3UXM
 
 | | Structure Guided Development of Novel Thymidine Mimetics targeting Pseudomonas aeruginosa Thymidylate Kinase: from Hit to Lead Generation | | Descriptor: | 5'-deoxy-5'-fluorothymidine, MAGNESIUM ION, Thymidylate kinase | | Authors: | Choi, J.Y, Plummer, M.S, Starr, J, Desbonnet, C.R, Soutter, H.H, Chang, J, Miller, J.R, Dillman, K, Miller, A.A, Roush, W.R. | | Deposit date: | 2011-12-05 | | Release date: | 2012-02-01 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure guided development of novel thymidine mimetics targeting Pseudomonas aeruginosa thymidylate kinase: from hit to lead generation.
J.Med.Chem., 55, 2012
|
|
1YT7
 
 | | Cathepsin K complexed with a constrained ketoamide inhibitor | | Descriptor: | (1R)-2,2-DIMETHYL-1-({5-[4-(TRIFLUOROMETHYL)PHENYL]-1,3,4-OXADIAZOL-2-YL}METHYL)PROPYL (1S)-1-{OXO[(2-OXO-1,3-OXAZOLIDIN-3-YL)AMINO]ACETYL}PENTYLCARBAMATE, Cathepsin K, SULFATE ION | | Authors: | Barrett, D.G, Boncek, V.M, Catalano, J.G, Deaton, D.N, Hassell, A.M, Jurgensen, C.H, Long, S.T, McFadyen, R.B, Miller, A.B, Miller, L.R, Payne, J.A, Ray, J.A, Samano, V, Shewchuk, L.M, Tavares, F.X, Wells-Knecht, K.J, Willard, D.H, Wright, L.L, Zhou, H.Q. | | Deposit date: | 2005-02-10 | | Release date: | 2005-07-19 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | P(2)-P(3) conformationally constrained ketoamide-based inhibitors of cathepsin K.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
8BAW
 
 | | X-ray structure of the CeuE Homologue from Geobacillus stearothermophilus - 5-LICAM siderophore analogue complex. | | Descriptor: | FE (III) ION, N,N'-pentane-1,5-diylbis(2,3-dihydroxybenzamide), Siderophore ABC transporter substrate-binding protein | | Authors: | Blagova, E.V, Miller, A, Booth, R, Dodson, E.J, Duhme-Klair, A.K, Wilson, K.S. | | Deposit date: | 2022-10-12 | | Release date: | 2023-07-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.471 Å) | | Cite: | Thermostable homologues of the periplasmic siderophore-binding protein CeuE from Geobacillus stearothermophilus and Parageobacillus thermoglucosidasius.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8BF6
 
 | | X-ray structure of the CeuE Homologue from Parageobacillus thermoglucosidasius - azotochelin complex | | Descriptor: | ABC transporter, Azotochelin, FE (III) ION, ... | | Authors: | Wilson, K.S, Duhme-Klair, A.-K, Blagova, E.V, Miller, A, Booth, R, Dodson, E.J. | | Deposit date: | 2022-10-24 | | Release date: | 2023-07-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.969 Å) | | Cite: | Thermostable homologues of the periplasmic siderophore-binding protein CeuE from Geobacillus stearothermophilus and Parageobacillus thermoglucosidasius.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
1YK8
 
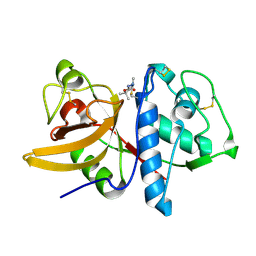 | | Cathepsin K complexed with a cyanamide-based inhibitor | | Descriptor: | Cathepsin K, TERT-BUTYL 2-CYANO-2-METHYLHYDRAZINECARBOXYLATE | | Authors: | Barrett, D.G, Deaton, D.N, Hassell, A.M, McFadyen, R.B, Miller, A.B, Miller, L.R, Payne, J.A, Shewchuk, L.M, Willard, D.H, Wright, L.L. | | Deposit date: | 2005-01-17 | | Release date: | 2005-07-19 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Acyclic cyanamide-based inhibitors of cathepsin K.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
2AUX
 
