8FO0
 
 | | The structure of a crystallizable variant of E. coli pyruvate formate-lyase activating enzyme bound to a partially cleaved SAM molecule | | Descriptor: | IRON/SULFUR CLUSTER, POTASSIUM ION, Pyruvate formate-lyase 1-activating enzyme, ... | | Authors: | Moody, J.D, Saxton, A.J, Galambas, A, Lawrence, C.M, Broderick, J.B. | | Deposit date: | 2022-12-29 | | Release date: | 2023-05-24 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Computational engineering of previously crystallized pyruvate formate-lyase activating enzyme reveals insights into SAM binding and reductive cleavage.
J.Biol.Chem., 299, 2023
|
|
8FOL
 
 | | The structure of a crystallizable variant of E. coli pyruvate formate-lyase activating enzyme bound to SAM, alternate crystal form | | Descriptor: | CHLORIDE ION, IRON/SULFUR CLUSTER, POTASSIUM ION, ... | | Authors: | Moody, J.D, Saxton, A.J, Galambas, A, Lawrence, C.M, Broderick, J.B. | | Deposit date: | 2022-12-31 | | Release date: | 2023-05-24 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Computational engineering of previously crystallized pyruvate formate-lyase activating enzyme reveals insights into SAM binding and reductive cleavage.
J.Biol.Chem., 299, 2023
|
|
8FSI
 
 | | The structure of a crystallizable variant of E. coli pyruvate formate-lyase activating enzyme bound to SAM | | Descriptor: | CHLORIDE ION, IRON/SULFUR CLUSTER, POTASSIUM ION, ... | | Authors: | Moody, J.D, Galambas, A, Lawrence, C.M, Broderick, J.B. | | Deposit date: | 2023-01-10 | | Release date: | 2023-05-24 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Computational engineering of previously crystallized pyruvate formate-lyase activating enzyme reveals insights into SAM binding and reductive cleavage.
J.Biol.Chem., 299, 2023
|
|
1BZL
 
 | | CRYSTAL STRUCTURE OF TRYPANOSOMA CRUZI TRYPANOTHIONE REDUCTASE IN COMPLEX WITH TRYPANOTHIONE, AND THE STRUCTURE-BASED DISCOVERY OF NEW NATURAL PRODUCT INHIBITORS | | Descriptor: | BIS(GAMMA-GLUTAMYL-CYSTEINYL-GLYCINYL)SPERMIDINE, FLAVIN-ADENINE DINUCLEOTIDE, TRYPANOTHIONE REDUCTASE (OXIDIZED FORM) | | Authors: | Bond, C.S, Zhang, Y, Berriman, M, Cunningham, M, Fairlamb, A.H, Hunter, W.N. | | Deposit date: | 1998-11-02 | | Release date: | 1999-11-10 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of Trypanosoma cruzi trypanothione reductase in complex with trypanothione, and the structure-based discovery of new natural product inhibitors.
Structure Fold.Des., 7, 1999
|
|
7P3I
 
 | | Crystal structure of human CD40/TNFRSF5 in complex with the anti-CD40 DARPin protein | | Descriptor: | Darpin, SODIUM ION, Tumor necrosis factor receptor superfamily member 5 | | Authors: | Malvezzi, F, Mangold, S, Hospodarsch, T, Reichen, C, Iss, C, Lammens, A, Krapp, S, Domke, C. | | Deposit date: | 2021-07-07 | | Release date: | 2022-04-06 | | Last modified: | 2022-05-11 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | A Multispecific Anti-CD40 DARPin Construct Induces Tumor-Selective CD40 Activation and Tumor Regression.
Cancer Immunol Res, 10, 2022
|
|
8B5U
 
 | |
2QFB
 
 | | Crystal structure of the regulatory domain of human RIG-I with bound Zn | | Descriptor: | Probable ATP-dependent RNA helicase DDX58, ZINC ION | | Authors: | Cui, S, Lammens, A, Lammens, K, Hopfner, K.P. | | Deposit date: | 2007-06-27 | | Release date: | 2008-02-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The C-Terminal Regulatory Domain Is the RNA 5'-Triphosphate Sensor of RIG-I.
Mol.Cell, 29, 2008
|
|
2QFD
 
 | | Crystal structure of the regulatory domain of human RIG-I with bound Hg | | Descriptor: | MERCURY (II) ION, Probable ATP-dependent RNA helicase DDX58 | | Authors: | Cui, S, Lammens, A, Lammens, K, Hopfner, K.P. | | Deposit date: | 2007-06-27 | | Release date: | 2008-02-12 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The C-Terminal Regulatory Domain Is the RNA 5'-Triphosphate Sensor of RIG-I.
Mol.Cell, 29, 2008
|
|
8B5T
 
