3BNF
 
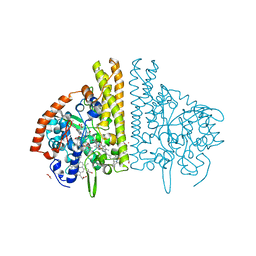 | | W. succinogenes NrfA Sulfite Complex | | 分子名称: | ACETATE ION, CALCIUM ION, Cytochrome c-552, ... | | 著者 | Lukat, P, Einsle, O. | | 登録日 | 2007-12-14 | | 公開日 | 2008-02-26 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Binding and Reduction of Sulfite by Cytochrome c Nitrite Reductase
Biochemistry, 47, 2008
|
|
3BTG
 
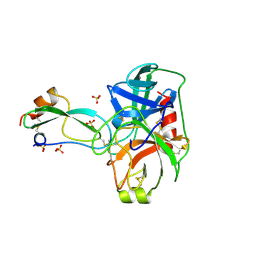 | | THE CRYSTAL STRUCTURES OF THE COMPLEXES BETWEEN BOVINE BETA-TRYPSIN AND TEN P1 VARIANTS OF BPTI | | 分子名称: | CALCIUM ION, PROTEIN (PANCREATIC TRYPSIN INHIBITOR), PROTEIN (TRYPSIN), ... | | 著者 | Helland, R, Otlewski, J, Sundheim, O, Dadlez, M, Smalas, A.O. | | 登録日 | 1999-03-10 | | 公開日 | 2000-03-13 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | The crystal structures of the complexes between bovine beta-trypsin and ten P1 variants of BPTI.
J.Mol.Biol., 287, 1999
|
|
3BTP
 
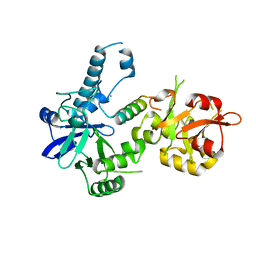 | | Crystal structure of Agrobacterium tumefaciens VirE2 in complex with its chaperone VirE1: a novel fold and implications for DNA binding | | 分子名称: | AMMONIUM ION, DI(HYDROXYETHYL)ETHER, Protein virE1, ... | | 著者 | Dym, O, Albeck, S, Unger, T, Elbaum, M, Israel Structural Proteomics Center (ISPC) | | 登録日 | 2007-12-30 | | 公開日 | 2008-08-19 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Crystal structure of the Agrobacterium virulence complex VirE1-VirE2 reveals a flexible protein that can accommodate different partners.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2QDS
 
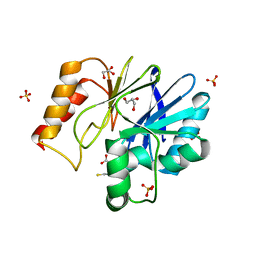 | |
3CA8
 
 | | Crystal structure of Escherichia coli YdcF, an S-adenosyl-L-methionine utilizing enzyme | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, Protein ydcF, SULFATE ION | | 著者 | Lim, K, Chao, K, Lehmann, C, Herzberg, O, Structure 2 Function Project (S2F) | | 登録日 | 2008-02-19 | | 公開日 | 2008-05-06 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | The Escherichia coli YdcF binds S-adenosyl-L-methionine and adopts an alpha/beta-fold characteristic of nucleotide-utilizing enzymes.
Proteins, 72, 2008
|
|
3CCK
 
 | | Human CD69 | | 分子名称: | CHLORIDE ION, Early activation antigen CD69 | | 著者 | Brynda, J, Vanek, O, Rezacova, P. | | 登録日 | 2008-02-26 | | 公開日 | 2008-11-25 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Soluble recombinant CD69 receptors optimized to have an exceptional physical and chemical stability display prolonged circulation and remain intact in the blood of mice
Febs J., 275, 2008
|
|
3C9N
 