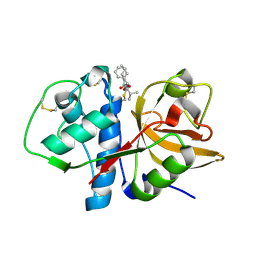 | | Cathepsin K complexed with a semicarbazone inhibitor | | Descriptor: | (1R)-2-METHYL-1-(PHENYLMETHYL)PROPYL[(1S)-1-FORMYLPENTYL]CARBAMATE, Cathepsin K | | Authors: | Adkison, K.K, Barrett, D.G, Deaton, D.N, Gampe, R.T, Hassell, A.M, Long, S.T, McFadyen, R.B, Miller, A.B, Miller, L.R, Shewchuk, L.M. | | Deposit date: | 2005-08-29 | | Release date: | 2006-08-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Semicarbazone-based inhibitors of cathepsin K, are they prodrugs for aldehyde inhibitors?
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2AUZ
 
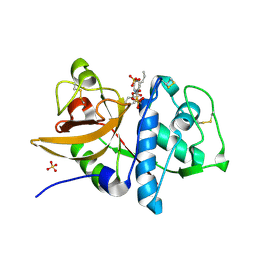 | | Cathepsin K complexed with a semicarbazone inhibitor | | Descriptor: | 1-(PHENYLMETHYL)CYCLOPENTYL[(1S)-1-FORMYLPENTYL]CARBAMATE, Cathepsin K, SULFATE ION | | Authors: | Adkison, K.K, Barrett, D.G, Deaton, D.N, Gampe, R.T, Hassell, A.M, Long, S.T, McFadyen, R.B, Miller, A.B, Miller, L.R, Shewchuk, L.M. | | Deposit date: | 2005-08-29 | | Release date: | 2006-08-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Semicarbazone-based inhibitors of cathepsin K, are they prodrugs for aldehyde inhibitors?
Bioorg.Med.Chem.Lett., 16, 2006
|
|
8BAX
 
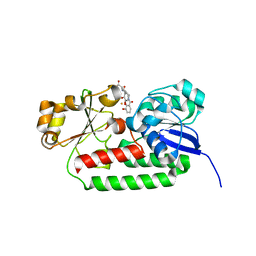 | | X-ray structure of the CeuE Homologue from Geobacillus stearothermophilus - azotochelin complex. | | Descriptor: | Azotochelin, FE (III) ION, Siderophore ABC transporter substrate-binding protein | | Authors: | Blagova, E.V, Miller, A, Dodson, E.J, Booth, R, Duhme-Klair, A.K, Wilson, K.S. | | Deposit date: | 2022-10-12 | | Release date: | 2023-07-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Thermostable homologues of the periplasmic siderophore-binding protein CeuE from Geobacillus stearothermophilus and Parageobacillus thermoglucosidasius.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
1YK7
 
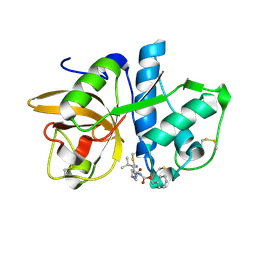 | | Cathepsin K complexed with a cyanopyrrolidine inhibitor | | Descriptor: | Cathepsin K, N2-[(BENZYLOXY)CARBONYL]-N1-[(3S)-1-CYANOPYRROLIDIN-3-YL]-L-LEUCINAMIDE | | Authors: | Barrett, D.G, Deaton, D.N, Hassell, A.M, McFadyen, R.B, Miller, A.B, Miller, L.R, Shewchuk, L.M, Tavares, F.X, Willard, D.H, Wright, L.L. | | Deposit date: | 2005-01-17 | | Release date: | 2005-03-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Novel and potent cyclic cyanamide-based cathepsin K inhibitors.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
5KW2
 
