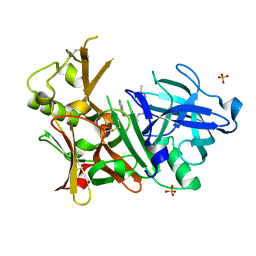 | | Crystal structure of BACE1 with its inhibitor | | Descriptor: | Beta-secretase 1, N-{N-[4-(acetylamino)-3,5-dichlorobenzyl]carbamimidoyl}-2-(1H-indol-1-yl)acetamide, SULFATE ION | | Authors: | Chen, T.T, Li, L, Chen, W.Y, Xu, Y.C. | | Deposit date: | 2013-01-23 | | Release date: | 2013-11-13 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Virtual screening and structure-based discovery of indole acylguanidines as potent beta-secretase (BACE1) inhibitors
Molecules, 18, 2013
|
|
3EHZ
 
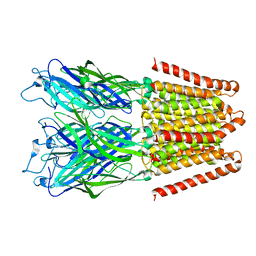 | |
4IZE
 
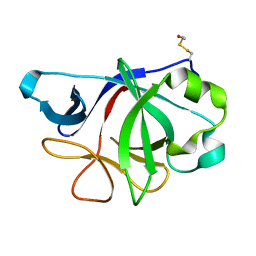 | | Crystal Structure of IL-36gamma | | Descriptor: | Interleukin-36 gamma | | Authors: | Guenther, S, Sundberg, E.J. | | Deposit date: | 2013-01-29 | | Release date: | 2014-06-25 | | Last modified: | 2014-07-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular Determinants of Agonist and Antagonist Signaling through the IL-36 Receptor.
J.Immunol., 193, 2014
|
|
3EBI
 
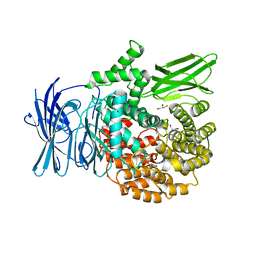 | | Structure of the M1 Alanylaminopeptidase from malaria complexed with the phosphinate dipeptide analog | | Descriptor: | (2S)-3-[(R)-[(1S)-1-amino-3-phenylpropyl](hydroxy)phosphoryl]-2-benzylpropanoic acid, GLYCEROL, M1 family aminopeptidase, ... | | Authors: | McGowan, S, Porter, C.J, Buckle, A.M, Whisstock, J.C. | | Deposit date: | 2008-08-27 | | Release date: | 2009-01-27 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for the inhibition of the essential Plasmodium falciparum M1 neutral aminopeptidase
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3ECD
 
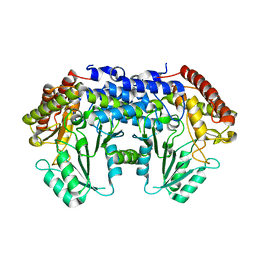 | |
3EG3
 
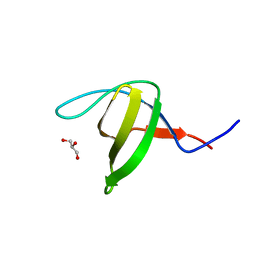 | | Crystal structure of the N114A mutant of ABL-SH3 domain | | Descriptor: | GLYCEROL, Proto-oncogene tyrosine-protein kinase ABL1 | | Authors: | Camara-Artigas, A. | | Deposit date: | 2008-09-10 | | Release date: | 2009-09-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Role of interfacial water molecules in proline-rich ligand recognition by the Src homology 3 domain of Abl.
J.Biol.Chem., 285, 2010
|
|
3EQQ
 
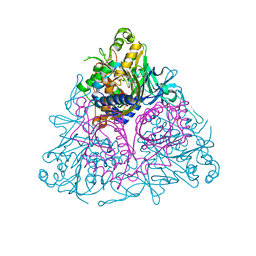 | | Apo Toluene 2,3-Dioxygenase | | Descriptor: | Benzene 1,2-dioxygenase subunit alpha, Benzene 1,2-dioxygenase subunit beta, FE (II) ION, ... | | Authors: | Friemann, R, Lee, K, Brown, E.N, Gibson, D.T, Eklund, H, Ramaswamy, S. | | Deposit date: | 2008-10-01 | | Release date: | 2009-03-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structures of the multicomponent Rieske non-heme iron toluene 2,3-dioxygenase enzyme system
Acta Crystallogr.,Sect.D, 65, 2009
|
|
4ISW
 