 | |
8BM3
 
 | | H207A mutant of E. coli PgpB, a PAP2 type phosphatidyl glycerol phosphate and C55-PP phosphatase, in complex with farnesyl pyrophosphate | | Descriptor: | FARNESYL DIPHOSPHATE, Phosphatidylglycerophosphatase B | | Authors: | Delbrassine, F, El Ghachi, M, Lambion, A, Herman, R, Kerff, F. | | Deposit date: | 2022-11-10 | | Release date: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | H207A mutant of E. coli PgpB, a PAP2 type phosphatidyl glycerol phosphate and C55-PP phosphatase, in complex with farnesyl pyrophosphate
To Be Published
|
|
1BI9
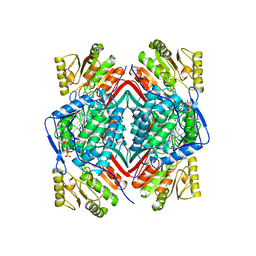 
 | | RETINAL DEHYDROGENASE TYPE TWO WITH NAD BOUND | | Descriptor: | CHLORIDE ION, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, RETINAL DEHYDROGENASE TYPE II | | Authors: | Newcomer, M.E, Lamb, A.L. | | Deposit date: | 1998-06-23 | | Release date: | 1999-07-22 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The structure of retinal dehydrogenase type II at 2.7 A resolution: implications for retinal specificity.
Biochemistry, 38, 1999
|
|
7L5P
 
 | | Crystal structure of the covalently bonded complex of rilzabrutinib with BTK | | Descriptor: | (2E)-2-{(3R)-3-[4-amino-3-(2-fluoro-4-phenoxyphenyl)-1H-pyrazolo[3,4-d]pyrimidin-1-yl]piperidine-1-carbonyl}-4-methyl-4-[4-(oxetan-3-yl)piperazin-1-yl]pent-2-enenitrile, 1,2-ETHANEDIOL, SULFATE ION, ... | | Authors: | Bradshaw, J.M, Brameld, K.A, Mrosek, M, Lammens, A, Blaesse, M. | | Deposit date: | 2020-12-22 | | Release date: | 2022-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | The Discovery Rilzabrutinib (PRN1008): A Reversible Covalent BTK Inhibitor for Immune Mediated Diseases
To Be Published
|
|
6U4L
 
 | | cysteine dioxygenase variant - C93E | | Descriptor: | ACETATE ION, Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Meneely, K.M, Chilton, A.S, Forbes, D.L, Ellis, H.R, Lamb, A.L. | | Deposit date: | 2019-08-26 | | Release date: | 2020-07-08 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.911 Å) | | Cite: | The 3-His Metal Coordination Site Promotes the Coupling of Oxygen Activation to Cysteine Oxidation in Cysteine Dioxygenase.
Biochemistry, 59, 2020
|
|
6U4V
 
 | | Non-crosslinked wild type cysteine dioxygenase | | Descriptor: | Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Meneely, K.M, Chilton, A.S, Forbes, D.L, Ellis, H.R, Lamb, A.L. | | Deposit date: | 2019-08-26 | | Release date: | 2020-07-08 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The 3-His Metal Coordination Site Promotes the Coupling of Oxygen Activation to Cysteine Oxidation in Cysteine Dioxygenase.
Biochemistry, 59, 2020
|
|
2W4R
 
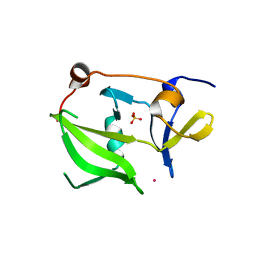 | | Crystal structure of the regulatory domain of human LGP2 | | Descriptor: | MERCURY (II) ION, PROBABLE ATP-DEPENDENT RNA HELICASE DHX58, SULFATE ION | | Authors: | Pippig, D.A, Hellmuth, J.C, Cui, S, Kirchhofer, A, Lammens, K, Lammens, A, Schmidt, A, Rothenfusser, S, Hopfner, K.P. | | Deposit date: | 2008-12-01 | | Release date: | 2009-02-24 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Regulatory Domain of the Rig-I Family ATPase Lgp2 Senses Double-Stranded RNA.
Nucleic Acids Res., 37, 2009
|
|
8AU5
 
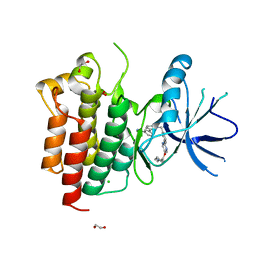 | | c-MET F1200I mutant in complex with Tepotinib | | Descriptor: | 1,2-ETHANEDIOL, 3-[1-(3-{5-[(1-methylpiperidin-4-yl)methoxy]pyrimidin-2-yl}benzyl)-6-oxo-1,6-dihydropyridazin-3-yl]benzonitrile, CHLORIDE ION, ... | | Authors: | Graedler, U, Lammens, A. | | Deposit date: | 2022-08-25 | | Release date: | 2023-09-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Biophysical and structural characterization of the impacts of MET phosphorylation on tepotinib binding.
J.Biol.Chem., 299, 2023
|
|
1DKW
 