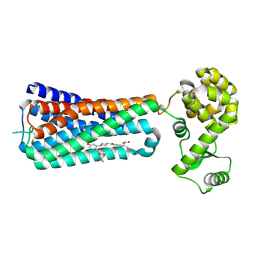 | | The extra-helical binding site of GPR40 and the structural basis for allosteric agonism and incretin stimulation | | Descriptor: | (3~{S})-3-cyclopropyl-3-[2-[1-[2-[2,2-dimethylpropyl-(6-methylpyridin-2-yl)carbamoyl]-5-methoxy-phenyl]piperidin-4-yl]-1-benzofuran-6-yl]propanoic acid, Free fatty acid receptor 1,Lysozyme,Free fatty acid receptor 1 | | Authors: | Ho, J.D, Chau, B, Rodgers, L, Lu, F, Wilbur, K.L, Otto, K.A, Chen, Y, Song, M, Riley, J.P, Yang, H.-C, Reynolds, N.A, Kahl, S.D, Lewis, A.P, Groshong, C, Madsen, R.E, Conners, K, Linswala, J.P, Gheyi, T, Saflor, M.D, Lee, M.R, Benach, J, Baker, K.A, Montrose-Rafizadeh, C, Genin, M.J, Miller, A.R, Hamdouchi, C. | | Deposit date: | 2016-07-15 | | Release date: | 2018-05-02 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Structural basis for GPR40 allosteric agonism and incretin stimulation.
Nat Commun, 9, 2018
|
|
7QHZ
 
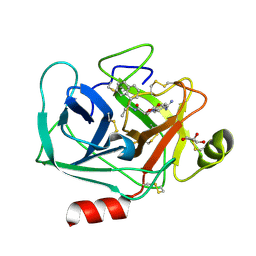 | | Crystal structure of KLK6 in complex with compound DKFZ917 | | Descriptor: | (5~{R})-3-(4-carbamimidoylphenyl)-~{N}-[(1~{S})-1-naphthalen-1-ylpropyl]-2-oxidanylidene-1,3-oxazolidine-5-carboxamide, GLYCEROL, Kallikrein-6 | | Authors: | Jagtap, P.K.A, Baumann, A, Lohbeck, J, Isak, D, Miller, A, Hennig, J. | | Deposit date: | 2021-12-14 | | Release date: | 2022-11-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Scalable synthesis and structural characterization of reversible KLK6 inhibitors.
Rsc Adv, 12, 2022
|
|
5J8D
 
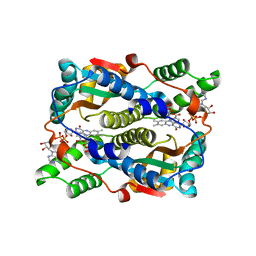 | | Structure of nitroreductase from E. cloacae complexed with nicotinic acid adenine dinucleotide | | Descriptor: | FLAVIN MONONUCLEOTIDE, NICOTINIC ACID ADENINE DINUCLEOTIDE, Oxygen-insensitive NAD(P)H nitroreductase | | Authors: | Haynes, C.A, Koder, R.L, Miller, A.F, Rodgers, D.W. | | Deposit date: | 2016-04-07 | | Release date: | 2017-05-17 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Mechanism-Informed Refinement Reveals Altered Substrate-Binding Mode for Catalytically Competent Nitroreductase.
Structure, 25, 2017
|
|
7QI0
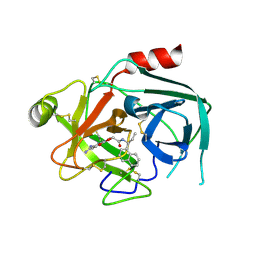 
 | | Crystal structure of KLK6 in complex with compound DKFZ918 | | Descriptor: | (5~{R})-3-(6-carbamimidoylpyridin-3-yl)-~{N}-[(1~{S})-1-naphthalen-1-ylpropyl]-2-oxidanylidene-1,3-oxazolidine-5-carboxamide, Kallikrein-6 | | Authors: | Jagtap, P.K.A, Baumann, A, Lohbeck, J, Isak, D, Miller, A, Hennig, J. | | Deposit date: | 2021-12-14 | | Release date: | 2022-11-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Scalable synthesis and structural characterization of reversible KLK6 inhibitors.
Rsc Adv, 12, 2022
|
|
6H47
 