 | | Crystal Structure of a SARS Corona Virus Derived Peptide Bound to the Human Major Histocompatibility Complex Class I molecule HLA-B*1501 | | 分子名称: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Beta-2-microglobulin, HLA class I histocompatibility antigen, ... | | 著者 | Roder, G.A, Kristensen, O, Kastrup, J.S, Buus, S, Gajhede, M. | | 登録日 | 2008-02-18 | | 公開日 | 2008-02-26 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (1.87 Å) | | 主引用文献 | Structure of a SARS coronavirus-derived peptide bound to the human major histocompatibility complex class I molecule HLA-B*1501.
ACTA CRYSTALLOGR.,SECT.F, 64, 2008
|
|
3CEJ
 
 | | Human glycogen phosphorylase (tense state) in complex with the allosteric inhibitor AVE2865 | | 分子名称: | 1-{2-[3-(2-Chloro-4,5-difluoro-benzoyl)-ureido]-4-fluoro-phenyl}-piperidine-4-carboxylic acid, Glycogen phosphorylase, liver form, ... | | 著者 | Wendt, K.U, Dreyer, M.K, Anderka, O, Klabunde, T, Loenze, P, Defossa, E, Schmoll, D. | | 登録日 | 2008-02-29 | | 公開日 | 2008-05-27 | | 最終更新日 | 2020-07-29 | | 実験手法 | X-RAY DIFFRACTION (3.3 Å) | | 主引用文献 | Thermodynamic characterization of allosteric glycogen phosphorylase inhibitors.
Biochemistry, 47, 2008
|
|
8Q93
 
 | | Crystal structure of the SARS-COV-2 RBD with neutralizing-VHHs Re30H02 and Re21D01 | | 分子名称: | Nanobody Re21D01, Nanobody Re30H02, Spike protein S1 | | 著者 | Aksu, M, Guttler, T, Rymarenko, O, Gorlich, D. | | 登録日 | 2023-08-19 | | 公開日 | 2023-12-20 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | Nanobodies to multiple spike variants and inhalation of nanobody-containing aerosols neutralize SARS-CoV-2 in cell culture and hamsters.
Antiviral Res., 221, 2023
|
|
8TY6
 
 | |
8Q95
 
 | | Crystal structure of the SARS-CoV-2 BA.1 RBD with neutralizing-VHHs Ma16B06 and Ma3F05 | | 分子名称: | Nanobody Ma16B06, Nanobody Ma3F05, Spike protein S1 | | 著者 | Aksu, M, Rymarenko, O, Guttler, T, Gorlich, D. | | 登録日 | 2023-08-19 | | 公開日 | 2023-12-20 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Nanobodies to multiple spike variants and inhalation of nanobody-containing aerosols neutralize SARS-CoV-2 in cell culture and hamsters.
Antiviral Res., 221, 2023
|
|
6GBL
 | | Repertoires of functionally diverse enzymes through computational design at epistatic active-site positions | | 分子名称: | 1,2-ETHANEDIOL, CACODYLATE ION, FORMIC ACID, ... | | 著者 | Khersonsky, O, Lipsh, R, Avizemer, Z, Goldsmith, M, Ashani, Y, Leader, H, Dym, O, Rogotner, S, Trudeau, D, Tawfik, D.S, Fleishman, S.J. | | 登録日 | 2018-04-15 | | 公開日 | 2018-10-24 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Automated Design of Efficient and Functionally Diverse Enzyme Repertoires.
Mol. Cell, 72, 2018
|
|
3CUO
 | | Crystal structure of the predicted DNA-binding transcriptional regulator from E. coli | | 分子名称: | Uncharacterized HTH-type transcriptional regulator ygaV | | 著者 | Zhang, R, Evdokimova, E, Kagan, O, Savchenko, A, Edwards, A.M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | 登録日 | 2008-04-16 | | 公開日 | 2008-06-17 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | The crystal structure of the predicted DNA-binding transcriptional regulator from E. coli.
To be Published
|
|
3CV6
 