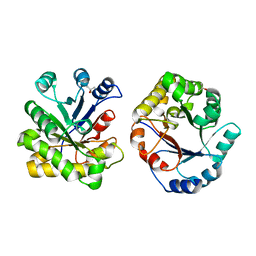 | | CRYSTAL STRUCTURE OF TRIOSE-PHOSPHATE ISOMERASE WITH MODIFIED SUBSTRATE BINDING SITE | | Descriptor: | TERTIARY-BUTYL ALCOHOL, TRIOSEPHOSPHATE ISOMERASE | | Authors: | Norledge, B.V, Lambeir, A.M, Abagyan, R.A, Rottman, A, Fernandez, A.M, Filimonov, V.V, Peter, M.G, Wierenga, R.K. | | Deposit date: | 1999-12-08 | | Release date: | 2000-11-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Modeling, mutagenesis, and structural studies on the fully conserved phosphate-binding loop (loop 8) of triosephosphate isomerase: toward a new substrate specificity.
Proteins, 42, 2001
|
|
8AU3
 
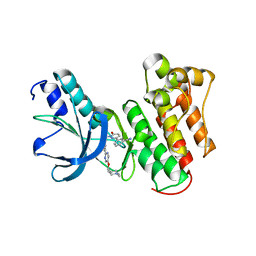 | | c-MET Y1234E,Y1235E mutant in complex with Tepotinib | | Descriptor: | 3-[1-(3-{5-[(1-methylpiperidin-4-yl)methoxy]pyrimidin-2-yl}benzyl)-6-oxo-1,6-dihydropyridazin-3-yl]benzonitrile, DI(HYDROXYETHYL)ETHER, Hepatocyte growth factor receptor, ... | | Authors: | Graedler, U, Lammens, A. | | Deposit date: | 2022-08-25 | | Release date: | 2023-09-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Biophysical and structural characterization of the impacts of MET phosphorylation on tepotinib binding.
J.Biol.Chem., 299, 2023
|
|
6PBT
 
 | | Pseudopaline Dehydrogenase with (R)-Pseudopaline Soaked 2 hours | | Descriptor: | 1,2-ETHANEDIOL, N-[(3S)-3-amino-3-carboxypropyl]-L-histidine, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | McFarlane, J.S, Lamb, A.L. | | Deposit date: | 2019-06-14 | | Release date: | 2019-10-30 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Staphylopine and pseudopaline dehydrogenase from bacterial pathogens catalyze reversible reactions and produce stereospecific metallophores.
J.Biol.Chem., 294, 2019
|
|
7UEZ
 
 | | Apo RibB from Vibrio cholera | | Descriptor: | 3,4-dihydroxy-2-butanone 4-phosphate synthase | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF5
 
 | | RibB from Vibrio cholera bound with intermediate 2 in the reaction cycle and the products DHBP and formate | | Descriptor: | (2R)-2-(dihydroxymethyl)-2-hydroxy-3-oxobutyl dihydrogen phosphate, (2R)-2-hydroxy-3-oxobutyl dihydrogen phosphate, 3,4-dihydroxy-2-butanone 4-phosphate synthase, ... | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF0
 
 | | RibB from Vibrio cholera bound with D-ribulose-5-phosphate (D-Ru5P) | | Descriptor: | 3,4-dihydroxy-2-butanone 4-phosphate synthase, RIBULOSE-5-PHOSPHATE | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF1
 
 | | RibB from Vibrio cholera bound with D-Ribose-5-phosphate (D-R5P) and manganese | | Descriptor: | 3,4-dihydroxy-2-butanone 4-phosphate synthase, MANGANESE (II) ION, RIBOSE-5-PHOSPHATE | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF4
 
 | | RibB from Vibrio cholera bound with intermediate 1 of the reaction cycle and D-ribulose-5-phosphate (D-Ru5P) | | Descriptor: | (2R)-2-hydroxy-3,4-dioxopentyl dihydrogen phosphate, 1,2-ETHANEDIOL, 3,4-dihydroxy-2-butanone 4-phosphate synthase, ... | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF2
 
 | | RibB from Vibrio cholera bound with D-xylulose-5-phosphate (D-Xy5P) and manganese | | Descriptor: | 1,2-ETHANEDIOL, 3,4-dihydroxy-2-butanone 4-phosphate synthase, 5-O-phosphono-D-xylulose, ... | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
