7OZG
 
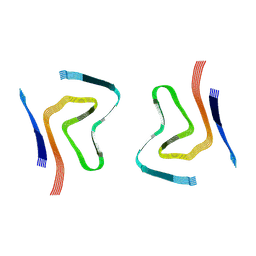 | |
8CSG
 
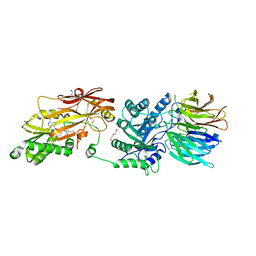 | | Human PRMT5:MEP50 structure with Fragment 1 and MTA Bound | | Descriptor: | 1,2-ETHANEDIOL, 5'-DEOXY-5'-METHYLTHIOADENOSINE, 6-bromo-1H-pyrrolo[3,2-b]pyridin-5-amine, ... | | Authors: | Gunn, R.J, Lawson, J.D, Smith, C.R. | | Deposit date: | 2022-05-12 | | Release date: | 2022-10-05 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Fragment optimization and elaboration strategies - the discovery of two lead series of PRMT5/MTA inhibitors from five fragment hits.
Rsc Med Chem, 13, 2022
|
|
7VQF
 
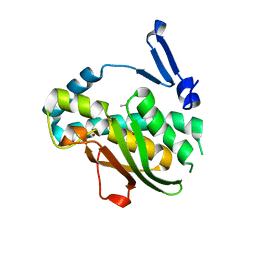 | | Phenol binding protein, MopR | | Descriptor: | ACETATE ION, PHENOL, Phenol sensing regulator, ... | | Authors: | Singh, J, Ray, S, Anand, R. | | Deposit date: | 2021-10-19 | | Release date: | 2022-09-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Phenol sensing in nature is modulated via a conformational switch governed by dynamic allostery.
J.Biol.Chem., 298, 2022
|
|
1ITM
 
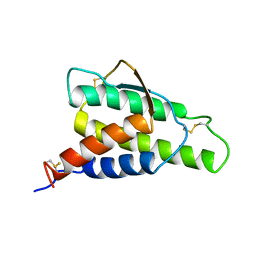 | | ANALYSIS OF THE SOLUTION STRUCTURE OF HUMAN INTERLEUKIN 4 DETERMINED BY HETERONUCLEAR THREE-DIMENSIONAL NUCLEAR MAGNETIC RESONANCE TECHNIQUES | | Descriptor: | INTERLEUKIN-4 | | Authors: | Redfield, C, Smith, L.J, Boyd, J, Lawrence, G.M.P, Edwards, R.G, Gershater, C.J, Smith, R.A.G, Dobson, C.M. | | Deposit date: | 1994-02-28 | | Release date: | 1994-05-31 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Analysis of the solution structure of human interleukin-4 determined by heteronuclear three-dimensional nuclear magnetic resonance techniques.
J.Mol.Biol., 238, 1994
|
|
3PSK
 
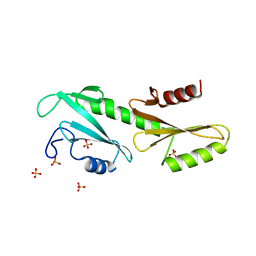 | |
1IRL
 
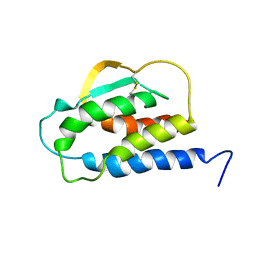 | | THE SOLUTION STRUCTURE OF THE F42A MUTANT OF HUMAN INTERLEUKIN 2 | | Descriptor: | INTERLEUKIN-2 | | Authors: | Mott, H.R, Baines, B.S, Hall, R.M, Cooke, R.M, Driscoll, P.C, Weir, M.P, Campbell, I.D. | | Deposit date: | 1995-08-25 | | Release date: | 1995-12-07 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the F42A mutant of human interleukin 2.
J.Mol.Biol., 247, 1995
|
|
1ITL
 
