8OMG
 
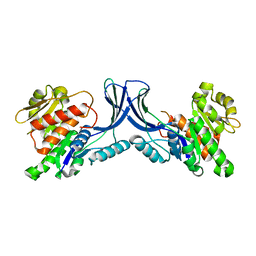 | | Crystal structure of mKHK (apo) | | Descriptor: | Ketohexokinase | | Authors: | Ebenhoch, E, Pautsch, P. | | Deposit date: | 2023-03-31 | | Release date: | 2023-09-27 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Crystal structures of human and mouse ketohexokinase provide a structural basis for species- and isoform-selective inhibitor design.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
8OMD
 
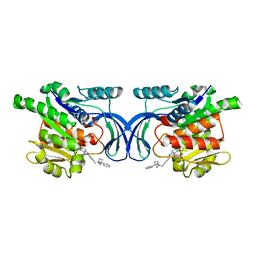 | | Crystal structure of mKHK in complex with compound-4 | | Descriptor: | Ketohexokinase, compound | | Authors: | Ebenhoch, R, Pautsch, A. | | Deposit date: | 2023-03-31 | | Release date: | 2023-09-27 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of human and mouse ketohexokinase provide a structural basis for species- and isoform-selective inhibitor design.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
7VSI
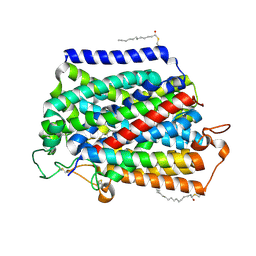 
 | | Structure of human SGLT2-MAP17 complex bound with empagliflozin | | Descriptor: | (2S,3R,4R,5S,6R)-2-[4-chloranyl-3-[[4-[(3S)-oxolan-3-yl]oxyphenyl]methyl]phenyl]-6-(hydroxymethyl)oxane-3,4,5-triol, PALMITIC ACID, PDZK1-interacting protein 1, ... | | Authors: | Chen, L, Niu, Y, Liu, R. | | Deposit date: | 2021-10-26 | | Release date: | 2021-12-15 | | Last modified: | 2022-02-16 | | Method: | ELECTRON MICROSCOPY (2.95 Å) | | Cite: | Structural basis of inhibition of the human SGLT2-MAP17 glucose transporter.
Nature, 601, 2022
|
|
6TX7
 
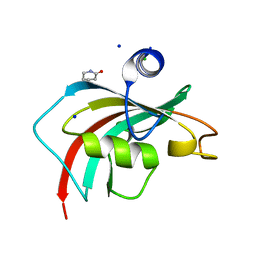 | |
6TX8
 
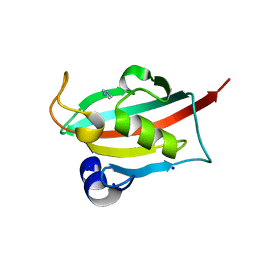 | |
6TX5
 
 | |
6TX6
 | |
6TXX
 | | CRYSTAL STRUCTURE OF HUMAN FKBP51 FK1 DOMAIN A19T MUTANT IN COMPLEX WITH SAFit2 | | Descriptor: | (1R)-3-(3,4-dimethoxyphenyl)-1-{3-[2-(morpholin-4-yl)ethoxy]phenyl}propyl (2S)-1-[(2S)-2-[(1S)-cyclohex-2-en-1-yl]-2-(3,4,5-trimethoxyphenyl)acetyl]piperidine-2-carboxylate, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Fiegen, D, Draxler, S.W. | | Deposit date: | 2020-01-14 | | Release date: | 2020-05-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Hybrid Screening Approach for Very Small Fragments: X-ray and Computational Screening on FKBP51.
J.Med.Chem., 63, 2020
|
|
1IAE
 