 | | Human KRAS in complex with darpin K19 | | Descriptor: | GTPase KRas, SULFATE ION, darpin K19 | | Authors: | Debreczeni, J.E, Bery, N, Legg, S, Breed, J, Embrey, K, Stubbs, C, Kolasinska-Zwierz, P, Barrett, N, Marwood, R, Watson, J, Tart, J, Overman, R, Miller, A, Phillips, C, Minter, R, Rabbitts, T.H. | | Deposit date: | 2018-07-20 | | Release date: | 2019-04-24 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | KRAS-specific inhibition using a DARPin binding to a site in the allosteric lobe.
Nat Commun, 10, 2019
|
|
6H46
 
 | | Human KRAS in complex with darpin K13 | | Descriptor: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, SULFATE ION, ... | | Authors: | Debreczeni, J.E, Bery, N, Legg, S, Breed, J, Embrey, K, Stubbs, C, Kolasinska-Zwierz, P, Barrett, N, Marwood, R, Watson, J, Tart, J, Overman, R, Miller, A, Phillips, C, Minter, R, Rabbitts, T.H. | | Deposit date: | 2018-07-20 | | Release date: | 2019-04-24 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | KRAS-specific inhibition using a DARPin binding to a site in the allosteric lobe.
Nat Commun, 10, 2019
|
|
1FM9
 
 | | THE 2.1 ANGSTROM RESOLUTION CRYSTAL STRUCTURE OF THE HETERODIMER OF THE HUMAN RXRALPHA AND PPARGAMMA LIGAND BINDING DOMAINS RESPECTIVELY BOUND WITH 9-CIS RETINOIC ACID AND GI262570 AND CO-ACTIVATOR PEPTIDES. | | Descriptor: | (9cis)-retinoic acid, 2-(2-BENZOYL-PHENYLAMINO)-3-{4-[2-(5-METHYL-2-PHENYL-OXAZOL-4-YL)-ETHOXY]-PHENYL}-PROPIONIC ACID, PEROXISOME PROLIFERATOR ACTIVATED RECEPTOR GAMMA, ... | | Authors: | Gampe Jr, R.T, Montana, V.G, Lambert, M.H, Miller, A.B, Bledsoe, R.K, Milburn, M.V, Kliewer, S.A, Willson, T.M, Xu, H.E. | | Deposit date: | 2000-08-16 | | Release date: | 2001-02-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Asymmetry in the PPARgamma/RXRalpha crystal structure reveals the molecular basis of heterodimerization among nuclear receptors.
Mol.Cell, 5, 2000
|
|
1FM6
 
 | | THE 2.1 ANGSTROM RESOLUTION CRYSTAL STRUCTURE OF THE HETERODIMER OF THE HUMAN RXRALPHA AND PPARGAMMA LIGAND BINDING DOMAINS RESPECTIVELY BOUND WITH 9-CIS RETINOIC ACID AND ROSIGLITAZONE AND CO-ACTIVATOR PEPTIDES. | | Descriptor: | (9cis)-retinoic acid, 2,4-THIAZOLIDIINEDIONE, 5-[[4-[2-(METHYL-2-PYRIDINYLAMINO)ETHOXY]PHENYL]METHYL]-(9CL), ... | | Authors: | Gampe Jr, R.T, Montana, V.G, Lambert, M.H, Miller, A.B, Bledsoe, R.K, Milburn, M.V, Kliewer, S.A, Willson, T.M, Xu, H.E. | | Deposit date: | 2000-08-16 | | Release date: | 2001-02-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Asymmetry in the PPARgamma/RXRalpha crystal structure reveals the molecular basis of heterodimerization among nuclear receptors.
Mol.Cell, 5, 2000
|
|
8SNZ
 
