1NVW
 
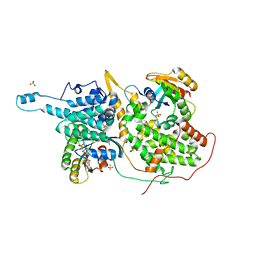 | | Structural evidence for feedback activation by RasGTP of the Ras-specific nucleotide exchange factor SOS | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Margarit, S.M, Sondermann, H, Hall, B.E, Nagar, B, Hoelz, A, Pirruccello, M, Bar-Sagi, D, Kuriyan, J. | | Deposit date: | 2003-02-04 | | Release date: | 2003-04-01 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural evidence for feedback activation by RasGTP of the Ras-specific nucleotide exchange factor SOS
Cell(Cambridge,Mass.), 112, 2003
|
|
1NVU
 
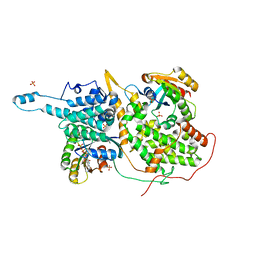 | | Structural evidence for feedback activation by RasGTP of the Ras-specific nucleotide exchange factor SOS | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Margarit, S.M, Sondermann, H, Hall, B.E, Nagar, B, Hoelz, A, Pirruccello, M, Bar-Sagi, D, Kuriyan, J. | | Deposit date: | 2003-02-04 | | Release date: | 2003-04-01 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural evidence for feedback activation by RasGTP of the Ras-specific nucleotide exchange factor SOS
Cell(Cambridge,Mass.), 112, 2003
|
|
1M52
 
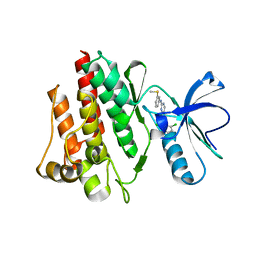 | | Crystal Structure of the c-Abl Kinase domain in complex with PD173955 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 6-(2,6-DICHLORO-PHENYL)-8-METHYL-2-(3-METHYLSULFANYL-PHENYLAMINO)-8H-PYRIDO[2,3-D]PYRIMIDIN-7-ONE, PROTO-ONCOGENE TYROSINE-PROTEIN KINASE ABL1 | | Authors: | Nagar, B, Bornmann, W, Pellicena, P, Schindler, T, Veach, D, Miller, W.T, Clarkson, B, Kuriyan, J. | | Deposit date: | 2002-07-08 | | Release date: | 2002-09-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structures of the Kinase Domain of c-Abl in Complex with the Small Molecule Inhibitors PD173955 and Imatinib (STI-571)
Cancer Res., 62, 2002
|
|
1QCF
 
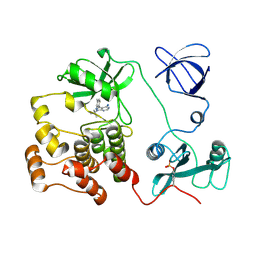 | | CRYSTAL STRUCTURE OF HCK IN COMPLEX WITH A SRC FAMILY-SELECTIVE TYROSINE KINASE INHIBITOR | | Descriptor: | 1-TER-BUTYL-3-P-TOLYL-1H-PYRAZOLO[3,4-D]PYRIMIDIN-4-YLAMINE, Tyrosine-protein kinase HCK | | Authors: | Schindler, T, Sicheri, F, Pico, A, Gazit, A, Levitzki, A, Kuriyan, J. | | Deposit date: | 1999-05-04 | | Release date: | 1999-06-08 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of Hck in complex with a Src family-selective tyrosine kinase inhibitor.
Mol.Cell, 3, 1999
|
|
1Q9C
 