 | | The crystal structure of mouse 17-alpha hydroxysteroid dehydrogenase GG225.226PP mutant in complex with inhibitor and cofactor NADP+. | | 分子名称: | 4-[(1R,2S)-1-ethyl-2-(4-hydroxyphenyl)butyl]phenol, Aldo-keto reductase family 1 member C21, BETA-MERCAPTOETHANOL, ... | | 著者 | Dhagat, U, El-Kabbani, O. | | 登録日 | 2008-04-17 | | 公開日 | 2009-03-03 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Structure of the G225P/G226P mutant of mouse 3(17)alpha-hydroxysteroid dehydrogenase (AKR1C21) ternary complex: implications for the binding of inhibitor and substrate.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3CXW
 
 | | Crystal structure of human proto-oncogene serine threonine kinase (PIM1) in complex with a consensus peptide and a beta carboline ligand I | | 分子名称: | (4R)-7,8-dichloro-1',9-dimethyl-1-oxo-1,2,4,9-tetrahydrospiro[beta-carboline-3,4'-piperidine]-4-carbonitrile, CHLORIDE ION, Pimtide peptide, ... | | 著者 | Filippakopoulos, P, Bullock, A, Fedorov, O, Huber, K, Bracher, F, Pike, A.C.W, Roos, A, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Knapp, S, Structural Genomics Consortium (SGC) | | 登録日 | 2008-04-25 | | 公開日 | 2008-07-15 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | 7,8-Dichloro-1-oxo-beta-carbolines as a Versatile Scaffold for the Development of Potent and Selective Kinase Inhibitors with Unusual Binding Modes
J.Med.Chem., 55, 2012
|
|
3CY4
 | | Crystal Structure cation-dependent mannose 6-phosphate receptor at pH 7.4 | | 分子名称: | Cation-dependent mannose-6-phosphate receptor, GLYCEROL, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | 著者 | Olson, L.J, Hindsgaul, O, Dahms, N.M, Kim, J.-J.P. | | 登録日 | 2008-04-25 | | 公開日 | 2008-05-13 | | 最終更新日 | 2021-10-20 | | 実験手法 | X-RAY DIFFRACTION (1.51 Å) | | 主引用文献 | Structural Insights into the Mechanism of pH-dependent Ligand Binding and Release by the Cation-dependent Mannose 6-Phosphate Receptor.
J.Biol.Chem., 283, 2008
|
|
3D08
 
 | | Human p53 core domain with hot spot mutation R249S and second-site suppressor mutation H168R | | 分子名称: | Cellular tumor antigen p53, ZINC ION | | 著者 | Suad, O, Rozenberg, H, Shimon, L.J.W, Frolow, F, Shakked, Z. | | 登録日 | 2008-05-01 | | 公開日 | 2009-01-20 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Structural basis of restoring sequence-specific DNA binding and transactivation to mutant p53 by suppressor mutations
J.Mol.Biol., 385, 2009
|
|
3CKD
 
 | | Crystal structure of the C-terminal domain of the Shigella type III effector IpaH | | 分子名称: | DI(HYDROXYETHYL)ETHER, GLYCEROL, Invasion plasmid antigen, ... | | 著者 | Lam, R, Singer, A.U, Cuff, M.E, Skarina, T, Kagan, O, DiLeo, R, Edwards, A.M, Joachimiak, A, Savchenko, A, Midwest Center for Structural Genomics (MCSG) | | 登録日 | 2008-03-14 | | 公開日 | 2008-03-25 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (2.65 Å) | | 主引用文献 | Structure of the Shigella T3SS effector IpaH defines a new class of E3 ubiquitin ligases.
Nat.Struct.Mol.Biol., 15, 2008
|
|
6GBK
 | | Repertoires of functionally diverse enzymes through computational design at epistatic active-site positions | | 分子名称: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, FORMIC ACID, Parathion hydrolase, ... | | 著者 | Khersonsky, O, Lipsh, R, Avizemer, Z, Goldsmith, M, Ashani, Y, Leader, H, Dym, O, Rogotner, S, Trudeau, D, Tawfik, D.S, Fleishman, S.J. | | 登録日 | 2018-04-15 | | 公開日 | 2018-10-24 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Automated Design of Efficient and Functionally Diverse Enzyme Repertoires.
Mol. Cell, 72, 2018
|
|
3CIF
 