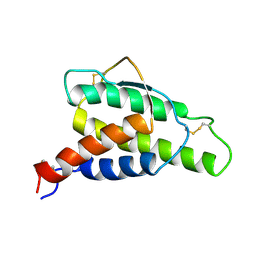 | | HUMAN INTERLEUKIN 4: THE SOLUTION STRUCTURE OF A FOUR-HELIX-BUNDLE PROTEIN | | Descriptor: | INTERLEUKIN-4 | | Authors: | Smith, L.J, Redfield, C, Boyd, J, Lawrence, G.M.P, Edwards, R.G, Smith, R.A.G, Dobson, C.M. | | Deposit date: | 1992-02-08 | | Release date: | 1993-04-15 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Human interleukin 4. The solution structure of a four-helix bundle protein.
J.Mol.Biol., 224, 1992
|
|
1HIJ
 
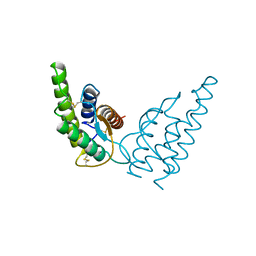 | |
8TO0
 
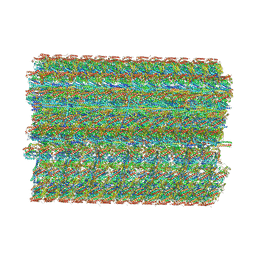 | | 48-nm repeating structure of doublets from mouse sperm flagella | | Descriptor: | Cilia- and flagella- associated protein 210, Cilia- and flagella-associated protein 107, Cilia- and flagella-associated protein 141, ... | | Authors: | Chen, Z, Shiozak, M, Hass, K.M, Skinner, W, Zhao, S, Guo, C, Polacco, B.J, Yu, Z, Krogan, N.J, Kaake, R.M, Vale, R.D, Agard, D.A. | | Deposit date: | 2023-08-02 | | Release date: | 2023-11-01 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (7.7 Å) | | Cite: | De novo protein identification in mammalian sperm using in situ cryoelectron tomography and AlphaFold2 docking.
Cell, 186, 2023
|
|
7EFA
 
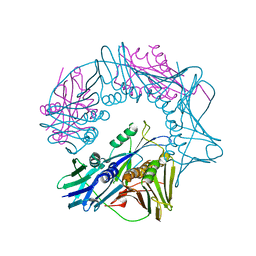 | |
5JCP
 
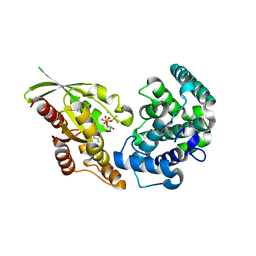 | | RhoGAP domain of ARAP3 in complex with RhoA in the transition state | | Descriptor: | Arf-GAP with Rho-GAP domain, ANK repeat and PH domain-containing protein 3,Linker,Transforming protein RhoA, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Bao, H, Li, F, Wang, C, Wang, N, Jiang, Y, Tang, Y, Wu, J, Shi, Y. | | Deposit date: | 2016-04-15 | | Release date: | 2016-06-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Basis for the Specific Recognition of RhoA by the Dual GTPase-activating Protein ARAP3
J.Biol.Chem., 291, 2016
|
|
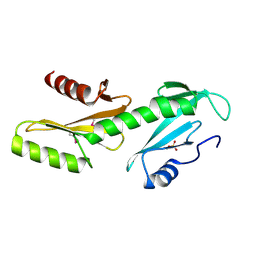
3PSJ
 | |
6C27
 