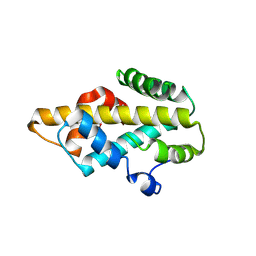 | |
1FPU
 
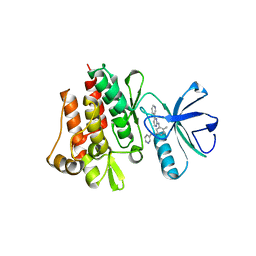 | | CRYSTAL STRUCTURE OF ABL KINASE DOMAIN IN COMPLEX WITH A SMALL MOLECULE INHIBITOR | | Descriptor: | N-[4-METHYL-3-[[4-(3-PYRIDINYL)-2-PYRIMIDINYL]AMINO]PHENYL]-3-PYRIDINECARBOXAMIDE, PROTO-ONCOGENE TYROSINE-PROTEIN KINASE ABL | | Authors: | Schindler, T, Bornmann, W, Pellicena, P, Miller, W.T, Clarkson, B, Kuriyan, J. | | Deposit date: | 2000-08-31 | | Release date: | 2000-09-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural mechanism for STI-571 inhibition of abelson tyrosine kinase.
Science, 289, 2000
|
|
1EQN
 
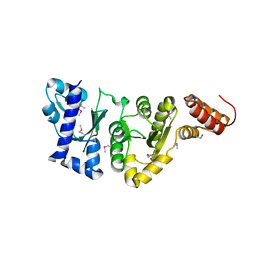 | | E.COLI PRIMASE CATALYTIC CORE | | Descriptor: | DNA PRIMASE | | Authors: | Podobnik, M, McInerney, P, O'Donnell, M, Kuriyan, J. | | Deposit date: | 2000-04-05 | | Release date: | 2000-06-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | A TOPRIM domain in the crystal structure of the catalytic core of Escherichia coli primase confirms a structural link to DNA topoisomerases.
J.Mol.Biol., 300, 2000
|
|
1DBH
 
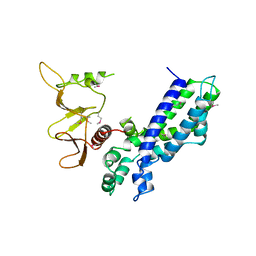 | | DBL AND PLECKSTRIN HOMOLOGY DOMAINS FROM HSOS1 | | Descriptor: | PROTEIN (HUMAN SOS 1) | | Authors: | Soisson, S.M, Kuriyan, J. | | Deposit date: | 1998-12-17 | | Release date: | 1998-12-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the Dbl and pleckstrin homology domains from the human Son of sevenless protein.
Cell(Cambridge,Mass.), 95, 1998
|
|
1DKG
 
 | |
1D5A
 
 | | CRYSTAL STRUCTURE OF AN ARCHAEBACTERIAL DNA POLYMERASE D.TOK. DEPOSITION OF SECOND NATIVE STRUCTURE AT 2.4 ANGSTROM | | Descriptor: | MAGNESIUM ION, PROTEIN (DNA POLYMERASE), SULFATE ION | | Authors: | Zhao, Y, Jeruzalmi, D, Leighton, L, Lasken, R, Kuriyan, J. | | Deposit date: | 1999-10-06 | | Release date: | 2000-03-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of an archaebacterial DNA polymerase.
Structure Fold.Des., 7, 1999
|
|
1FJL
 
 | | HOMEODOMAIN FROM THE DROSOPHILA PAIRED PROTEIN BOUND TO A DNA OLIGONUCLEOTIDE | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*AP*TP*CP*TP*GP*AP*TP*TP*AP*C)-3'), DNA (5'-D(*TP*GP*TP*AP*AP*TP*CP*AP*GP*AP*TP*TP*AP*T)-3'), DNA (5'-D(*TP*GP*TP*AP*AP*TP*CP*TP*GP*AP*TP*TP*AP*C)-3'), ... | | Authors: | Wilson, D.S, Guenther, B, Desplan, C, Kuriyan, J. | | Deposit date: | 1995-12-17 | | Release date: | 1996-06-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | High resolution crystal structure of a paired (Pax) class cooperative homeodomain dimer on DNA.
Cell(Cambridge,Mass.), 82, 1995
|
|
1QA4
 | |
4XI2
 