 | | CRYSTAL STRUCTURES, SPECTROSCOPIC FEATURES, AND CATALYTIC PROPERTIES OF COBALT(II), COPPER(II), NICKEL(II), AND MERCURY(II) DERIVATIVES OF THE ZINC ENDOPEPTIDASE ASTACIN. A CORRELATION OF STRUCTURE AND PROTEOLYTIC ACTIVITY | | Descriptor: | ASTACIN, NICKEL (II) ION | | Authors: | Grams, F, Stoecker, W, Bode, W. | | Deposit date: | 1994-05-09 | | Release date: | 1994-08-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Crystal structures, spectroscopic features, and catalytic properties of cobalt(II), copper(II), nickel(II), and mercury(II) derivatives of the zinc endopeptidase astacin. A correlation of structure and proteolytic activity.
J.Biol.Chem., 269, 1994
|
|
1IAB
 
 | | CRYSTAL STRUCTURES, SPECTROSCOPIC FEATURES, AND CATALYTIC PROPERTIES OF COBALT(II), COPPER(II), NICKEL(II), AND MERCURY(II) DERIVATIVES OF THE ZINC ENDOPEPTIDASE ASTACIN. A CORRELATION OF STRUCTURE AND PROTEOLYTIC ACTIVITY | | Descriptor: | ASTACIN, COBALT (II) ION | | Authors: | Gomis-Rueth, F.-X, Stoecker, W, Bode, W. | | Deposit date: | 1994-05-09 | | Release date: | 1994-08-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Crystal structures, spectroscopic features, and catalytic properties of cobalt(II), copper(II), nickel(II), and mercury(II) derivatives of the zinc endopeptidase astacin. A correlation of structure and proteolytic activity.
J.Biol.Chem., 269, 1994
|
|
1IAA
 
 | | CRYSTAL STRUCTURES, SPECTROSCOPIC FEATURES, AND CATALYTIC PROPERTIES OF COBALT(II), COPPER(II), NICKEL(II), AND MERCURY(II) DERIVATIVES OF THE ZINC ENDOPEPTIDASE ASTACIN. A CORRELATION OF STRUCTURE AND PROTEOLYTIC ACTIVITY | | Descriptor: | ASTACIN, COPPER (II) ION | | Authors: | Gomis-Rueth, F.-X, Stoecker, W, Bode, W. | | Deposit date: | 1994-05-09 | | Release date: | 1994-08-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structures, spectroscopic features, and catalytic properties of cobalt(II), copper(II), nickel(II), and mercury(II) derivatives of the zinc endopeptidase astacin. A correlation of structure and proteolytic activity.
J.Biol.Chem., 269, 1994
|
|
6TX4
 | |
6TX9
 
 | |
6GPS
 
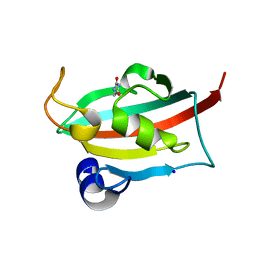
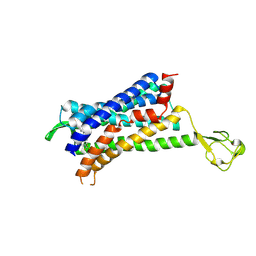 | | CRYSTAL STRUCTURE OF CCR2A IN COMPLEX WITH MK-0812 | | Descriptor: | C-C chemokine receptor type 2,Rubredoxin,C-C chemokine receptor type 2, ZINC ION, [(3~{S},4~{S})-3-methoxyoxan-4-yl]-[(1~{R},3~{S})-3-propan-2-yl-3-[[3-(trifluoromethyl)-7,8-dihydro-5~{H}-1,6-naphthyridin-6-yl]carbonyl]cyclopentyl]azanium | | Authors: | Pautsch, A, Schnapp, G. | | Deposit date: | 2018-06-07 | | Release date: | 2019-01-02 | | Last modified: | 2019-03-13 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Crystal Structure of CC Chemokine Receptor 2A in Complex with an Orthosteric Antagonist Provides Insights for the Design of Selective Antagonists.
Structure, 27, 2019
|
|
6GPX
 