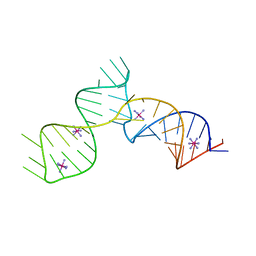 | | SAM-III riboswitch ON-state | | Descriptor: | COBALT HEXAMMINE(III), SAM-III riboswitch | | Authors: | Grigg, J.C, Price, I.R, Ke, A. | | Deposit date: | 2018-01-07 | | Release date: | 2019-01-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3.601 Å) | | Cite: | Evidence for two-tiered conformation selection in the SAM-III riboswitch
To Be Published
|
|
7WLR
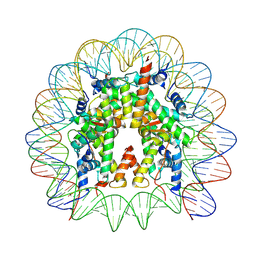 
 | | Cryo-EM structure of the nucleosome containing Komagataella pastoris histones | | Descriptor: | DNA (145-MER), Histone H2A, Histone H2B, ... | | Authors: | Fukushima, Y, Hatazawa, S, Hirai, S, Kujirai, T, Takizawa, Y, Kurumizaka, H. | | Deposit date: | 2022-01-13 | | Release date: | 2022-07-13 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.54 Å) | | Cite: | Structural and biochemical analyses of the nucleosome containing Komagataella pastoris histones.
J.Biochem., 172, 2022
|
|
5JHG
 
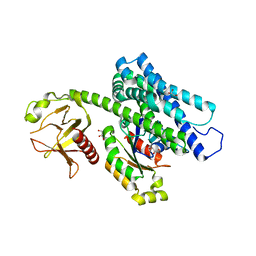 | | Crystal structure of the complex between the human RhoA and the DH/PH domain of human ARHGEF11 | | Descriptor: | GLYCEROL, Rho guanine nucleotide exchange factor 11, Transforming protein RhoA | | Authors: | Wang, R, Chen, Q, Zhang, H, Yan, Z, Li, J, Miao, L, Wang, F. | | Deposit date: | 2016-04-21 | | Release date: | 2017-04-26 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystallization and preliminary X-ray crystallographic analysis of a small GTPase RhoA bound with its inhibitor and ARHGEF11
To Be Published
|
|
1CF7
 
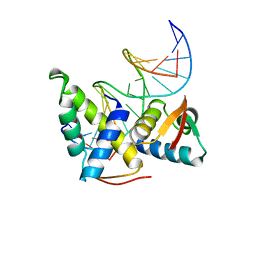 | | STRUCTURAL BASIS OF DNA RECOGNITION BY THE HETERODIMERIC CELL CYCLE TRANSCRIPTION FACTOR E2F-DP | | Descriptor: | DNA (5'-D(*AP*TP*TP*TP*TP*CP*GP*CP*GP*CP*GP*GP*TP*TP*TP*T)-3'), DNA (5'-D(*TP*AP*AP*AP*AP*CP*CP*GP*CP*GP*CP*GP*AP*AP*AP*A)-3'), PROTEIN (TRANSCRIPTION FACTOR DP-2), ... | | Authors: | Zheng, N, Fraenkel, E, Pabo, C.O, Pavletich, N.P. | | Deposit date: | 1999-03-24 | | Release date: | 1999-04-02 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis of DNA recognition by the heterodimeric cell cycle transcription factor E2F-DP.
Genes Dev., 13, 1999
|
|
4AQF
 
 | | X-ray crystallographic structure of Crimean-congo haemorrhagic fever virus nucleoprotein | | Descriptor: | NUCLEOPROTEIN, SULFATE ION | | Authors: | Wang, Y, Dutta, S, Karlberg, H, Devignot, S, Weber, F, Hao, Q, Tan, Y.J, Mirazimi, A, Kotaka, M. | | Deposit date: | 2012-04-17 | | Release date: | 2012-09-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure of Crimean-Congo Haemorraghic Fever Virus Nucleoprotein: Superhelical Homo-Oligomers and the Role of Caspase-3 Cleavage.
J.Virol., 86, 2012
|
|
8CTB
 