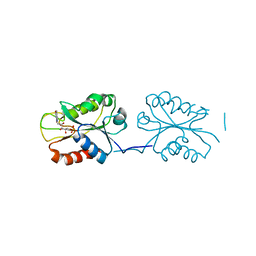 | | X-ray Crystal Structure of FMN-bound long-chain flavodoxin from Rhodopseudomonas palustris | | Descriptor: | FLAVIN MONONUCLEOTIDE, Flavodoxin | | Authors: | Ansari, A, Khan, S.A, Miller, A.F. | | Deposit date: | 2023-04-28 | | Release date: | 2024-03-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Structure, dynamics, and redox reactivity of an all-purpose flavodoxin.
J.Biol.Chem., 300, 2024
|
|
8FE4
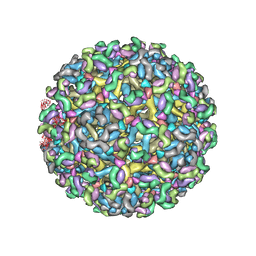 
 | | Structure of dengue virus (DENV2) in complex with prM13, an anti-PrM monoclonal antibody | | Descriptor: | Envelope protein E, prM protein, prM13 Fab Heavy Chain, ... | | Authors: | Dowd, A.D, Sirohi, D, Speer, S, Mukherjee, S, Govero, J, Aleshnick, M, Larman, B, Sukupolvi-Petty, S, Sevvana, M, Miller, A.S, Klose, T, Zheng, A, Kielian, M, Kuhn, R.J, Diamond, M.S, Pierson, T.C. | | Deposit date: | 2022-12-05 | | Release date: | 2023-02-01 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (9.8 Å) | | Cite: | prM-reactive antibodies reveal a role for partially mature virions in dengue virus pathogenesis.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8FE3
 
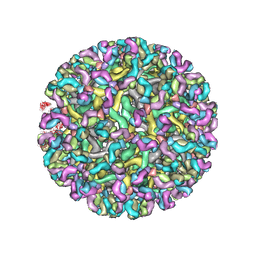 | | Structure of dengue virus (DENV2) in complex with prM12, an anti-PrM monoclonal antibody | | Descriptor: | Envelope protein E, prM protein, prM12 Fab Heavy Chain, ... | | Authors: | Dowd, A.D, Sirohi, D, Speer, S, Mukherjee, S, Govero, J, Aleshnick, M, Larman, B, Sukupolvi-Petty, S, Sevvana, M, Miller, A.S, Klose, T, Zheng, A, Kielian, M, Kuhn, R.J, Diamond, M.S, Pierson, T.C. | | Deposit date: | 2022-12-05 | | Release date: | 2023-02-01 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (10.2 Å) | | Cite: | prM-reactive antibodies reveal a role for partially mature virions in dengue virus pathogenesis.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
5CLS
 
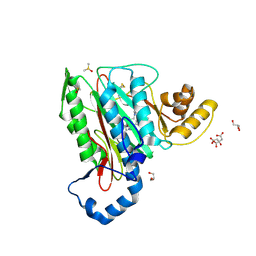 | | Structure of human methionine aminopeptidase-2 complexed with spiroepoxytriazole inhibitor (+)-31a | | Descriptor: | (4R,7R)-7-hydroxy-1-(4-methoxybenzyl)-7-methyl-4,5,6,7-tetrahydro-1H-benzotriazol-4-yl propan-2-ylcarbamate, CHLORIDE ION, CITRIC ACID, ... | | Authors: | Janowski, R, Miller, A.K, Niessing, D. | | Deposit date: | 2015-07-16 | | Release date: | 2016-01-13 | | Last modified: | 2019-11-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Spiroepoxytriazoles Are Fumagillin-like Irreversible Inhibitors of MetAP2 with Potent Cellular Activity.
Acs Chem.Biol., 11, 2016
|
|
5D6F
 