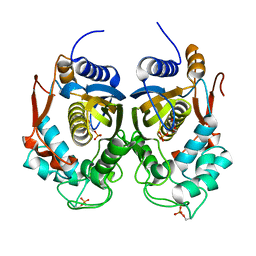 | | Crystal Structure of Phosphorylated C.elegans Thymidylate Synthase in Complex with dUMP | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, Thymidylate synthase | | Authors: | Wilk, P, Dowiercial, A, Banaszak, K, Jarmula, A, Rypniewski, W, Rode, W. | | Deposit date: | 2013-01-17 | | Release date: | 2013-12-11 | | Last modified: | 2014-01-15 | | Method: | X-RAY DIFFRACTION (3.14 Å) | | Cite: | Crystal structure of phosphoramide-phosphorylated thymidylate synthase reveals pSer127, reflecting probably pHis to pSer phosphotransfer.
Bioorg.Chem., 52C, 2013
|
|
4IVF
 
 | | Crystal structure of glutathione transferase homolog from Lodderomyces elongisporus, target EFI-501753, with two GSH per subunit | | Descriptor: | CITRIC ACID, GLUTATHIONE, Putative uncharacterized protein | | Authors: | Vetting, M.W, Toro, R, Bhosle, R, Al Obaidi, N.F, Morisco, L.L, Wasserman, S.R, Sojitra, S, Washington, E, Scott Glenn, A, Chowdhury, S, Evans, B, Hammonds, J, Hillerich, B, Love, J, Seidel, R.D, Imker, H.J, Armstrong, R.N, Gerlt, J.A, Almo, S.C, Enzyme Function Initiative (EFI) | | Deposit date: | 2013-01-22 | | Release date: | 2013-02-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of glutathione transferase homolog from Lodderomyces elongisporus, target EFI-501753, with two GSH per subunit
To be Published
|
|
4IWH
 
 | |
4IYQ
 
 | |
4IYD
 
 | |
4J3Z
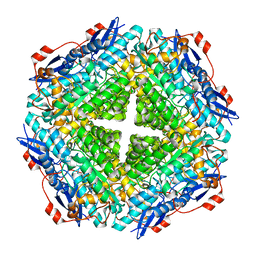 
 | | Crystal structure of mandelate racemase/muconate lactonizing enzyme from Jannaschia sp. CCS1 | | Descriptor: | Mandelate racemase/muconate lactonizing enzyme | | Authors: | Malashkevich, V.N, Bhosle, R, Toro, R, Hillerich, B, Gizzi, A, Garforth, S, Kar, A, Chan, M.K, Lafluer, J, Patel, H, Matikainen, B, Chamala, S, Lim, S, Celikgil, A, Villegas, G, Evans, B, Zenchek, W, Love, J, Fiser, A, Khafizov, K, Seidel, R, Bonanno, J.B, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2013-02-06 | | Release date: | 2013-03-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of mandelate racemase/muconate lactonizing enzyme from Jannaschia sp. CCS1
To be Published
|
|
3E2X
 
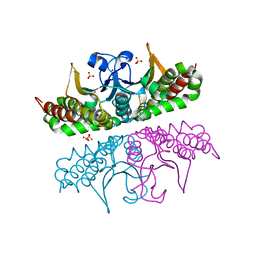 | | H. influenzae beta-carbonic anhydrase, variant V47A | | Descriptor: | Carbonic anhydrase 2, SULFATE ION, ZINC ION | | Authors: | Rowlett, R.S, Lee, J. | | Deposit date: | 2008-08-06 | | Release date: | 2009-08-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Evidence for a bicarbonate "escort" site in Haemophilus influenzae beta-carbonic anhydrase .
Biochemistry, 49, 2010
|
|
4IZV
 
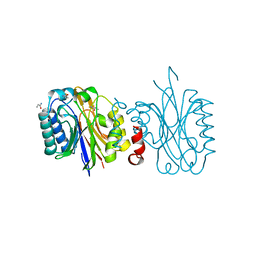 | |
3E4U
 
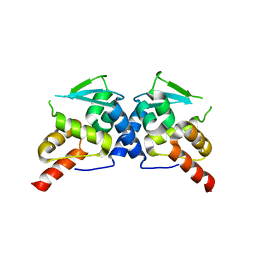 | | Crystal Structure of the Wild-Type Human BCL6 BTB/POZ Domain | | Descriptor: | B-cell lymphoma 6 protein | | Authors: | Stead, M.A, Rosbrook, G.O, Hadden, J.M, Trinh, C.H, Carr, S.B, Wright, S.C. | | Deposit date: | 2008-08-12 | | Release date: | 2008-12-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of the wild-type human BCL6 POZ domain.
Acta Crystallogr.,Sect.F, 64, 2008
|
|
4I4L
 
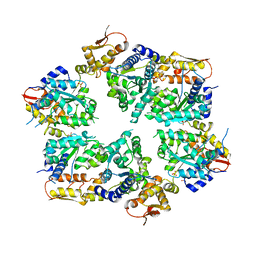 | | Crystal Structure of Nucleotide-Bound W-W-W ClpX Hexamer | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ATP-dependent Clp protease ATP-binding subunit ClpX, SULFATE ION | | Authors: | Glynn, S.E, Nager, A.R, Stinson, B.S, Schmitz, K.R, Baker, T.A, Sauer, R.T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-05-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.6981 Å) | | Cite: | Nucleotide Binding and Conformational Switching in the Hexameric Ring of a AAA+ Machine.
Cell(Cambridge,Mass.), 153, 2013
|
|