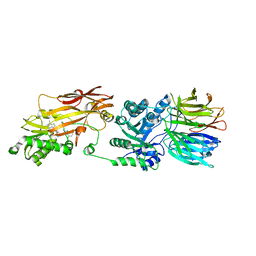 | | Human PRMT5:MEP50 structure with Fragment 3 and MTA Bound | | Descriptor: | 5'-DEOXY-5'-METHYLTHIOADENOSINE, 7-chloro-1-methyl-1H-benzimidazol-2-amine, Methylosome protein 50, ... | | Authors: | Gunn, R.J, Lawson, J.D, Smith, C.R. | | Deposit date: | 2022-05-13 | | Release date: | 2022-10-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Fragment optimization and elaboration strategies - the discovery of two lead series of PRMT5/MTA inhibitors from five fragment hits.
Rsc Med Chem, 13, 2022
|
|
7EYQ
 
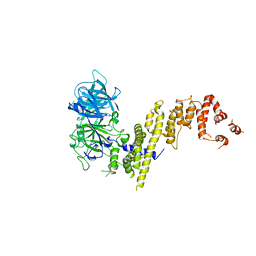 | |
1IAR
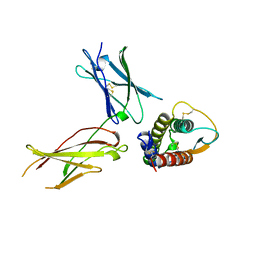 
 | | INTERLEUKIN-4 / RECEPTOR ALPHA CHAIN COMPLEX | | Descriptor: | PROTEIN (INTERLEUKIN-4 RECEPTOR ALPHA CHAIN), PROTEIN (INTERLEUKIN-4) | | Authors: | Hage, T, Sebald, W, Reinemer, P. | | Deposit date: | 1999-02-25 | | Release date: | 2000-03-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the interleukin-4/receptor alpha chain complex reveals a mosaic binding interface.
Cell(Cambridge,Mass.), 97, 1999
|
|
8CYI
 
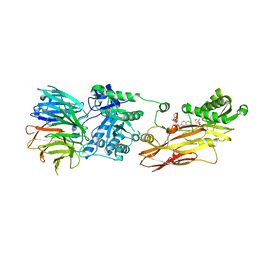 | | Cryo-EM structures and computational analysis for enhanced potency in MTA-synergic inhibition of human protein arginine methyltransferase 5 | | Descriptor: | 5'-DEOXY-5'-METHYLTHIOADENOSINE, Methylosome protein 50, N-[(2-aminoquinolin-7-yl)methyl]-9-(2-hydroxyethyl)-2,3,4,9-tetrahydro-1H-carbazole-6-carboxamide, ... | | Authors: | Yadav, G.P, Wei, Z, Xiaozhi, Y, Chenglong, L, Jiang, Q. | | Deposit date: | 2022-05-23 | | Release date: | 2023-04-12 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | Cryo-EM structure-based selection of computed ligand poses enables design of MTA-synergic PRMT5 inhibitors of better potency.
Commun Biol, 5, 2022
|
|
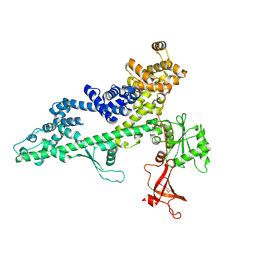
3PSI
 
 | |
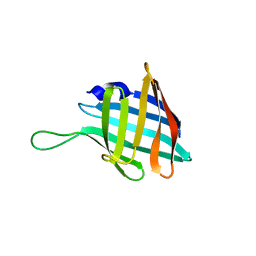
7QH3
 
 | |
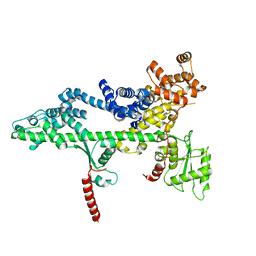
3PSF
 
 | |
8T5I
 
 | | Crystal structure of human WDR5 in complex with MR4397 | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, N-[(2S)-1-(6,7-dihydrothieno[3,2-c]pyridin-5(4H)-yl)-1-oxopentan-2-yl]-3-[(1H-imidazol-1-yl)methyl]benzamide, ... | | Authors: | Kimani, S, Dong, A, Li, F, Loppnau, P, Ackloo, S, Vedadi, M, Brown, P.J, Arrowsmith, C.H, Edwards, A.M, Halabelian, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2023-06-13 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of human WDR5 in complex with MR4397
To be published
|
|