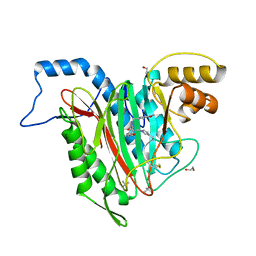 | | Structure of human methionine aminopeptidase-2 complexed with spiroepoxytriazole inhibitor (+)-31b | | Descriptor: | (4S,7R)-7-hydroxy-1-(4-methoxybenzyl)-7-methyl-4,5,6,7-tetrahydro-1H-benzotriazol-4-yl propan-2-ylcarbamate, 1,2-ETHANEDIOL, COBALT (II) ION, ... | | Authors: | Janowski, R, Miller, A.K, Niessing, D. | | Deposit date: | 2015-08-12 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Spiroepoxytriazoles Are Fumagillin-like Irreversible Inhibitors of MetAP2 with Potent Cellular Activity.
Acs Chem.Biol., 11, 2016
|
|
5D6E
 
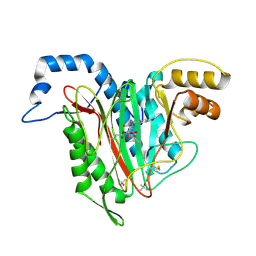 | | Structure of human methionine aminopeptidase 2 with covalent spiroepoxytriazole inhibitor (-)-31b | | Descriptor: | (4R,7S)-7-hydroxy-1-(4-methoxybenzyl)-7-methyl-4,5,6,7-tetrahydro-1H-benzotriazol-4-yl propan-2-ylcarbamate, COBALT (II) ION, Methionine aminopeptidase 2, ... | | Authors: | Janowski, R, Miller, A.K, Niessing, D. | | Deposit date: | 2015-08-12 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Spiroepoxytriazoles Are Fumagillin-like Irreversible Inhibitors of MetAP2 with Potent Cellular Activity.
Acs Chem.Biol., 11, 2016
|
|
2BKB
 
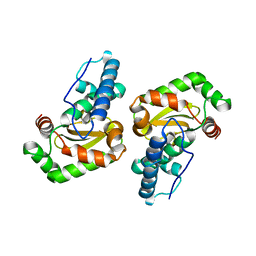 | | q69e-FeSOD | | Descriptor: | FE (II) ION, SUPEROXIDE DISMUTASE [FE] | | Authors: | Yikilmaz, E, Rodgers, D.W, Miller, A.-F. | | Deposit date: | 2005-02-14 | | Release date: | 2007-01-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Crucial Importance of Chemistry in the Structure-Function Link: Manipulating Hydrogen Bonding in Iron-Containing Superoxide Dismutase.
Biochemistry, 45, 2006
|
|
5UHY
 
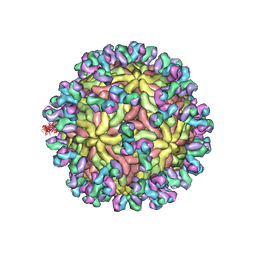 | | A Human Antibody Against Zika Virus Crosslinks the E Protein to Prevent Infection | | Descriptor: | ZV67 Fab chain 1, ZV67 Fab chain 2, envelope protein | | Authors: | Hasan, S.S, Miller, A, Sapparapu, G, Fernandez, E, Klose, T, Long, F, Fokine, A, Porta, J.C, Jiang, W, Diamond, M.S, Crowe Jr, J.E, Kuhn, R.J, Rossmann, M.G. | | Deposit date: | 2017-01-12 | | Release date: | 2017-03-29 | | Last modified: | 2019-11-27 | | Method: | ELECTRON MICROSCOPY (6.2 Å) | | Cite: | A human antibody against Zika virus crosslinks the E protein to prevent infection.
Nat Commun, 8, 2017
|
|