 | |
8CZI
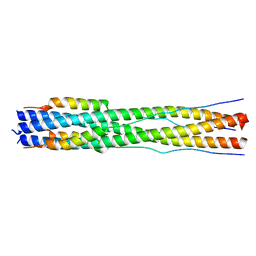 
 | |
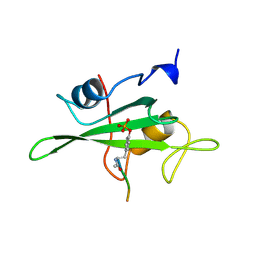
1HCT
 
 | |
1HCS
 
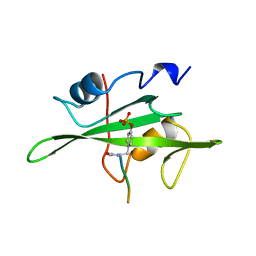 | |
5NLV
 
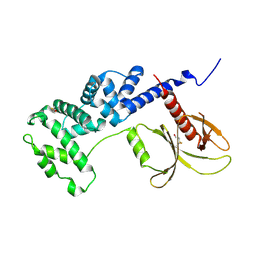 | | Brag2 Sec7-PH (390-763) | | Descriptor: | IQ motif and SEC7 domain-containing protein 1 | | Authors: | Nawrotek, A, Cherfils, J. | | Deposit date: | 2017-04-05 | | Release date: | 2017-09-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Multiple interactions between an Arf/GEF complex and charged lipids determine activation kinetics on the membrane.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5NLY
 
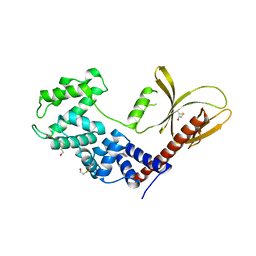 | | Brag2 Sec7-PH (390-763), P212121 | | Descriptor: | IQ motif and SEC7 domain-containing protein 1, PHOSPHATE ION | | Authors: | Nawrotek, A, Cherfils, J. | | Deposit date: | 2017-04-05 | | Release date: | 2017-09-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Multiple interactions between an Arf/GEF complex and charged lipids determine activation kinetics on the membrane.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
8UK9
 
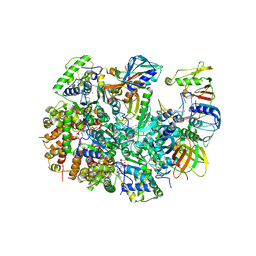 | | Structure of T4 Bacteriophage clamp loader mutant D110C bound to the T4 clamp, primer-template DNA, and ATP analog | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ALUMINUM FLUORIDE, DNA primer, ... | | Authors: | Marcus, K, Ghaffari-Kashani, S, Gee, C.L. | | Deposit date: | 2023-10-12 | | Release date: | 2023-12-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Autoinhibition of a clamp-loader ATPase revealed by deep mutagenesis and cryo-EM.
Nat.Struct.Mol.Biol., 31, 2024
|
|
8UH7
 
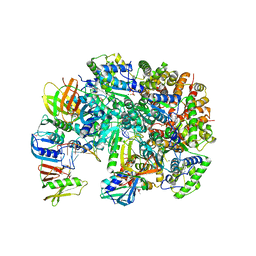 | | Structure of T4 Bacteriophage clamp loader bound to the T4 clamp, primer-template DNA, and ATP analog | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Primer DNA strand, ... | | Authors: | Gee, C.L, Marcus, K, Kelch, B.A, Makino, D.L. | | Deposit date: | 2023-10-07 | | Release date: | 2023-12-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.628 Å) | | Cite: | Autoinhibition of a clamp-loader ATPase revealed by deep mutagenesis and cryo-EM.
Nat.Struct.Mol.Biol., 31, 2024
|
|
4FTH
 