 | | Crystal Structure of C153S mutant glyceraldehyde 3-phosphate dehydrogenase from Cryptosporidium parvum | | 分子名称: | GLYCERALDEHYDE-3-PHOSPHATE, GLYCEROL, Glyceraldehyde-3-phosphate dehydrogenase, ... | | 著者 | Cook, W.J, Senkovich, O, Chattopadhyay, D. | | 登録日 | 2008-03-11 | | 公開日 | 2009-03-24 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | An unexpected phosphate binding site in Glyceraldehyde 3-Phosphate Dehydrogenase: Crystal structures of apo, holo and ternary complex of Cryptosporidium parvum enzyme
BMC STRUCT.BIOL., 9, 2009
|
|
3CPH
 
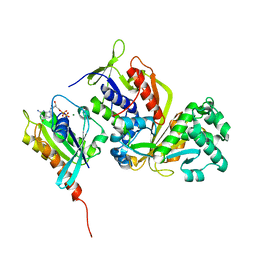 | | Crystal structure of Sec4 in complex with Rab-GDI | | 分子名称: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Rab GDP-dissociation inhibitor, ... | | 著者 | Kravchenko, S, Ignatev, A, Goody, R.S, Rak, A, Pylypenko, O. | | 登録日 | 2008-03-31 | | 公開日 | 2008-05-06 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.9 Å) | | 主引用文献 | A structural model of the GDP dissociation inhibitor rab membrane extraction mechanism.
J.Biol.Chem., 283, 2008
|
|
6GBJ
 
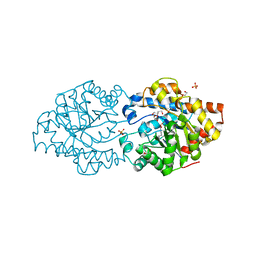 | | Repertoires of functionally diverse enzymes through computational design at epistatic active-site positions | | 分子名称: | 1,2-ETHANEDIOL, FORMIC ACID, Parathion hydrolase, ... | | 著者 | Khersonsky, O, Lipsh, R, Avizemer, Z, Goldsmith, M, Ashani, Y, Leader, H, Dym, O, Rogotner, S, Trudeau, D, Tawfik, D.S, Fleishman, S.J. | | 登録日 | 2018-04-15 | | 公開日 | 2018-10-24 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.63 Å) | | 主引用文献 | Automated Design of Efficient and Functionally Diverse Enzyme Repertoires.
Mol. Cell, 72, 2018
|
|
3D06
 
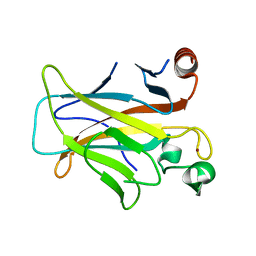 | | Human p53 core domain with hot spot mutation R249S (I) | | 分子名称: | Cellular tumor antigen p53, ZINC ION | | 著者 | Rozenberg, H, Suad, O, Shimon, L.J.W, Frolow, F, Shakked, Z. | | 登録日 | 2008-05-01 | | 公開日 | 2009-01-20 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.2 Å) | | 主引用文献 | Structural basis of restoring sequence-specific DNA binding and transactivation to mutant p53 by suppressor mutations
J.Mol.Biol., 385, 2009
|
|
3D1F
 
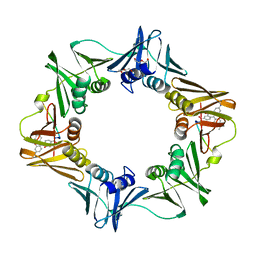 | | Crystal structure of E. coli sliding clamp (beta) bound to a polymerase III peptide | | 分子名称: | 2-[3,6-bis(dimethylamino)xanthen-9-yl]-5-methanoyl-benzoate, DI(HYDROXYETHYL)ETHER, DNA polymerase III subunit beta, ... | | 著者 | Georgescu, R.E, Yurieva, O, Seung-Sup, K, Kuriyan, J, Kong, X.-P, O'Donnell, M. | | 登録日 | 2008-05-05 | | 公開日 | 2008-07-29 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structure of a small-molecule inhibitor of a DNA polymerase sliding clamp.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3D3M
 
 | |