 | | CRYSTAL STRUCTURE OF CCR2A IN COMPLEX WITH MK-0812 | | Descriptor: | C-C chemokine receptor type 2,Rubredoxin,C-C chemokine receptor type 2, OLEIC ACID, ZINC ION, ... | | Authors: | Pautsch, A, Schnapp, G, Cheng, R, Apel, A. | | Deposit date: | 2018-06-07 | | Release date: | 2019-01-02 | | Last modified: | 2019-03-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of CC Chemokine Receptor 2A in Complex with an Orthosteric Antagonist Provides Insights for the Design of Selective Antagonists.
Structure, 27, 2019
|
|
5EU1
 | | CRYSTAL STRUCTURE OF BRD9 IN COMPLEX WITH BI-7273 | | Descriptor: | 4-[4-[(dimethylamino)methyl]-3,5-dimethoxy-phenyl]-2-methyl-2,7-naphthyridin-1-one, BRD9 | | Authors: | Bader, G, Martin, L.M, Steurer, S, Weiss-Puxbaum, A, Zoephel, A. | | Deposit date: | 2015-11-18 | | Release date: | 2016-03-09 | | Last modified: | 2016-06-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure-Based Design of an in Vivo Active Selective BRD9 Inhibitor.
J.Med.Chem., 59, 2016
|
|
5F25
 
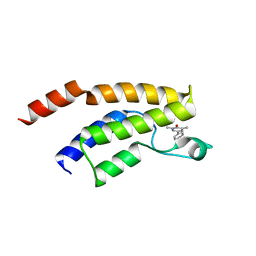 | | Crystal structure of the BRD9 bromodomain in complex with compound 4. | | Descriptor: | 4-(1,5-dimethyl-6-oxidanylidene-pyridin-3-yl)benzamide, BRD9 | | Authors: | Bader, G, Martin, L.J, Steurer, S, Weiss-Puxbaum, A, Zoephel, A. | | Deposit date: | 2015-12-01 | | Release date: | 2016-03-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Structure-Based Design of an in Vivo Active Selective BRD9 Inhibitor.
J.Med.Chem., 59, 2016
|
|
5F1H
 | | Crystal structure of the BRD9 bromodamian in complex with BI-9564. | | Descriptor: | 4-[4-[(dimethylamino)methyl]-2,5-dimethoxy-phenyl]-2-methyl-2,7-naphthyridin-1-one, Bromodomain-containing protein 9 | | Authors: | Bader, G, Martin, L.J, Steurer, S, Weiss-Puxbaum, A, Zoephel, A. | | Deposit date: | 2015-11-30 | | Release date: | 2016-03-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structure-Based Design of an in Vivo Active Selective BRD9 Inhibitor.
J.Med.Chem., 59, 2016
|
|
5F1L
 | | Crystal structure of the bromodomain of BRD9 in complex with compound 9. | | Descriptor: | 5-[3,5-dimethoxy-4-[(3-oxidanylazetidin-1-yl)methyl]phenyl]-1,3-dimethyl-pyridin-2-one, Bromodomain-containing protein 9 | | Authors: | Bader, G, Martin, L.J, Steurer, S, Weiss-Puxbaum, A, Zoephel, A. | | Deposit date: | 2015-11-30 | | Release date: | 2016-03-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure-Based Design of an in Vivo Active Selective BRD9 Inhibitor.
J.Med.Chem., 59, 2016
|
|
4EOW
 
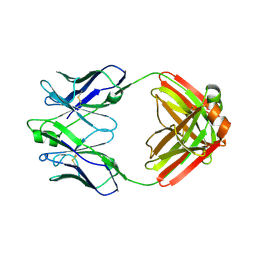 | | Crystal structure of a disease-associated anti-human GM-CSF autoantibody MB007 | | Descriptor: | MB007 IgG1 Fab fragment light chain, MB007 human IgG1 Fab fragment heavy chain | | Authors: | Blech, M. | | Deposit date: | 2012-04-16 | | Release date: | 2012-08-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Molecular structure of human GM-CSF in complex with a disease-associated anti-human GM-CSF autoantibody and its potential biological implications.
Biochem.J., 447, 2012
|
|
2FT6
 | |
2FT8
 
 | |
2FT7
 | |
2FTA
 
 | |
4G6J
 
 | |