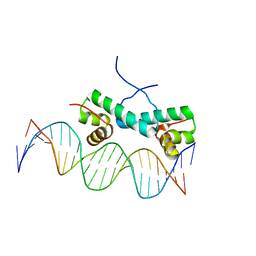 | | Crystal Structure of NtrC4 DNA-binding domain bound to double-stranded DNA | | Descriptor: | 5'-D(*AP*CP*TP*TP*GP*CP*AP*AP*AP*TP*TP*TP*GP*CP*AP*AP*AP*TP*GP*CP*AP*T)-3', 5'-D(P*GP*AP*TP*GP*CP*AP*TP*TP*TP*GP*CP*AP*AP*AP*TP*TP*TP*GP*CP*AP*A)-3', Transcriptional regulator (NtrC family) | | Authors: | Vidangos, N.K, Heideker, J, Lyubimov, A.Y, Lamers, M, Huo, Y, Pelton, J.G, Ton, J, Gralla, J.D, Kuriyan, J, Berger, J.M, Wemmer, D.E. | | Deposit date: | 2012-06-27 | | Release date: | 2012-08-29 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.004 Å) | | Cite: | DNA Recognition by a sigma (54) Transcriptional Activator from Aquifex aeolicus.
J.Mol.Biol., 426, 2014
|
|
4D8D
 
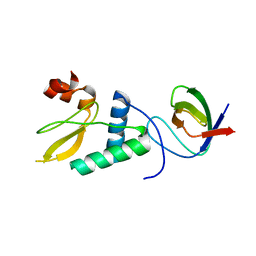 | |
7ROY
 
 | | The structure of the Fem1B:FNIP1 complex | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Folliculin-interacting protein 1, Protein fem-1 homolog B, ... | | Authors: | Gee, C.L, Mena, E.L, Manford, A.G, Rape, M. | | Deposit date: | 2021-08-02 | | Release date: | 2021-10-13 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural basis and regulation of the reductive stress response.
Cell, 184, 2021
|
|
6M94
 
 | | Monophosphorylated pSer33 b-Catenin peptide bound to b-TrCP/Skp1 Complex | | Descriptor: | Catenin beta-1, DIMETHYL SULFOXIDE, F-box/WD repeat-containing protein 1A, ... | | Authors: | Simonetta, K.R, Clifton, M.C, Walter, R.L, Ranieri, G.M, Carter, J.J. | | Deposit date: | 2018-08-22 | | Release date: | 2019-04-03 | | Last modified: | 2019-04-10 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Prospective discovery of small molecule enhancers of an E3 ligase-substrate interaction.
Nat Commun, 10, 2019
|
|
6M91
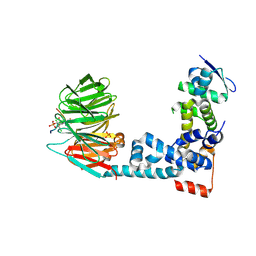 
 | | Monophosphorylated pSer33 b-Catenin peptide, b-TrCP/Skp1, NRX-103094 ternary complex | | Descriptor: | 3-({4-[(2,6-dichlorophenyl)sulfanyl]-2-oxo-6-(trifluoromethyl)-1,2-dihydropyridine-3-carbonyl}amino)benzoic acid, CHLORIDE ION, Catenin beta-1, ... | | Authors: | Simonetta, K.R, Clifton, M.C, Walter, R.L, Ranieri, G.M, Carter, J.J. | | Deposit date: | 2018-08-22 | | Release date: | 2019-04-03 | | Last modified: | 2019-04-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Prospective discovery of small molecule enhancers of an E3 ligase-substrate interaction.
Nat Commun, 10, 2019
|
|
